Amino acid

Amino acids are
Amino acids can be classified according to the locations of the core structural functional groups (
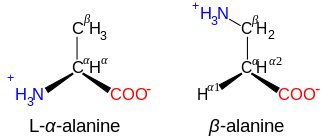
Amino acids are formally named by the IUPAC-IUBMB Joint Commission on Biochemical Nomenclature in terms of the fictitious "neutral" structure shown in the illustration. For example, the systematic name of alanine is 2-aminopropanoic acid, based on the formula CH3−CH(NH2)−COOH. The Commission justified this approach as follows:[6]
The systematic names and formulas given refer to hypothetical forms in which amino groups are unprotonated and carboxyl groups are undissociated. This convention is useful to avoid various nomenclatural problems but should not be taken to imply that these structures represent an appreciable fraction of the amino-acid molecules.
History
The first few amino acids were discovered in the early 1800s.
The unity of the chemical category was recognized by
General structure
2-, alpha-, or α-amino acids[20] have the generic formula H2NCHRCOOH in most cases,[b] where R is an organic substituent known as a "side chain".[21]
Of the many hundreds of described amino acids, 22 are proteinogenic ("protein-building").[22][23][24] It is these 22 compounds that combine to give a vast array of peptides and proteins assembled by ribosomes.[25] Non-proteinogenic or modified amino acids may arise from post-translational modification or during nonribosomal peptide synthesis.
Chirality
The
A few D-amino acids ("right-handed") have been found in nature, e.g., in
Side chains
Polar charged side chains
Five amino acids possess a charge at neutral pH. Often these side chains appear at the surfaces on proteins to enable their solubility in water, and side chains with opposite charges form important electrostatic contacts called
The two negatively charged amino acids at neutral pH are aspartate (Asp, D) and glutamate (Glu, E). The anionic carboxylate groups behave as Brønsted bases in most circumstances.[31] Enzymes in very low pH environments, like the aspartic protease pepsin in mammalian stomachs, may have catalytic aspartate or glutamate residues that act as Brønsted acids.

There are three amino acids with side chains that are cations at neutral pH: arginine (Arg, R), lysine (Lys, K) and histidine (His, H). Arginine has a charged guanidino group and lysine a charged alkyl amino group, and are fully protonated at pH 7. Histidine's imidazole group has a pKa of 6.0, and is only around 10 % protonated at neutral pH. Because histidine is easily found in its basic and conjugate acid forms it often participates in catalytic proton transfers in enzyme reactions.[31]
Polar uncharged side chains
The polar, uncharged amino acids serine (Ser, S), threonine (Thr, T), asparagine (Asn, N) and glutamine (Gln, Q) readily form hydrogen bonds with water and other amino acids.[31] They do not ionize in normal conditions, a prominent exception being the catalytic serine in serine proteases. This is an example of severe perturbation, and is not characteristic of serine residues in general. Threonine has two chiral centers, not only the L (2S) chiral center at the α-carbon shared by all amino acids apart from achiral glycine, but also (3R) at the β-carbon. The full stereochemical specification is (2S,3R)-L-threonine.
Hydrophobic side chains
Nonpolar amino acid interactions are the primary driving force behind the processes that fold proteins into their functional three dimensional structures.[31] None of these amino acids' side chains ionize easily, and therefore do not have pKas, with the exception of tyrosine (Tyr, Y). The hydroxyl of tyrosine can deprotonate at high pH forming the negatively charged phenolate. Because of this one could place tyrosine into the polar, uncharged amino acid category, but its very low solubility in water matches the characteristics of hydrophobic amino acids well.
Special case side chains
Several side chains are not described well by the charged, polar and hydrophobic categories.
