Adeno-associated virus
| Adeno-associated virus | |
|---|---|
| |
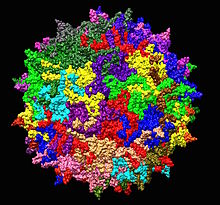
| Adeno-associated virus serotype 2 structure from 1LP3. One fivefold axis shown center. | |
| Scientific classification | |
| (unranked): | Virus |
| Realm: | Monodnaviria |
| Kingdom: | Shotokuvirae |
| Phylum: | Cossaviricota |
| Class: | Quintoviricetes |
| Order: | Piccovirales |
| Family: | Parvoviridae |
| Subfamily: | Parvovirinae |
| Genus: | Dependoparvovirus |
| Viruses included: | |
| |
Adeno-associated viruses (AAV) are small viruses that infect humans and some other
Several features make AAV an attractive candidate for creating
In March 2023, a series of Nature papers detected high titres of adeno-associated virus 2 (AAV2), alongside adenovirus and herpesvirus, in samples from a wave of childhood hepatitis.[6] One paper suggested that AAV2 co-infection may contribute to more serious liver disease than infection with only adeno- or herpesviruses and that the causal link remains to be established.[7]
History
The adeno-associated virus (AAV), previously thought to be a contaminant in adenovirus preparations, was first identified as a dependoparvovirus in the 1960s in the laboratories of Bob Atchison at Pittsburgh and Wallace Rowe at NIH. Serological studies in humans subsequently indicated that, despite being present in people infected by helper viruses such as adenovirus or herpes virus, AAV itself did not cause any disease.[8]
Use in gene therapy
Advantages and drawbacks
Wild-type AAV has attracted considerable interest from gene therapy researchers due to a number of features. Chief amongst these was the virus's apparent lack of pathogenicity. It can also infect non-dividing cells and has the ability to stably integrate into the host cell genome at a specific site (designated AAVS1) in the human
Use of the virus does present some disadvantages. The cloning capacity of the vector is relatively limited and most therapeutic genes require the complete replacement of the virus's 4.8 kilobase genome. Large genes are, therefore, not suitable for use in a standard AAV vector. Options are currently being explored to overcome the limited coding capacity.[15] The AAV ITRs of two genomes can anneal to form head-to-tail concatemers, almost doubling the capacity of the vector. Insertion of splice sites allows for the removal of the ITRs from the transcript.[citation needed]
Because of AAV's specialized gene therapy advantages, researchers have created an altered version of AAV termed
Humoral immunity instigated by infection with the wild type is thought to be common. The associated neutralising activity limits the usefulness of the most commonly used serotype AAV2 in certain applications. Accordingly, the majority of clinical trials under way involve delivery of AAV2 into the brain, a relatively immunologically privileged organ. In the brain, AAV2 is strongly neuron-specific.[citation needed]
Clinical trials
As of 2019, AAV vectors have been used in over 250 clinical trials worldwide, approximately 8.3% of virus-vectored gene-therapy trials.
| Indication | Gene | Route of administration | Phase | Subject number | Status |
| Cystic fibrosis | CFTR | Lung, via aerosol | I | 12 | Complete |
| CFTR | Lung, via aerosol | II | 38 | Complete | |
| CFTR | Lung, via aerosol | II | 100 | Complete | |
Hemophilia B |
FIX | Intramuscular | I | 9 | Complete |
| FIX | Hepatic artery | I | 6 | Ended | |
| Arthritis | TNFR:Fc |
Intraarticular | I | 1 | Ongoing |
| Hereditary emphysema | AAT |
Intramuscular | I | 12 | Ongoing |
Leber's congenital amaurosis |
RPE65 | Subretinal | I–II | Multiple | Several ongoing and complete (voretigene neparvovec) |
Age-related macular degeneration |
sFlt-1 |
Subretinal | I–II | 24 | Ongoing |
| Duchenne muscular dystrophy | SGCA | Intramuscular | I | 10 | Ongoing |
| Parkinson's disease | GAD67 |
Intracranial | I | 12 | Complete[27] |
| Canavan disease | AAC | Intracranial | I | 21 | Ongoing |
| Batten disease | CLN2 |
Intracranial | I | 10 | Ongoing |
| Alzheimer's disease | NGF | Intracranial | I | 6 | Ongoing |
| Spinal muscular atrophy | SMN1 | Intravenous, intrathecal | I–III | 150+ | Several ongoing and complete (onasemnogene abeparvovec) |
Congestive heart failure |
SERCA2a | Intra-coronary | IIb | 250 | Ongoing |
Fundamental biology
Genomics, transcriptomics and proteomics
The AAV genome is built of single-stranded deoxyribonucleic acid (ss
ITR sequences
