Apolipoprotein E
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 19: 44.91 – 44.91 Mb | Chr 7: 19.43 – 19.43 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Apolipoprotein E (Apo-E) is a protein involved in the metabolism of fats in the body of mammals. A subtype is implicated in Alzheimer's disease and cardiovascular diseases.[5] It is encoded in humans by the gene APOE.
Apo-E belongs to a family of fat-binding proteins called
Evolution
Apolipoproteins are not unique to mammals. Many terrestrial and marine
The three major human alleles (E4, E3, E2) arose after the primate-human split around 7.5 million years ago. These alleles are the by-product of non-synonymous mutations which led to changes in functionality. The first allele to emerge was E4. After the primate-human split, there were four amino acid changes in the human lineage, three of which had no effect on protein function (V174L, A18T, A135V). The fourth substitution (T61R) traded a threonine for an arginine altering the protein's functionality. This substitution occurred somewhere in the 6 million year gap between the primate-human split and the Denisovan-human split, since exactly the same substitutions were found in Denisovan APOE.[14]
About 220,000 years ago, a cysteine to arginine substitution took place at amino acid 112 (Cys112Arg) of the APOE4 gene, and this resulted in the E3 allele. Finally, 80,000 years ago, another arginine to cysteine substitution at amino acid 158 (Arg158Cys) of the APOE3 gene created the E2 allele.[15][13]
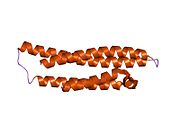
Structure
Gene
The gene, APOE, is mapped to
Protein
APOE is 299
Polymorphisms
| SNP: rs429358 | |
|---|---|
HapMap | 429358 |
| SNPedia | 429358 |
APOE is polymorphic,[19][20] with three major alleles (epsilon 2, epsilon 3, and epsilon 4): APOE-ε2 (cys112, cys158), APOE-ε3 (cys112, arg158), and APOE-ε4 (arg112, arg158).[5][21][22] Although these allelic forms differ from each other by only one or two amino acids at positions 112 and 158,[23][24][25] these differences alter APOE structure and function.
There are several low-frequency polymorphisms of APOE. APOE5 comes in two subtypes E5f and E5s, based on migration rates. APOE5 E5f and APOE7 combined were found in 2.8% of Japanese males.[26][unreliable medical source] APOE7 is a mutation of APOE3 with two lysine residues replacing glutamic acid residues at positions 244 and 245.[27]
| Polymorphism | Worldwide allele frequency | Disease relevance |
|---|---|---|
| ε2 (rs7412-T, rs429358-T) | 8.4%[9] | This variant of the apoprotein binds poorly to cell surface receptors while E3 and E4 bind well. type III hyperlipoproteinemia—94.4% of people with such disease are E2/E2 but only ~2% of E2/E2 develop it, so other environmental and genetic factors are likely to be involved (such as cholesterol in the diet and age).[29][30][31] E2 has also been implicated in Parkinson's disease,[32] but this finding was not replicated in a larger population association study.[33]
|
| ε3 (rs7412-C, rs429358-T) | 77.9%[9] | This variant is considered the "neutral" APOE genotype. |
| ε4 (rs7412-C, rs429358-C) | 13.7%[9] |
E4 has been implicated in |
Much remains to be learned about the APOE isoforms, including the interaction of other protective genes.[58] Indeed, the apolipoprotein ε4 isoform is more protective against cognitive decline than other isoforms in some cases,[58] so caution is advised before making determinant statements about the influence of APOE polymorphisms on cognition, development of Alzheimer's disease, cardiovascular disease, telomere shortening, etc. Many of the studies cited that purport these adverse outcomes are from single studies that have not been replicated and the research is based on unchecked assumptions about this isoform. As of 2007, there was no evidence that APOE polymorphisms influence cognition in younger age groups (other than possible increased episodic memory ability and neural efficiency in younger APOE4 age groups), nor that the APOE4 isoform places individuals at increased risk for any infectious disease.[59]
However, the association between the APOE4 allele and Alzheimer's disease has been shown to be weaker in minority groups differently compared to their Caucasian counterparts.