β- and γ-amino acids
Amino acids with the structure NH+3−CXY−CXY−CO−2, such as
Zwitterions

The common natural forms of amino acids have a
In strongly acidic conditions (pH below 3), the carboxylate group becomes protonated and the structure becomes an ammonio carboxylic acid, NH+3−CHR−CO2H. This is relevant for enzymes like pepsin that are active in acidic environments such as the mammalian stomach and
Although various definitions of acids and bases are used in chemistry, the only one that is useful for chemistry in aqueous solution is that of Brønsted:[36][37] an acid is a species that can donate a proton to another species, and a base is one that can accept a proton. This criterion is used to label the groups in the above illustration. The carboxylate side chains of aspartate and glutamate residues are the principal Brønsted bases in proteins. Likewise, lysine, tyrosine and cysteine will typically act as a Brønsted acid. Histidine under these conditions can act both as a Brønsted acid and a base.
Isoelectric point

For amino acids with uncharged side-chains the zwitterion predominates at pH values between the two pKa values, but coexists in equilibrium with small amounts of net negative and net positive ions. At the midpoint between the two pKa values, the trace amount of net negative and trace of net positive ions balance, so that average net charge of all forms present is zero.[38] This pH is known as the isoelectric point pI, so pI = 1/2(pKa1 + pKa2).
For amino acids with charged side chains, the pKa of the side chain is involved. Thus for aspartate or glutamate with negative side chains, the terminal amino group is essentially entirely in the charged form −NH+3, but this positive charge needs to be balanced by the state with just one C-terminal carboxylate group is negatively charged. This occurs halfway between the two carboxylate pKa values: pI = 1/2(pKa1 + pKa(R)), where pKa(R) is the side chain pKa.[37]
Similar considerations apply to other amino acids with ionizable side-chains, including not only glutamate (similar to aspartate), but also cysteine, histidine, lysine, tyrosine and arginine with positive side chains.
Amino acids have zero mobility in electrophoresis at their isoelectric point, although this behaviour is more usually exploited for peptides and proteins than single amino acids. Zwitterions have minimum solubility at their isoelectric point, and some amino acids (in particular, with nonpolar side chains) can be isolated by precipitation from water by adjusting the pH to the required isoelectric point.
Physicochemical properties
The 20 canonical amino acids can be classified according to their properties. Important factors are charge,
Some amino acids have special properties. Cysteine can form covalent
to the polypeptide backbone, and glycine is more flexible than other amino acids.Glycine and proline are strongly present within low complexity regions of both eukaryotic and prokaryotic proteins, whereas the opposite is the case with cysteine, phenylalanine, tryptophan, methionine, valine, leucine, isoleucine, which are highly reactive, or complex, or hydrophobic.[39][41][42]
Many proteins undergo a range of
Table of standard amino acid abbreviations and properties
Although one-letter symbols are included in the table, IUPAC–IUBMB recommend[6] that "Use of the one-letter symbols should be restricted to the comparison of long sequences".
The one-letter notation was chosen by IUPAC-IUB based on the following rules:[46]
- Initial letters are used where there is no ambuiguity: C cysteine, H histidine, I isoleucine, M methionine, S serine, V valine,[46]
- Where arbitrary assignment is needed, the structurally simpler amino acids are given precedence: A Alanine, G glycine, L leucine, P proline, T threonine,[46]
- F PHenylalanine and R aRginine are assigned by being phonetically suggestive,[46]
- W tryptophane is assigned based on the double ring being visually suggestive to the bulky letter W,[46]
- K lysine and Y tyrosine are assigned as alphabetically nearest to their initials L and T (note that U was avoided for its similarity with V, while X was reserved for undetermined or atypical amino acids); for tyrosine the mnemonic tYrosine was also proposed,[47]