The inverted terminal repeat (ITR) sequences comprise 145 bases each. They were named so because of their symmetry, which was shown to be required for efficient multiplication of the AAV genome.[29] The feature of these sequences that gives them this property is their ability to form a hairpin, which contributes to so-called self-priming that allows primase-independent synthesis of the second DNA strand. The ITRs were also shown to be required for both integration of the AAV DNA into the host cell genome (19th chromosome in humans) and rescue from it,[30][31] as well as for efficient encapsidation of the AAV DNA combined with generation of a fully assembled, deoxyribonuclease-resistant AAV particles.[32]
With regard to gene therapy, ITRs seem to be the only sequences required
rep gene and Rep proteins
On the "left side" of the genome there are two
cap gene and VP proteins
The right side of a positive-sensed AAV genome encodes overlapping sequences of three capsid proteins, VP1, VP2 and VP3, and two accessory proteins, MAAP & AAP, which start from one promoter, designated p40. The molecular weights of these proteins are 87, 72 and 62 kiloDaltons, respectively.[42] The AAV capsid is composed of a mixture of VP1, VP2, and VP3 totaling 60 monomers arranged in icosahedral symmetry in a ratio of 1:1:10,[43] with an empty mass of approximately 3.8 MDa.[44] The crystal structure of the VP3 protein was determined by Xie, Bue, et al.[45]

The cap gene produces an additional, non-structural protein called the Assembly-Activating Protein (AAP). This protein is produced from ORF2 and is essential for the capsid-assembly process.[46] The exact function of this protein in the assembly process and its structure have not been solved to date.[citation needed]
All three VPs are translated from one mRNA. After this mRNA is synthesized, it can be spliced in two different manners: either a longer or shorter intron can be excised resulting in the formation of two pools of mRNAs: a 2.3 kb- and a 2.6 kb-long mRNA pool. Usually, especially in the presence of adenovirus, the longer intron is preferred, so the 2.3-kb-long mRNA represents the so-called "major splice". In this form the first AUG codon, from which the synthesis of VP1 protein starts, is cut out, resulting in a reduced overall level of VP1 protein synthesis. The first AUG codon that remains in the major splice is the initiation codon for VP3 protein. However, upstream of that codon in the same open reading frame lies an ACG sequence (encoding threonine) which is surrounded by an optimal Kozak context. This contributes to a low level of synthesis of VP2 protein, which is actually VP3 protein with additional N terminal residues, as is VP1.[47][48][49][50]
Since the bigger intron is preferred to be spliced out, and since in the major splice the ACG codon is a much weaker translation initiation signal, the ratio at which the AAV structural proteins are synthesized in vivo is about 1:1:20, which is the same as in the mature virus particle.[51] The unique fragment at the N terminus of VP1 protein was shown to possess the phospholipase A2 (PLA2) activity, which is probably required for the releasing of AAV particles from late endosomes.[52] Muralidhar et al. reported that VP2 and VP3 are crucial for correct virion assembly.[49] More recently, however, Warrington et al. showed VP2 to be unnecessary for the complete virus particle formation and an efficient infectivity, and also presented that VP2 can tolerate large insertions in its N terminus, while VP1 can not, probably because of the PLA2 domain presence.[53]
Post-translational modifications
Recent discoveries made through use of high-throughput 'omics approaches include the fact that AAV capsids are post-translationally modified (PTM) during production such as acetylation, methylation, phosphorylation, deamidation, O-GlycNAcylation[54] and SUMOylation throughout capsid proteins VP1, VP2 and VP3. These PTMs differ depending on the manufacturing production platform. Another such discovery is the fact that AAV genomes are epigenetically methylated during production. Besides price, these findings might affect expression kinetics, rAAV receptor binding, trafficking, vector immunogenicity, and expression durability.[55][56]
Classification, serotypes, receptors and native tropism
| Serotype | Tissue tropism[57] | Pass the blood brain barrier | Cell tropism | Synthetic | Comment | Published cap / rep |
|---|---|---|---|---|---|---|
| 2 | smooth muscle, CNS, liver | No | No | |||
| 5 | CNS, smooth muscle | No | No | |||
| 8 | CNS, Brain, Liver, smooth muscle | No | No | |||
| 9 | CNS, liver, smooth muscle | Yes | No | |||
| PAL2 | CNS | Yes | Yes | Liver expression is 1/4 of AAV9 | No | |
| 9P1/AAVMYO | CNS | Yes | astrocytes | Yes | high musculature including skeletal muscle, heart and diaphragmsckeletal transduction | No |
Two species of AAV were recognised by the International Committee on Taxonomy of Viruses in 2013: adeno-associated dependoparvovirus A (formerly AAV-1, −2, −3 and −4) and adeno-associated dependoparvovirus B (formerly AAV-5).[58][59]