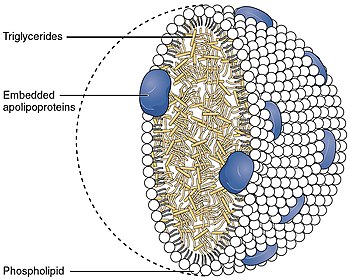
Function
As a component of the lipoprotein lipid transport system, APOE facilitates the transport of
In the nervous system, non-neuronal cell types, most notably astroglia and microglia, are the primary producers of APOE, while neurons preferentially express the receptors for APOE.[62] There are seven currently identified mammalian receptors for APOE which belong to the evolutionarily conserved LDLR family.[63]
APOE was initially recognized for its importance in lipoprotein
In the field of immune regulation, a growing number of studies point to APOE's interaction with many immunological processes, including suppressing
Clinical significance
Alzheimer's disease
As of 2012, the E4 variant was the largest known genetic risk factor for late-onset sporadic
Recently, the amyloid hypothesis of Alzheimer's disease has been questioned, and an article in
Although 40–65% of AD patients have at least one copy of the ε4 allele, APOE4 is not a determinant of the disease. At least one-third of patients with AD are APOE4 negative and some APOE4 homozygotes never develop the disease. Yet those with two ε4 alleles have up to 20 times the risk of developing AD.[78] There is also evidence that the APOE2 allele may serve a protective role in AD.[79] Thus, the genotype most at risk for Alzheimer's disease and at an earlier age is APOE4,4. Using genotype APOE3,3 as a benchmark (with the persons who have this genotype regarded as having a risk level of 1.0) and for white populations only, individuals with genotype APOE4,4 have an odds ratio of 14.9 of developing Alzheimer's disease. Individuals with the APOE3,4 genotype face an odds ratio of 3.2, and people with a copy of the 2 allele and the 4 allele (APOE2,4), have an odds ratio of 2.6. Persons with one copy each of the 2 allele and the 3 allele (APOE2,3) have an odds ratio of 0.6. Persons with two copies of the 2 allele (APOE2,2) also have an odds ratio of 0.6.[80]
| Estimated worldwide human allele frequencies of APOE in Caucasian population[80] | ||||
| Allele | ε2 | ε3 | ε4 | |
|---|---|---|---|---|
| General frequency | 8.4% | 77.9% | 13.7% | |
| AD frequency | 3.9% | 59.4% | 36.7% | |
While ApoE4 has been found to greatly increase the odds that an individual will develop Alzheimer's, a 2002 study concluded, that in persons with any combination of APOE alleles, high serum total cholesterol and high blood pressure in mid-life are independent risk factors which together can nearly triple the risk that the individual will later develop AD.[73] Projecting from their data, some researchers have suggested that lowering serum cholesterol levels may reduce a person's risk for Alzheimer's disease, even if they have two ApoE4 alleles, thus reducing the risk from nine or ten times the odds of getting AD down to just two times the odds.[73]
Women are more likely to develop AD than men across most ages and APOE genotypes. Premorbid women with the ε4 allele have significantly more neurological dysfunction than men.[81]
APOE-ε4 increases the risk not only for AD but also for dementia in pure alpha-synucleinopathies.[82] The influence of APOE-ε4 on hippocampal atrophy was suggested to be more predominant early in the course of AD at milder stages prior to more widespread neurodegeneration.[83]
Atherosclerosis
Knockout mice that lack the apolipoprotein-E gene (APOE−/−) develop extreme hypercholesterolemia when fed a high-fat diet.[84]
Malaria
APOE−/− knockout mice show marked attenuation of cerebral malaria and increased survival, as well as decreased sequestration of parasites and T cells within the brain, likely due to protection of the blood–brain barrier.[85] Human studies have shown that the APOE2 polymorphism correlates with earlier infection, and APOE3/4 polymorphisms increase likelihood of severe malaria.[86]
Lyme disease
Borrelia burgdorferi, the causative agent of Lyme disease, is a host-adapted pathogen that acquires environmental cholesterol to form glycolipids for use in cell membrane maintenance. In one experiment in 2015, mice engineered with apoE deficiency were infected with Borrelia spirochetes. The knockout mice suffered from an increased spirochete burden in joints, as well as inflamed ankles, when compared with wild-type mice. This study suggests that apoE deficiency (and potentially other hyperlipidemias) may be a risk factor in the pathogenicity of Lyme disease.
Interactions
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "Statin_Pathway_WP430".
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000130203 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000002985 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ ISBN 978-3540686989.