- D aspartate was assigned arbitrarily, with the proposed mnemonic asparDic acid;[48] E glutamate was assigned in alphabetical sequence being larger by merely one methylene –CH2– group,[47]
- N asparagine was assigned arbitrarily, with the proposed mnemonic asparagiNe;[48] Q glutamine was assigned in alphabetical sequence of those still available (note again that O was avoided due to similarity with D), with the proposed mnemonic Qlutamine.[48]
| Amino acid | 3- and 1-letter symbols | Side chain | Hydropathy
index[49] |
Molar absorptivity[50]
|
Molecular mass | Abundance in proteins (%)[51] |
Standard genetic coding, IUPAC notation | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | 1 | Class | Chemical polarity[52] | Net charge at pH 7.4[52] |
Wavelength, λmax (nm) |
Coefficient ε (mM−1·cm−1) | |||||
| Alanine | Ala | A | Aliphatic | Nonpolar | Neutral | 1.8 | 89.094 | 8.76 | GCN | ||
| Arginine | Arg | R | Fixed cation | Basic polar | Positive | −4.5 | 174.203 | 5.78 | MGR, CGY[53] | ||
| Asparagine | Asn | N | Amide | Polar | Neutral | −3.5 | 132.119 | 3.93 | AAY | ||
Aspartate
|
Asp | D | Anion | Brønsted base
|
Negative | −3.5 | 133.104 | 5.49 | GAY | ||
| Cysteine | Cys | C | Thiol | Brønsted acid | Neutral | 2.5 | 250 | 0.3 | 121.154 | 1.38 | UGY |
| Glutamine | Gln | Q | Amide | Polar | Neutral | −3.5 | 146.146 | 3.9 | CAR | ||
Glutamate
|
Glu | E | Anion | Brønsted base | Negative | −3.5 | 147.131 | 6.32 | GAR | ||
| Glycine | Gly | G | Aliphatic | Nonpolar | Neutral | −0.4 | 75.067 | 7.03 | GGN | ||
| Histidine | His | H | Cationic | Brønsted acid and base | Positive, 10% Neutral, 90% |
−3.2 | 211 | 5.9 | 155.156 | 2.26 | CAY |
| Isoleucine | Ile | I | Aliphatic | Nonpolar | Neutral | 4.5 | 131.175 | 5.49 | AUH | ||
| Leucine | Leu | L | Aliphatic | Nonpolar | Neutral | 3.8 | 131.175 | 9.68 | YUR, CUY[54] | ||
| Lysine | Lys | K | Cation | Brønsted acid | Positive | −3.9 | 146.189 | 5.19 | AAR | ||
| Methionine | Met | M | Thioether | Nonpolar | Neutral | 1.9 | 149.208 | 2.32 | AUG | ||
| Phenylalanine | Phe | F | Aromatic | Nonpolar | Neutral | 2.8 | 257, 206, 188 | 0.2, 9.3, 60.0 | 165.192 | 3.87 | UUY |
| Proline | Pro | P | Cyclic | Nonpolar | Neutral | −1.6 | 115.132 | 5.02 | CCN | ||
| Serine | Ser | S | Hydroxylic | Polar | Neutral | −0.8 | 105.093 | 7.14 | UCN, AGY | ||
| Threonine | Thr | T | Hydroxylic | Polar | Neutral | −0.7 | 119.119 | 5.53 | ACN | ||
| Tryptophan | Trp | W | Aromatic | Nonpolar | Neutral | −0.9 | 280, 219 | 5.6, 47.0 | 204.228 | 1.25 | UGG |
| Tyrosine | Tyr | Y | Aromatic | Brønsted acid | Neutral | −1.3 | 274, 222, 193 | 1.4, 8.0, 48.0 | 181.191 | 2.91 | UAY |
| Valine | Val | V | Aliphatic | Nonpolar | Neutral | 4.2 | 117.148 | 6.73 | GUN | ||
Two additional amino acids are in some species coded for by
| 21st and 22nd amino acids | 3-letter | 1-letter | Molecular mass |
|---|---|---|---|
| Selenocysteine | Sec | U | 168.064 |
| Pyrrolysine | Pyl | O | 255.313 |
In addition to the specific amino acid codes, placeholders are used in cases where chemical or crystallographic analysis of a peptide or protein cannot conclusively determine the identity of a residue. They are also used to summarize conserved protein sequence motifs. The use of single letters to indicate sets of similar residues is similar to the use of abbreviation codes for degenerate bases.[55][56]
| Ambiguous amino acids | 3-letter | 1-letter | Amino acids included | Codons included |
|---|---|---|---|---|
| Any / unknown | Xaa | X | All | NNN |
aspartate
|
Asx | B | D, N | RAY |
glutamate
|
Glx | Z | E, Q | SAR |
| Leucine or isoleucine | Xle | J | I, L | YTR, ATH, CTY[57] |
Hydrophobic
|
Φ | V, I, L, F, W, Y, M | NTN, TAY, TGG | |
Aromatic
|
Ω | F, W, Y, H | YWY, TTY, TGG[58] | |
Aliphatic (non-aromatic)
|
Ψ | V, I, L, M | VTN, TTR[59] | |
| Small | π | P, G, A, S | BCN, RGY, GGR | |
Hydrophilic
|
ζ | S, T, H, N, Q, E, D, K, R | VAN, WCN, CGN, AGY[60] | |
Positively-charged
|
+ | K, R, H | ARR, CRY, CGR | |
Negatively-charged
|
− | D, E | GAN |
Unk is sometimes used instead of Xaa, but is less standard.
