5S ribosomal RNA
| 5S ribosomal RNA | |
|---|---|
| SO | SO:0000652 |
| PDB structures | PDBe |
The 5S ribosomal RNA (5S rRNA) is an approximately 120 nucleotide-long

Biosynthesis
In prokaryotes, the 5S rRNA gene is typically located in the rRNA
Structure
The secondary structure of 5S rRNA consists of five helices (denoted I–V in roman numerals), four loops (B-E), and one hinge (A), which form together a Y-like structure. Loops C and D are terminal hairpins and loops B and E are internal.[4] According to phylogenetic studies, helices I and III are likely ancestral.[9] Helix III includes two highly conserved adenosines.[10] Helix V, with its hairpin structure, is thought to interact with TFIIIA.[4]
Location within the ribosome
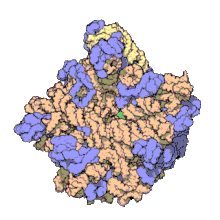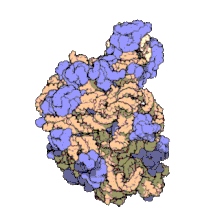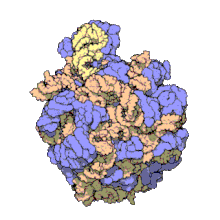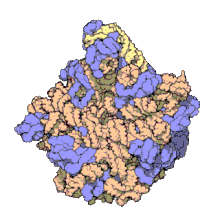
Using a variety of molecular techniques, including
In eukaryotes, the LSU contains 5S, 5.8S, and 28S rRNAs and even more proteins.[13][14] The structure of LSU in 3-dimensions shows one relatively smooth surface and the opposite surface having three projections, notably the L1 protuberance, the central protuberance (CP), and the L7/L12 stalk. The L1 protuberance and L7/L12 stalk are arranged laterally surrounding CP. The 5S rRNA is located in the CP and participates in formation and structure of this projection. The other major constituents of the central protuberance include the 23S rRNA (or alternatively 28S in eukaryotes) and several proteins including L5, L18, L25, and L27.[15]
Ribosomal functions
The exact function of 5S rRNA is not yet clear. In Escherichia coli, 5S rRNA gene deletions reduce the protein synthesis rate and have a more profound detrimental effect on cell fitness than deletions of a comparable number of copies of other (16S and 23S) rRNA genes.[16] Crystallographic studies indicate that 5S rRNA-binding proteins and other proteins of the central protuberance of the LSU plays a role in binding tRNAs.[15] Also, the topographical and physical proximity between 5S rRNA and 23S rRNA, which forms the peptidyl transferase and GTPase-associating center, suggests that 5S rRNA acts as a mediator between the two functional centers of the ribosome by forming, together with 5S rRNA-binding proteins and other components of the central protuberance, intersubunit bridges and tRNA-binding sites.[15]
Roles in ribosomal assembly
In eukaryotes, the cytosolic ribosome is assembled from four rRNAs and over 80 proteins.
Interactions with proteins
Several important proteins which interact with 5S rRNA are listed below.
La protein
Interaction of 5S rRNA with the La protein prevents the RNA from degradation by exonucleases in the cell.[19] La protein is found in the nucleus in all eukaryotic organisms and associates with several types of RNAs transcribed by RNA pol III. La protein interacts with these RNAs (including the 5S rRNA) through their 3' oligo-uridine tract, aiding stability and folding of the RNA.[4][20]
L5 protein
In eukaryotic cells, ribosomal protein L5 associates and stabilizes the 5S rRNA forming a pre-ribosomal ribonucleoprotein particle (RNP) that is found in both cytosol and the nucleus. L5 deficiency prevents transport of 5S rRNA to the nucleus and results in decreased ribosomal assembly.[4]
Other ribosomal proteins
In prokaryotes the 5S rRNA binds to the L5, L18 and L25 ribosomal proteins, whereas in eukaryotes 5S rRNA is only known to bind the
Presence in organelle ribosomes
| Permuted mitochondrial genome encoded 5S rRNA | |
|---|---|
| Identifiers | |
| Symbol | mtPerm-5S |
GO | GO:0005840 GO:0003735 |
| SO | SO:0000652 |
| PDB structures | PDBe |

Translation machineries of mitochondria and plastids (organelles of endosymbiotic bacterial origin), and their bacterial relatives share many features but also display marked differences. Organelle genomes encode SSU and LSU rRNAs without exception, yet the distribution of 5S rRNA genes (rrn5) is most uneven. Rrn5 is easily identified and common in genomes of most plastids. In contrast, mitochondrial rrn5 initially appeared to be restricted to plants and a small number of protists.[22][23] Additional, more divergent organellar 5S rRNAs were only identified with specialized covariance models that incorporate information on the pronounced sequence composition bias and structural variation.[24] This analysis pinpointed additional 5S rRNA genes not only in mitochondrial genomes of most protist lineages, but also in genomes of certain apicoplasts (non-photosynthetic plastids of pathogenic protozoa such as Toxoplasma gondii and Eimeria tenella).

Mitochondrial 5S rRNAs of most stramenopiles comprise the largest diversity of secondary structures.[24] The permuted mitochondrial 5S rRNAs in brown algae represent the most unconventional case, where the closing helix I, which otherwise brings together the molecule's 5′ and 3′ ends, is replaced by a (closed) hairpin resulting in an open three-way junction.
Current evidence indicates that mitochondrial DNA of only a few groups, notably animals, fungi, alveolates and euglenozoans lacks the gene.[24] The central protuberance, otherwise occupied by 5S rRNA and its associated proteins (see Figure 2), was remodeled in various ways. In the fungal mitochondrial ribosomes, 5S rRNA is replaced by LSU rRNA expansion sequences.[25] In kinetoplastids (euglenozoans), the central protuberance is made entirely of evolutionarily novel mitochondrial ribosomal proteins.[26] Lastly, animal mitochondrial ribosomes have coopted a specific mitochondrial tRNA (Val in vertebrates) to substitute the missing 5S rRNA.[27][28]
See also
References
- PMID 11752286.
- PMID 10756104.
- ^ PMID 17804668.
- ^ PMID 21957041.
- PMID 17487217.
- PMID 7731045.
- PMID 10926492.
- PMID 7001457.
- S2CID 21572917.
- PMID 11601989.
- PMID 10937989.
- PMID 32865340.
- PMID 25176256.
- ^ PMID 24580643.
- ^ S2CID 41669737.
- PMID 9862991.
- ^ S2CID 1934099.
- PMID 10716935.
- PMID 12045101.
- PMID 11868987.
- PMID 11258942.
- PMID 12592002.
- PMID 20304999.
- ^ PMID 25429974.
- PMID 24675956.
- PMID 19497863.
- PMID 25278503.
- S2CID 4402484.
External links
- Page for 5S ribosomal RNA at Rfam
- 5SData Archived 2010-04-27 at the Wayback Machine
- 5S+Ribosomal+RNA at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- 5S_rRNA human gene location in the UCSC Genome Browser.
- Halococcus morrhuae (archaebacterium) 5S rRNA
