16S ribosomal RNA
This article is missing information about Rfam SSU_rRNA_bacteria, SSU_rRNA_archaea. (December 2020) |
16
The genes coding for it are referred to as 16S rRNA genes and are used in reconstructing
Functions
- Like the large (23S) ribosomal RNA, it has a structural role, acting as a scaffold defining the positions of the ribosomal proteins.
- The protein synthesis[5]
- Interacts with 23S, aiding in the binding of the two ribosomal subunits (30S)
- Stabilizes correct codon-anticodon pairing in the A-site by forming a hydrogen bond between the N1 atom of adenine residues 1492 and 1493 and the 2′OH group of the mRNA backbone.
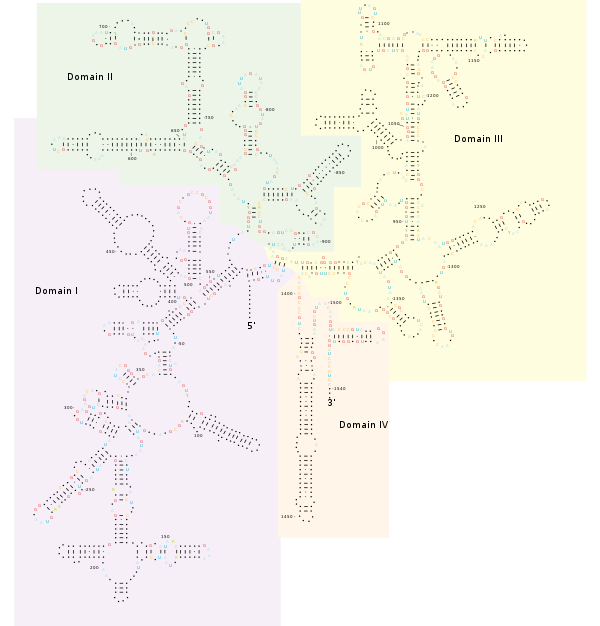
Structure
Universal primers
The 16S rRNA gene is used for
The most common primer pair was devised by Weisburg et al. (1991)[7] and is currently referred to as 27F and 1492R; however, for some applications shorter amplicons may be necessary, for example for 454 sequencing with titanium chemistry the primer pair 27F-534R covering V1 to V3.[12] Often 8F is used rather than 27F. The two primers are almost identical, but 27F has an M instead of a C. AGAGTTTGATCMTGGCTCAG compared with 8F.[13]
| Primer name | Sequence (5′–3′) | Ref. |
|---|---|---|
| 8F | AGA GTT TGA TCC TGG CTC AG | [14][15] |
| 27F | AGA GTT TGA TCM TGG CTC AG | [13] |
| 336R | ACT GCT GCS YCC CGT AGG AGT CT | [16] |
| 337F | GAC TCC TAC GGG AGG CWG CAG | [17] |
| 518R | GTA TTA CCG CGG CTG CTG G | |
| 533F | GTG CCA GCM GCC GCG GTA A | |
| 785F | GGA TTA GAT ACC CTG GTA | |
| 806R | GGA CTA CVS GGG TAT CTA AT | [18][19] |
| 907R | CCG TCA ATT CCT TTR AGT TT | |
| 928F | TAA AAC TYA AAK GAA TTG ACG GG | [16] |
| 1100F | YAA CGA GCG CAA CCC | |
| 1100R | GGG TTG CGC TCG TTG | |
| U1492R | GGT TAC CTT GTT ACG ACT T | [14][15] |
| 1492R | CGG TTA CCT TGT TAC GAC TT | [20] |
PCR and NGS applications
In addition to highly conserved primer binding sites, 16S rRNA gene sequences contain hypervariable regions that can provide species-specific signature sequences useful for identification of bacteria.[21][22] As a result, 16S rRNA gene sequencing has become prevalent in medical microbiology as a rapid and cheap alternative to phenotypic methods of bacterial identification.[23] Although it was originally used to identify bacteria, 16S sequencing was subsequently found to be capable of reclassifying bacteria into completely new species,[24] or even genera.[7][25] It has also been used to describe new species that have never been successfully cultured.[26][27] With
Hypervariable regions
The bacterial 16S gene contains nine hypervariable regions (V1–V9), ranging from about 30 to 100
While 16S hypervariable regions can vary dramatically between bacteria, the 16S gene as a whole maintains greater length homogeneity than its eukaryotic counterpart (
While 16S hypervariable region analysis is a powerful tool for bacterial taxonomic studies, it struggles to differentiate between closely related species.[34] In the families Enterobacteriaceae, Clostridiaceae, and Peptostreptococcaceae, species can share up to 99% sequence similarity across the full 16S gene.[36] As a result, the V4 sequences can differ by only a few nucleotides, leaving reference databases unable to reliably classify these bacteria at lower taxonomic levels.[36] By limiting 16S analysis to select hypervariable regions, these studies can fail to observe differences in closely related taxa and group them into single taxonomic units, therefore underestimating the total diversity of the sample.[34] Furthermore, bacterial genomes can house multiple 16S genes, with the V1, V2, and V6 regions containing the greatest intraspecies diversity.[8] While not the most precise method of classifying bacterial species, analysis of the hypervariable regions remains one of the most useful tools available to bacterial community studies.[36]
Promiscuity of 16S rRNA genes
Under the assumption that evolution is driven by
16S ribosomal databases
The 16S rRNA gene is used as the standard for classification and identification of microbes, because it is present in most microbes and shows proper changes.[41] Type strains of 16S rRNA gene sequences for most bacteria and archaea are available on public databases, such as NCBI. However, the quality of the sequences found on these databases is often not validated. Therefore, secondary databases that collect only 16S rRNA sequences are widely used. The most frequently used databases are listed below:
MIMt
MIMt is a compact non-redundant 16S database for a rapid metagenomic samples identification. It is composed of 39.940 full 16S sequences belonging to 17,625 well classified bacteria and archaea species. All sequences were obtained from complete genomes deposited in NCBI and for each of the sequences full taxonomic hierarchy is provided. It contains no redundancy, so only one representative for each species was considered avoiding same sequences from differente strains, isolates or patovars resulting in a very fast tool for microorganisms identification, compatible with any classification software (QIIME, Mothur, DADA, etc).[42]
EzBioCloud
EzBioCloud database, formerly known as EzTaxon, consists of a complete hierarchical taxonomic system containing 62,988 bacteria and archaea species/phylotypes which includes 15,290 valid published names as of September 2018. Based on the phylogenetic relationship such as maximum-likelihood and OrthoANI, all species/subspecies are represented by at least one 16S rRNA gene sequence. The EzBioCloud database is systematically curated and updated regularly which also includes novel candidate species. Moreover, the website provides bioinformatics tools such as ANI calculator, ContEst16S and 16S rRNA DB for QIIME and Mothur pipeline.[43]^^
Ribosomal Database Project
The Ribosomal Database Project (RDP) is a curated database that offers ribosome data along with related programs and services. The offerings include phylogenetically ordered alignments of ribosomal RNA (rRNA) sequences, derived phylogenetic trees, rRNA secondary structure diagrams and various software packages for handling, analyzing and displaying alignments and trees. The data are available via ftp and electronic mail. Certain analytic services are also provided by the electronic mail server.[44] Due to its large size the RDP database is often used as the basis for bioinformatic tool development and creating manually curated databases.[45]
SILVA
GreenGenes
GreenGenes is a quality controlled, comprehensive 16S rRNA gene reference database and taxonomy based on a de novo phylogeny that provides standard operational taxonomic unit sets. Beware that it utilizes taxonomic terms proposed from phylogenetic methods applied years ago between 2012 and 2013. Since then, a variety of novel phylogenetic methods have been proposed for Archaea and Bacteria.[47][48]
References
- S2CID 1024446.
- ^ PMID 270744.

- PMID 2112744.
- PMID 17071787.
- S2CID 22941368.
- PMID 2439888.
- ^ PMID 1987160.
- ^ PMID 14612235.
- PMID 28855596.
- PMID 26156036.
- PMID 33004967.
- ^ "Human Microbiome Project DACC - Home". www.hmpdacc.org. Archived from the original on 2010-10-30.
- ^ a b "Primers, 16S ribosomal DNA - François Lutzoni's Lab". lutzonilab.net. Archived from the original on 2012-12-27.
- ^ PMID 1854644.
- ^ ISBN 978-90-481-9038-6.
- ^ (PDF) from the original on 2011-07-15.
- PMID 34721319.
- S2CID 27232975.
- PMID 22267877.
- PMID 16751487.
- PMID 20923781.
- PMID 10383862.
- PMID 15489351.
- PMID 19395563.
- PMID 9542103.
- PMID 7520119.
- PMID 8899989.
- PMID 25226019.
- PMID 6462918.
- ^ PMID 27000765.
- PMID 21460107.
- ^ PMID 27688981.
- PMID 8811093.
- ^ PMID 23460914.
- ^ PMID 17391789.
- ^ PMID 27148170.
- PMID 23112186.
- PMID 28855596.
- PMID 31375780.
- PMID 31375780.
- S2CID 21895693.
- ^ "MIMt - (Mass Identification of Metagenomics tests)". mimt.bu.biopolis.pt. Retrieved 11 February 2024.
- ^ Yoon, S. H., Ha, S. M., Kwon, S., Lim, J., Kim, Y., Seo, H. and Chun, J. (2017). Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol. 67:1613–1617
- ^ Larsen N, Olsen GJ, Maidak BL, McCaughey MJ, Overbeek R, Macke TJ, Marsh TL, Woese CR. (1993) The ribosomal database project. Nucleic Acids Res. Jul 1;21(13):3021-3.
- PMID 26450747.
- ^ Elmar Pruesse, Christian Quast, Katrin Knittel, Bernhard M. Fuchs, Wolfgang Ludwig, Jörg Peplies, Frank Oliver Glöckner (2007) Nucleic Acids Res. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. December; 35(21): 7188–7196.
- PMID 16820507.
- PMID 22134646.