Ter or * (from termination) is used in notation for mutations in proteins when a stop codon occurs. It corresponds to no amino acid at all.[61]
In addition, many
Occurrence and functions in biochemistry
Proteinogenic amino acids
Amino acids are the precursors to proteins.[25] They join by condensation reactions to form short polymer chains called peptides or longer chains called either polypeptides or proteins. These chains are linear and unbranched, with each amino acid residue within the chain attached to two neighboring amino acids. In nature, the process of making proteins encoded by DNA/RNA genetic material is called translation and involves the step-by-step addition of amino acids to a growing protein chain by a ribozyme that is called a ribosome.[63] The order in which the amino acids are added is read through the genetic code from an mRNA template, which is an RNA copy of one of the organism's genes.
Twenty-two amino acids are naturally incorporated into polypeptides and are called
Several independent evolutionary studies have suggested that Gly, Ala, Asp, Val, Ser, Pro, Glu, Leu, Thr may belong to a group of amino acids that constituted the early genetic code, whereas Cys, Met, Tyr, Trp, His, Phe may belong to a group of amino acids that constituted later additions of the genetic code.[67][68][69]
Standard vs nonstandard amino acids
The 20 amino acids that are encoded directly by the codons of the universal genetic code are called standard or canonical amino acids. A modified form of methionine (N-formylmethionine) is often incorporated in place of methionine as the initial amino acid of proteins in bacteria, mitochondria and chloroplasts. Other amino acids are called nonstandard or non-canonical. Most of the nonstandard amino acids are also non-proteinogenic (i.e. they cannot be incorporated into proteins during translation), but two of them are proteinogenic, as they can be incorporated translationally into proteins by exploiting information not encoded in the universal genetic code.
The two nonstandard proteinogenic amino acids are selenocysteine (present in many non-eukaryotes as well as most eukaryotes, but not coded directly by DNA) and
Non-proteinogenic amino acids
Aside from the 22
Non-proteinogenic amino acids that are found in proteins are formed by post-translational modification. Such modifications can also determine the localization of the protein, e.g., the addition of long hydrophobic groups can cause a protein to bind to a phospholipid membrane.[78] Examples:
- the glutamate allows for better binding of calcium cations,[79]
- Hydroxyproline, generated by hydroxylation of proline, is a major component of the connective tissue collagen.[80]
- Hypusine in the translation initiation factor EIF5A, contains a modification of lysine.[81]
Some non-proteinogenic amino acids are not found in proteins. Examples include
In mammalian nutrition
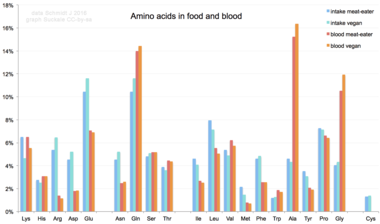
Amino acids are not typical component of food: animals eat proteins. The protein is broken down into amino acids in the process of digestion. They are then used to synthesize new proteins, other biomolecules, or are oxidized to urea and carbon dioxide as a source of energy.[84] The oxidation pathway starts with the removal of the amino group by a transaminase; the amino group is then fed into the urea cycle. The other product of transamidation is a keto acid that enters the citric acid cycle.[85] Glucogenic amino acids can also be converted into glucose, through gluconeogenesis.[86]