- PMID 19106071.
- ^ a b "Entrez Gene: APOE apolipoprotein E".
- S2CID 220044671.
- ^ PMID 23296339.
- S2CID 5407666.
- PMID 30692699.
- PMID 9238027.
- ^ S2CID 3758905.
- PMID 23112842.
- S2CID 14225962.
- PMID 11172721.
- S2CID 24698373.
- S2CID 6159310.
- S2CID 34696595.
- PMID 20734437.
- ^ ISBN 978-1849964715.
- PMID 16204452.
- ^ Online Mendelian Inheritance in Man (OMIM): APOE3 isoform, hyperlipoproteinemia, type III, autosomal recessive - 107741#0015
- ^ Online Mendelian Inheritance in Man (OMIM): APOE3 isoform, APOE, CYS112 and ARG158 - 107741#0001
- PMID 16603077.
- S2CID 20361600.
- PMID 11060347.
- PMID 6277903.
- PMID 7175379.
- S2CID 39103399.
- PMID 9125318.
- S2CID 10445412.
- PMID 22349451.
- S2CID 6159310.
- PMID 11701639.
- PMID 8346443.
- PMID 8446617.
- S2CID 4418270.
- S2CID 21131734.
- S2CID 236451232.
- PMID 18562290.
- S2CID 40761206.
- PMID 11836653.
- S2CID 131908791.
- S2CID 23443430.
- PMID 11401610.
- S2CID 12280483.
- PMID 23418430.
- S2CID 27820056.
- S2CID 73120632.
- PMID 31541246.
- PMID 18395206.
- PMID 32451547.
- S2CID 22483645.
- PMID 25673673.
- PMID 20401432.
- PMID 18280677.
- ^ a b Sundermann EE, Wang C, Katz M, et al. Cholesteryl ester transfer protein genotype modifies the effect of apolipoprotein ε4 on memory decline in older adults. Neurobiol Aging. 2016;41:200.e7-200.e12. doi:10.1016/j.neurobiolaging.2016.02.006
- PMID 17077159.
- ^ PMID 28373857.
- PMID 35641666.
- PMID 23171142.
- PMID 19674510.
- PMID 25551803.
- S2CID 42716198.
- S2CID 3109596.
- PMID 20182542.
- PMID 24244577.
- PMID 23022896.
- ^ PMID 2491016.
- ^ S2CID 71543885.
- S2CID 35969696.
- ^ S2CID 23780605.
- S2CID 8839627.
- PMID 18549781.
- Lay summary in: "Mechanism Explains Link Between Apolipoprotein E And Alzheimer's Disease". ScienceDaily (Press release). June 13, 2008.
- S2CID 240152869.
- ^ PMID 35008886.
- PMID 23919769.
- S2CID 11137478.
- ^ PMID 9343467.
- PMID 22699906.
- PMID 23407718.
- S2CID 236451232.
- PMID 20210786.
- PMID 27647324.
- PMID 14985370.
Further reading
- Ashford JW (2004). "APOE genotype effects on Alzheimer's disease onset and epidemiology". Journal of Molecular Neuroscience. 23 (3): 157–165. S2CID 14864342.
- Beffert U, Danik M, Krzywkowski P, Ramassamy C, Berrada F, Poirier J (July 1998). "The neurobiology of apolipoproteins and their receptors in the CNS and Alzheimer's disease". Brain Research. Brain Research Reviews. 27 (2): 119–142. S2CID 28731779.
- Bennet AM, Di Angelantonio E, Ye Z, Wensley F, Dahlin A, Ahlbom A, et al. (September 2007). "Association of apolipoprotein E genotypes with lipid levels and coronary risk". JAMA. 298 (11): 1300–1311. PMID 17878422.
- Bocksch L, Stephens T, Lucas A, Singh B (December 2001). "Apolipoprotein E: possible therapeutic target for atherosclerosis". Current Drug Targets - Cardiovascular & Hematological Disorders. 1 (2): 93–106. PMID 12769659.
- de Knijff P, van den Maagdenberg AM, Frants RR, Havekes LM (1995). "Genetic heterogeneity of apolipoprotein E and its influence on plasma lipid and lipoprotein levels". Human Mutation. 4 (3): 178–194. S2CID 41959843.