Until the 1990s, virtually all AAV biology was studied using AAV serotype 2. However, AAV is highly prevalent in humans and other primates and several serotypes have been isolated from various tissue samples. Serotypes 2, 3, 5, and 6 were discovered in human cells, AAV serotypes 1, 4, and 7–11 in nonhuman primate samples.[60] As of 2006 there have been 11 AAV serotypes described, the 11th in 2004.[61] AAV capsid proteins contain 12 hypervariable surface regions, with most variability occurring in the threefold proximal peaks, but the parvovirus genome in general presents highly conserved replication and structural genes across serotypes.[60] All of the known serotypes can infect cells from multiple diverse tissue types. Tissue specificity is determined by the capsid serotype and pseudotyping of AAV vectors to alter their tropism range will likely be important to their use in therapy.
Serotype 2
Serotype 2 (AAV2) has been the most extensively examined so far.[62][63][64][65][66][67] AAV2 presents natural tropism towards skeletal muscles,[68] neurons,[62] vascular smooth muscle cells[69] and hepatocytes.[70]
Three cell receptors have been described for AAV2: heparan sulfate proteoglycan (HSPG), aVβ5 integrin and fibroblast growth factor receptor 1 (FGFR-1). The first functions as a primary receptor, while the latter two have a co-receptor activity and enable AAV to enter the cell by receptor-mediated endocytosis.[71][72][73] These study results have been disputed by Qiu, Handa, et al.[74] HSPG functions as the primary receptor, though its abundance in the extracellular matrix can scavenge AAV particles and impair the infection efficiency.[75]
Studies have shown that serotype 2 of the virus (AAV-2) apparently kills cancer cells without harming healthy ones. "Our results suggest that adeno-associated virus type 2, which infects the majority of the population but has no known ill effects, kills multiple types of cancer cells yet has no effect on healthy cells," said Craig Meyers,
In March 2023, a series of Nature papers linked infection of adeno-associated virus 2 (AAV2) to a wave of childhood hepatitis.[6]
Other serotypes
This section needs to be updated. (December 2023) |
Although AAV2 is the most popular serotype in various AAV-based research, it has been shown that other serotypes can be more effective as gene delivery vectors. AAV9 passes the blood-brain-barrier in humans, AAV6 appears much better in infecting airway epithelial cells,[78][79] AAV7 presents very high transduction rate of murine skeletal muscle cells (similar to AAV1 and AAV5), AAV8 transduce hepatocytes[80][81][82] and AAV1 and 5 were shown to be very efficient in gene delivery to vascular endothelial cells.[83] In the brain, most AAV serotypes show neuronal tropism, while AAV5 also transduces astrocytes.[84] AAV6, a hybrid of AAV1 and AAV2,[82] also shows lower immunogenicity than AAV2.[81]
Serotypes can differ with the respect to the receptors they are bound to. For example, AAV4 and AAV5 transduction can be inhibited by soluble sialic acids (of different form for each of these serotypes),[85] and AAV5 was shown to enter cells via the platelet-derived growth factor receptor.[86]
Synthetic serotypes
There have been many efforts to engineer and improve new AAV variants for both clinical and research purposes. Such modifications include new tropisms to target specific tissues, and modified surface residues to evade detection by the immune system. Beyond opting for particular strains of recombinant AAV (rAAV) to target particular cells, researchers have also explored AAV pseudotyping, the practice of creating hybrids of certain AAV strains to approach an even more refined target. The hybrid is created by taking a capsid from one strain and the genome from another strain. For example, research involving AAV2/5, a hybrid with the genome of AAV2 and the capsid of AAV5, was able to achieve more accuracy and range in brain cells than AAV2 would be able to achieve unhybridized. Researchers have continued to experiment with pseudotyping by creating strains with hybrid capsids. AAV-DJ has a hybrid capsid from eight different strains of AAV; as such, it can infect different cells throughout many areas of the body, a property which a single strain of AAV with a limited tropism would not have.[87] Other efforts to engineer and improve new AAV variants have involved the ancestral reconstruction of virus variants to generate new vectors with enhanced properties for clinical applications and the study of AAV biology.[88]
Immunology
AAV is of particular interest to gene therapists due to its apparent limited capacity to induce immune responses in humans, a factor which should positively influence vector transduction efficiency while reducing the risk of any immune-associated pathology.