Of the 20 standard amino acids, nine (His, Ile, Leu, Lys, Met, Phe, Thr, Trp and Val) are called essential amino acids because the human body cannot synthesize them from other compounds at the level needed for normal growth, so they must be obtained from food.[87][88][89]
Semi-essential and conditionally essential amino acids, and juvenile requirements
In addition, cysteine, tyrosine, and arginine are considered semiessential amino acids, and taurine a semi-essential aminosulfonic acid in children. Some amino acids are conditionally essential for certain ages or medical conditions. Essential amino acids may also vary from species to species.[d] The metabolic pathways that synthesize these monomers are not fully developed.[90][91]
Non-protein functions
Many proteinogenic and non-proteinogenic amino acids have biological functions beyond being precursors to proteins and peptides.In humans, amino acids also have important roles in diverse biosynthetic pathways. Defenses against herbivores in plants sometimes employ amino acids.[95] Examples:
Standard amino acids
- Tryptophan is a precursor of the neurotransmitter serotonin.[96]
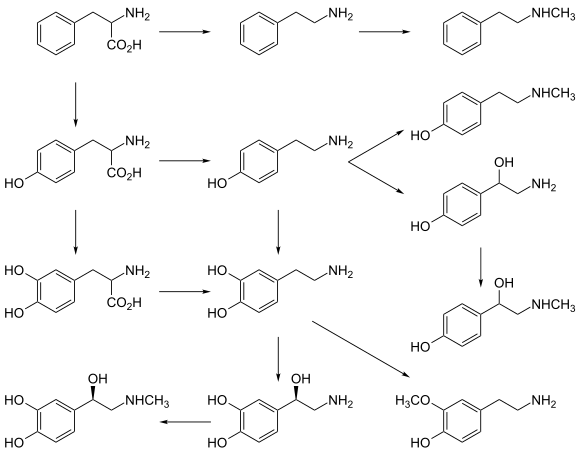
- epinephrine and norepinephrine and various trace amines.
- Phenylalanine is a precursor of phenethylamine and tyrosine in humans. In plants, it is a precursor of various phenylpropanoids, which are important in plant metabolism.
- Glycine is a precursor of porphyrins such as heme.[97]
- Arginine is a precursor of nitric oxide.[98]
- Ornithine and S-adenosylmethionine are precursors of polyamines.[99]
- However, not all of the functions of other abundant nonstandard amino acids are known.
Roles for nonstandard amino acids
- Carnitine is used in lipid transport.
- gamma-aminobutyric acid is a neurotransmitter.[101]
- 5-HTP (5-hydroxytryptophan) is used for experimental treatment of depression.[102]
- Parkinson's treatment,[103]
- Eflornithine inhibits ornithine decarboxylase and used in the treatment of sleeping sickness.[104]
- Canavanine, an analogue of arginine found in many legumes is an antifeedant, protecting the plant from predators.[105]
- Mimosine found in some legumes, is another possible antifeedant.[106] This compound is an analogue of tyrosine and can poison animals that graze on these plants.
Uses in industry
Animal feed
Amino acids are sometimes added to
Food
The
Chemical building blocks
Amino acids are low-cost
Amino acids are used in the synthesis of some cosmetics.[107]
Aspirational uses
Fertilizer
The chelating ability of amino acids is sometimes used in fertilizers to facilitate the delivery of minerals to plants in order to correct mineral deficiencies, such as iron chlorosis. These fertilizers are also used to prevent deficiencies from occurring and to improve the overall health of the plants.[114]
Biodegradable plastics
Amino acids have been considered as components of biodegradable polymers, which have applications as
Synthesis

Chemical synthesis
The commercial production of amino acids usually relies on mutant bacteria that overproduce individual amino acids using glucose as a carbon source. Some amino acids are produced by enzymatic conversions of synthetic intermediates.