- Gunzburg MJ, Perugini MA, Howlett GJ (December 2007). "Structural basis for the recognition and cross-linking of amyloid fibrils by human apolipoprotein E". The Journal of Biological Chemistry. 282 (49): 35831–35841. PMID 17916554.
- Huang Y, Weisgraber KH, Mucke L, Mahley RW (2004). "Apolipoprotein E: diversity of cellular origins, structural and biophysical properties, and effects in Alzheimer's disease". Journal of Molecular Neuroscience. 23 (3): 189–204. S2CID 40135107.
- Itzhaki RF, Dobson CB, Shipley SJ, Wozniak MA (June 2004). "The role of viruses and of APOE in dementia". Annals of the New York Academy of Sciences. 1019 (1): 15–18. S2CID 28979273.
- Kolbe D, da Silva NA, Dose J, Torres GG, Caliebe A, Krause-Kyora B, Nebel A (May 2023). "Current allele distribution of the human longevity gene APOE in Europe can mainly be explained by ancient admixture". Aging Cell. 22 (5). Wiley: e13819. PMID 36951219.
- Kolovou GD, Anagnostopoulou KK (August 2007). "Apolipoprotein E polymorphism, age and coronary heart disease". Ageing Research Reviews. 6 (2): 94–108. S2CID 35607578.
- Lambert JC, Amouyel P (August 2007). "Genetic heterogeneity of Alzheimer's disease: complexity and advances". Psychoneuroendocrinology. 32 (Suppl 1): S62–S70. S2CID 8114580.
- Liu CC, Liu CC, Kanekiyo T, Xu H, Bu G (February 2013). "Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy". Nature Reviews. Neurology. 9 (2): 106–118. PMID 23296339.
- Mahley RW, Ji ZS (January 1999). "Remnant lipoprotein metabolism: key pathways involving cell-surface heparan sulfate proteoglycans and apolipoprotein E". Journal of Lipid Research. 40 (1): 1–16. PMID 9869645.
- Mahley RW, Rall SC (2002). "Apolipoprotein E: far more than a lipid transport protein". Annual Review of Genomics and Human Genetics. 1 (1): 507–537. PMID 11701639.
- Mahley RW (April 1988). "Apolipoprotein E: cholesterol transport protein with expanding role in cell biology". Science. 240 (4852): 622–630. PMID 3283935.
- Masterman T, Hillert J (June 2004). "The telltale scan: APOE epsilon4 in multiple sclerosis". The Lancet. Neurology. 3 (6): 331. S2CID 54404547.
- Moriyama K, Sasaki J, Matsunaga A, Arakawa F, Takada Y, Araki K, et al. (September 1992). "Apolipoprotein E1 Lys-146----Glu with type III hyperlipoproteinemia". Biochimica et Biophysica Acta. 1128 (1): 58–64. PMID 1356443.
- Parasuraman R, Greenwood PM, Sunderland T (April 2002). "The apolipoprotein E gene, attention, and brain function". Neuropsychology. 16 (2): 254–274. PMID 11949718.
- Raber J (2007). "Role of apolipoprotein E in anxiety". Neural Plasticity. 2007: 91236. PMID 17710250.
- Roses AD, Einstein G, Gilbert J, Goedert M, Han SH, Huang D, et al. (January 1996). "Morphological, biochemical, and genetic support for an apolipoprotein E effect on microtubular metabolism". Annals of the New York Academy of Sciences. 777 (1): 146–157. S2CID 9145181.
- Strittmatter WJ, Roses AD (May 1995). "Apolipoprotein E and Alzheimer disease". Proceedings of the National Academy of Sciences of the United States of America. 92 (11): 4725–4727. PMID 7761390.
- Utermann G, Pruin N, Steinmetz A (January 1979). "Polymorphism of apolipoprotein E. III. Effect of a single polymorphic gene locus on plasma lipid levels in man". Clinical Genetics. 15 (1): 63–72. S2CID 34127430.
- Ye J (August 2007). "Reliance of host cholesterol metabolic pathways for the life cycle of hepatitis C virus". PLOS Pathogens. 3 (8): e108. PMID 17784784.
External links
- Apolipoproteins+E at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- apoe4.info – website for APOE-epsilon-4 carriers
- Human APOE genome location and APOE gene details page in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: P02649 (Apolipoprotein E) at the PDBe-KB.