AAV is not considered to have any known role in disease.[89] However, host immune system response and immune tolerance reduce the efficacy of AAV-mediated gene therapy. Host immune response has been shown to respond to the AAV vectors, the transduced cells, and the transduced proteins.[90] The immune response can be subdivided into two categories: innate and adaptive, the latter of which is divided into humoral and cell-mediated.[91][92]
Innate
The
In-vivo studies indicate that AAV vectors interact with the Toll-like receptor (TLR)9- and TLR2-MyD88 pathways to trigger the innate immune response by stimulating the production of interferons.[94] It's shown that mice deficient in TLR9 are more receptive to AAV treatment and demonstrate higher levels of transgene expression[95]
Humoral
Due to previous natural infection, many people have preexisting neutralizing antibodies (NAbs) against AAV's, which can significantly hinder its application in gene therapy.[96] Even though AAV's are highly variable among wild-type and synthetic variants, antibody recognition sites may be conserved evolutionarily.[97]
The virus is known to instigate robust
Cell-mediated
The cell-mediated response to the virus and to vectors is poorly characterised, and has been largely ignored in the literature as recently as 2005.[98] Clinical trials using an AAV2-based vector to treat haemophilia B seem to indicate that targeted destruction of transduced cells may be occurring.[100] Combined with data that shows that CD8+ T-cells can recognise elements of the AAV capsid in vitro,[101] it appears that there may be a cytotoxic T lymphocyte response to AAV vectors. Cytotoxic responses would imply the involvement of CD4+ T helper cells in the response to AAV and in vitro data from human studies suggests that the virus may indeed induce such responses, including both Th1 and Th2 memory responses.[99] A number of candidate T cell stimulating epitopes have been identified within the AAV capsid protein VP1, which may be attractive targets for modification of the capsid if the virus is to be used as a vector for gene therapy.[99][100]
Infection cycle
There are several steps in the AAV infection cycle, from infecting a cell to producing new infectious particles:[citation needed]
- attachment to the cell membrane
- receptor-mediated endocytosis
- endosomal trafficking
- escape from the late endosome or lysosome
- translocation to the nucleus
- uncoating
- formation of double-stranded DNA replicative form of the AAV genome
- expression of rep genes
- genome replication
- expression of cap genes, synthesis of progeny ssDNA particles
- assembly of complete virions, and
- release from the infected cell.
Some of these steps may look different in various types of cells, which, in part, contributes to the defined and quite limited native tropism of AAV. Replication of the virus can also vary in one cell type, depending on the cell's current cell cycle phase.[102]
The characteristic feature of the adeno-associated virus is a deficiency in replication and thus its inability to multiply in unaffected cells. Adeno-associated virus spreads by co-infecting a cell with a helper virus. The first helper virus that was described as providing successful generation of new AAV particles, was the adenovirus, from which the AAV name originated. It was then shown that AAV replication can be facilitated by selected proteins derived from the adenovirus genome,
The minimal set of the adenoviral genes required for efficient generation, of progeny AAV particles, was discovered by Matsushita, Ellinger et al.[103] This discovery allowed for new production methods of recombinant AAV, which do not require adenoviral co-infection of the AAV-producing cells. In the absence of helper virus or genotoxic factors, AAV DNA can either integrate into the host genome or persist in episomal form. In the former case integration is mediated by Rep78 and Rep68 proteins and requires the presence of ITRs flanking the region being integrated. In mice, the AAV genome has been observed persisting for long periods of time in quiescent tissues, such as skeletal muscles, in episomal form (a circular head-to-tail conformation).[111]
See also
References
- PMID 28669112.