Biosynthesis
In plants, nitrogen is first assimilated into organic compounds in the form of
Nonstandard amino acids are usually formed through modifications to standard amino acids. For example,
Primordial synthesis
The formation of amino acids and peptides are assumed to precede and perhaps induce the emergence of life on earth. Amino acids can form from simple precursors under various conditions.[124] Surface-based chemical metabolism of amino acids and very small compounds may have led to the build-up of amino acids, coenzymes and phosphate-based small carbon molecules.[125][additional citation(s) needed] Amino acids and similar building blocks could have been elaborated into proto-peptides, with peptides being considered key players in the origin of life.[126]
In the famous
According to a review, amino acids, and even peptides, "turn up fairly regularly in the
Reactions
Amino acids undergo the reactions expected of the constituent functional groups.[130][131]
Peptide bond formation

As both the amine and carboxylic acid groups of amino acids can react to form amide bonds, one amino acid molecule can react with another and become joined through an amide linkage. This polymerization of amino acids is what creates proteins. This condensation reaction yields the newly formed peptide bond and a molecule of water. In cells, this reaction does not occur directly; instead, the amino acid is first activated by attachment to a transfer RNA molecule through an ester bond. This aminoacyl-tRNA is produced in an ATP-dependent reaction carried out by an aminoacyl tRNA synthetase.[132] This aminoacyl-tRNA is then a substrate for the ribosome, which catalyzes the attack of the amino group of the elongating protein chain on the ester bond.[133] As a result of this mechanism, all proteins made by ribosomes are synthesized starting at their N-terminus and moving toward their C-terminus.
However, not all peptide bonds are formed in this way. In a few cases, peptides are synthesized by specific enzymes. For example, the tripeptide
In chemistry, peptides are synthesized by a variety of reactions. One of the most-used in solid-phase peptide synthesis uses the aromatic oxime derivatives of amino acids as activated units. These are added in sequence onto the growing peptide chain, which is attached to a solid resin support.[136] Libraries of peptides are used in drug discovery through high-throughput screening.[137]
The combination of functional groups allow amino acids to be effective polydentate ligands for metal–amino acid chelates.[138] The multiple side chains of amino acids can also undergo chemical reactions.
Catabolism
* Amino acids catabolized into both glucogenic and ketogenic products.
Degradation of an amino acid often involves
Complexation
Amino acids are bidentate ligands, forming transition metal amino acid complexes.[140]
Chemical analysis
The total nitrogen content of organic matter is mainly formed by the amino groups in proteins. The Total Kjeldahl Nitrogen (
See also
Notes
- ^ The late discovery is explained by the fact that cysteine becomes oxidized to cystine in air.
- ^ Proline and other cyclic amino acids are an exception to this general formula. Cyclization of the α-amino acid creates the corresponding secondary amine. These are occasionally referred to as imino acids.
- ^ The L and D convention for amino acid configuration refers not to the optical activity of the amino acid itself but rather to the optical activity of the isomer of glyceraldehyde from which that amino acid can, in theory, be synthesized (D-glyceraldehyde is dextrorotatory; L-glyceraldehyde is levorotatory). An alternative convention is to use the (S) and (R) designators to specify the absolute configuration.[29] Almost all of the amino acids in proteins are (S) at the α carbon, with cysteine being (R) and glycine non-chiral.[30] Cysteine has its side chain in the same geometric location as the other amino acids, but the R/S terminology is reversed because sulfur has higher atomic number compared to the carboxyl oxygen which gives the side chain a higher priority by the Cahn-Ingold-Prelog sequence rules.
- microbes in the first two stomach chambers.
References
- ISBN 0-7167-4339-6.
- PMID 31691799.
- ^ Richard Cammack, ed. (2009). "Newsletter 2009". Biochemical Nomenclature Committee of IUPAC and NC-IUBMB. Pyrrolysine. Archived from the original on 12 September 2017. Retrieved 16 April 2012.
- PMID 20847933.
- ^ Latham MC (1997). "Chapter 8. Body composition, the functions of food, metabolism and energy". Human nutrition in the developing world. Food and Nutrition Series – No. 29. Rome: Food and Agriculture Organization of the United Nations. Archived from the original on 8 October 2012. Retrieved 9 September 2012.
- ^ a b c "Nomenclature and Symbolism for Amino Acids and Peptides". IUPAC-IUB Joint Commission on Biochemical Nomenclature. 1983. Archived from the original on 9 October 2008. Retrieved 17 November 2008.
- .
- ^ Hansen S (May 2015). "Die Entdeckung der proteinogenen Aminosäuren von 1805 in Paris bis 1935 in Illinois" (PDF) (in German). Berlin. Archived from the original (PDF) on 1 December 2017.
- ^ Vauquelin LN, Robiquet PJ (1806). "The discovery of a new plant principle in Asparagus sativus". Annales de Chimie. 57: 88–93.
- ^ ISBN 978-0-12-034226-6.
- S2CID 110151163.
- ^ Baumann E (1884). "Über cystin und cystein". Z Physiol Chem. 8 (4): 299–305. Archived from the original on 14 March 2011. Retrieved 28 March 2011.
- ^ Braconnot HM (1820). "Sur la conversion des matières animales en nouvelles substances par le moyen de l'acide sulfurique". Annales de Chimie et de Physique. 2nd Series. 13: 113–125.
- from the original on 10 June 2019. Retrieved 4 July 2015.
- .
- ^ Menten, P. Dictionnaire de chimie: Une approche étymologique et historique. De Boeck, Bruxelles. link Archived 28 December 2019 at the Wayback Machine.
- ^ Harper D. "amino-". Online Etymology Dictionary. Archived from the original on 2 December 2017. Retrieved 19 July 2010.
- doi:10.1002/cber.189402701205. Archived from the originalon 25 July 2020.
- ISBN 978-0-87169-191-0.
- ^ "Alpha amino acid". Merriam-Webster Medical. Archived from the original on 3 January 2015. Retrieved 3 January 2015..
- ^ Clark, Jim (August 2007). "An introduction to amino acids". chemguide. Archived from the original on 30 April 2015. Retrieved 4 July 2015.
- ISBN 9783527621170. Archivedfrom the original on 17 May 2016. Retrieved 5 January 2016 – via Google Books.
- S2CID 3705304.
- S2CID 5359077.

- ^ a b "Chapter 1: Proteins are the Body's Worker Molecules". The Structures of Life. National Institute of General Medical Sciences. 27 October 2011. Archived from the original on 7 June 2014. Retrieved 20 May 2008.
- ISBN 978-0-470-14684-2.
- ^ ISBN 978-0-7167-7030-5.
- S2CID 254088816.
- .
- ^ Hatem SM (2006). "Gas chromatographic determination of Amino Acid Enantiomers in tobacco and bottled wines". University of Giessen. Archived from the original on 22 January 2009. Retrieved 17 November 2008.
- ^ OCLC 297392560.
- S2CID 265572280.
- PMID 17508906.
- S2CID 35519996.
- ISBN 978-0126654509.
- .
- ^ OCLC 61448218.
- ISBN 978-0-8247-9691-4.
- ^ PMID 31504783.
- .
- PMID 10512723.
- PMID 20962881.
- PMID 15780596.
- PMID 17500024.
- PMID 15189153.
- ^ .
- ^ .
- ^ .
- PMID 7108955.
- ISBN 978-0-7167-1315-9.[page needed]
- PMID 27789699.
- ^ ISBN 978-0-87893-214-6.
- ^ Codons can also be expressed by: CGN, AGR
- ^ codons can also be expressed by: CUN, UUR
- PMID 11911894.
- PMID 5080161.
- ^ Codons can also be expressed by: CTN, ATH, TTR; MTY, YTR, ATA; MTY, HTA, YTG
- ^ Codons can also be expressed by: TWY, CAY, TGG
- ^ Codons can also be expressed by: NTR, VTY
- ^ Codons can also be expressed by: VAN, WCN, MGY, CGP
- ^ "HGVS: Sequence Variant Nomenclature, Protein Recommendations". Archived from the original on 24 September 2021. Retrieved 23 September 2021.