- PMID 19904234.
- PMID 16568890.
- PMID 19649989.
- ^ PMID 18441370.
- ^ PMID 36997704.
- PMID 36996871.
- PMID 15564130.
- PMID 2156265.
- PMID 9311886.
- ^ PMID 18854481.
- PMID 10490767.
- PMID 10482608.
- PMID 9185868.
- PMID 28723575.
- PMID 11509958.
- PMID 24460861.
- ^ "Vectors used in Gene Therapy Clinical Trials". Journal of Gene Medicine. Wiley. December 2019. Archived from the original on 21 October 2019. Retrieved 4 January 2012.
- PMID 18441371.
- PMID 18774912.
- PMID 22149959.
- PMID 21709064.
- Avexis. 6 May 2016. Archived from the originalon 11 June 2017. Retrieved 8 May 2016.
- PMID 22717743.
- S2CID 37154043.
- PMID 15916479.
- S2CID 24491886.
- ISBN 978-0-585-39515-9.
- PMID 2845646.
- PMID 7623375.
- ^ PMID 8016070.
- PMID 9525651.
- PMID 11559833.
- PMID 12477885.
- PMID 11991970.
- PMID 11090148.
- ^ PMID 8151765.
- S2CID 27997617.
- PMID 1309894.
- )
- PMID 2824856.
- PMID 6265925.
- PMID 20479244.
- PMID 27310298.
- PMID 12136130.
- PMID 21917944.
- PMID 2999784.
- PMID 2847413.
- ^ PMID 8254726.
- PMID 2841488.
- PMID 11118354.
- PMID 11961250.
- PMID 15163751.
- PMID 32995354.
- PMID 32669341.
- PMID 30343890.
- ^ "AAV tropisms".
- ^ "ICTV Taxonomy history: Adeno-associated dependoparvovirus A". ICTV. Retrieved 31 January 2020.
- ^ "ICTV Taxonomy history: Adeno-associated dependoparvovirus B". ICTV. Retrieved 31 January 2020.
- ^ ISBN 978-1-61779-370-7.
- PMID 15567432.
- ^ PMID 9625257.
- PMID 14664794.
- PMID 11545607.
- PMID 10600599.
- PMID 12727115.
- PMID 10954565.
- PMID 12515715.
- S2CID 14150468.
- PMID 9037069.
- S2CID 31602526.
- PMID 9445046.
- S2CID 23326070.
- PMID 10725206.
- PMID 12388714.
- ^ "Common virus 'kills cancer'". CNN. 22 June 2005. Retrieved 5 August 2009.
- ^ Penn State (23 June 2005). "Harmless Virus Kills Some Cancers". Science Daily.
- PMID 19066597.
- PMID 25845025.
- PMID 12192090.
- ^ PMID 11413329.
- ^ PMID 15078923.
- PMID 15761263.
- PMID 20418874.
- PMID 11435568.
- S2CID 32547188.
- ^ "Adeno-associated Virus (AAV) Guide".
- PMID 26235624.
- PMID 15871671.
- PMID 32362898.
- PMID 28835127.
- PMID 30807882.
- PMID 11932423.
- PMID 19587448.
- PMID 21474674.
- PMID 21775517.
- PMID 28559317.
- ^ PMID 15975009.
- ^ PMID 19641045.
- ^ S2CID 14664182.
- PMID 16263332.
- PMID 12270659.
- ^ PMID 9813665.
- PMID 6251278.
- PMID 221504.
- PMID 2835153.
- PMID 3029431.
- PMID 2536816.
- ^ "Introduction to Adeno-Associated Virus (AAV)".
- ^ "Adeno-Associated Virus and Adeno-associated Viral Vectors". Archived from the original on 20 September 2018. Retrieved 19 September 2018.
- PMID 9765395.
External links
- Kimball JW (17 May 2015). "Gene Therapy II". Kimball's Biology Pages. Archived from the original on 18 March 2005. Retrieved 13 May 2005.
- "Adeno-associated virus". NCBI Taxonomy Browser. 272636.