- PMID 15782218.
- PMID 17157507.
- PMID 12524431.
- PMID 16256420.
- PMID 16164991.
- PMID 1057181.
- PMID 11164045.
- S2CID 9039622.
- from the original on 23 July 2018. Retrieved 12 June 2019.
- S2CID 1944741.
- ^ Tjong H (2008). Modeling Electrostatic Contributions to Protein Folding and Binding (PhD thesis). Florida State University. p. 1 footnote. Archived from the original on 28 January 2020. Retrieved 28 January 2020.
- from the original on 14 April 2021. Retrieved 5 January 2016.
- ^ Elzanowski A, Ostell J (7 April 2008). "The Genetic Codes". National Center for Biotechnology Information (NCBI). Archived from the original on 20 August 2016. Retrieved 10 March 2010.
- PMID 16260173.
- PMID 19318213.
- ISBN 978-0-387-22046-8.
- PMID 8129952.
- PMID 2183788.
- S2CID 7211864.
- PMID 16452303.
- S2CID 23877884.
- PMID 16042590.
- PMID 14144484.
- PMID 10736367.
- PMID 11533293.
- PMID 8064412.
- PMID 15173430.
- PMID 10867060.
- PMID 9437700.
- PMID 12514918.
- PMID 19948186.
- PMID 15860375.
- PMID 24374199.
- .
- PMID 18923670.
- from the original on 7 May 2022. Retrieved 3 November 2008.
- PMID 18815130.
- PMID 16709566.
- ^ ISBN 978-0-7167-4684-3.
- S2CID 84891972.
- from the original on 13 April 2020. Retrieved 12 June 2019.
- S2CID 33603501.
- S2CID 26273053.
- S2CID 3144019.
- from the original on 7 May 2022. Retrieved 7 May 2022.
- ^ S2CID 24161808.
- ^ Ashmead HD (1993). The Role of Amino Acid Chelates in Animal Nutrition. Westwood: Noyes Publications.
- PMID 10736350.
- PMID 3300262.
- ^ ISBN 978-3527306732.
- S2CID 43992655.
- .
- ^ Ashmead HD (1986). Foliar Feeding of Plants with Amino Acid Chelates. Park Ridge: Noyes Publications.
- .
- from the original on 25 July 2020. Retrieved 12 June 2019.
- ^ Low KC, Wheeler AP, Koskan LP (1996). Commercial poly(aspartic acid) and Its Uses. Advances in Chemistry Series. Vol. 248. Washington, D.C.: American Chemical Society.
- S2CID 94818855.
- ISBN 978-0-943088-39-6.
- PMID 16702333.
- PMID 9559057.
- S2CID 24638475.
- PMID 12324528.
- ^ S2CID 102659869.
- PMID 28684991.
- ^ S2CID 211536416.
- PMID 31641431.
- PMID 35743865.
- S2CID 227037164.
- ISBN 978-0-521-46827-5.
- PMID 16214343.
- PMID 11375928.
- PMID 4896351.
- PMID 14988435.
- PMID 3053703.
- .
- PMID 18855697.
- S2CID 94822047.
- ^ Stipanuk MH (2006). Biochemical, physiological, & molecular aspects of human nutrition (2nd ed.). Saunders Elsevier.
- PMID 29712039.
- PMID 23959242.
- doi:10.4141/S01-054.
Further reading
- Tymoczko JL (2012). "Protein Composition and Structure". Biochemistry. New York: W. H. Freeman and company. pp. 28–31. ISBN 9781429229364.
- LCCN 89008555.
- Nelson DL, Cox MM (2000). Lehninger Principles of Biochemistry (3rd ed.). LCCN 99049137.
- LCCN 2008930865. Archived from the original(PDF) on 12 January 2012.
External links
 Media related to Amino acids at Wikimedia Commons
Media related to Amino acids at Wikimedia Commons