Phi X 174
| Escherichia virus ΦX174 | |
|---|---|
| |
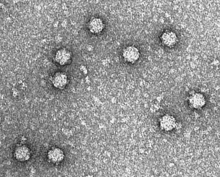
| Electron micrograph of phage ΦX174 | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Monodnaviria |
| Kingdom: | Sangervirae |
| Phylum: | Phixviricota |
| Class: | Malgrandaviricetes |
| Order: | Petitvirales |
| Family: | Microviridae |
| Genus: | Sinsheimervirus |
| Species: | Escherichia virus ΦX174
|

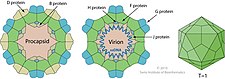
The phi X 174 (or ΦX174)
Genome

This bacteriophage has a [+] sense circular single-stranded DNA genome of 5,386 nucleotides.[10] The genome GC-content is 44% and 95% of nucleotides belong to coding genes. Because of the balance base pattern of the genome, it is used as the control DNA for Illumina sequencers.[citation needed]
Genes
ΦX174 encodes 11 genes, named as consecutive letters of the alphabet in the order they were discovered, with the exception of A* which is an alternative start codon within the large A genes. Only genes A* and K are thought to be non-essential, although there is some doubt about A* because its start codon could be changed to ATT but not any other sequence.[11] It is now known that the ATT is still likely capable of producing protein[12] within E. coli and therefore this gene may in fact be essential.
The first half of the ΦX174 genome features high levels of gene overlap[13] with eight out of 11 genes overlapping by at least one nucleotide.[2] These overlaps have been shown to be non-essential [9] although the refactored phage with all gene overlaps removed had decreased fitness from wild-type.[14]
Phage ΦX174 has been used to try to establish the absence of undiscovered genetic information through a "proof by synthesis" approach.[15]
Transcriptome
In 2020, the transcriptome of ΦX174 was generated.[16] Notable features of the ΦX174 transcriptome is a series of up to four relatively weak promoters in series with up to four Rho-independent (intrinsic) terminators and one Rho-dependent terminator.[citation needed]
Proteins
ΦX174 encodes 11 proteins.
| Protein | Copies | Function[17] |
|---|---|---|
| A | — | Nicks RF DNA to initiate rolling circle replication; ligates ends of linear phage DNA to form single-stranded circular DNA |
| A* | — | Inhibits host cell DNA replication; blocks superinfecting phage; not essential |
| B | 60 in procapsid |
Internal scaffolding protein involved in procapsid assembly |
| C | — | DNA packaging |
| D | 240 in procapsid | External scaffolding protein involved in procapsid assembly |
| E | — | Host cell lysis |
| F | 60 in virion | Major capsid protein |
| G | 60 in virion | Major spike protein |
| H | 12 in virion | DNA pilot protein (or minor spike protein) |
| J | 60 in virion | Binds to new single-stranded phage DNA; accompanies phage DNA into procapsid |
| K | — | Optimizes burst size; not essential |
Proteome
Identification of all ΦX174 proteins using mass spectrometry has recently been reported.[14]
Infection Cycle
Infection begins when G protein binds to
The DNA is ejected through a hydrophilic channel at the 5-fold vertex.
As D protein is the most abundant gene transcript, it is the most protein in the viral procapsid. Similarly, gene transcripts for F, J, and G are more abundant than for H as the stoichiometry for these structural proteins is 5:5:5:1. The primosomes are protein complexes which attach/bind the enzyme helicase on the template. Primosomes gives RNA primers for DNA synthesis to strands.[citation needed]
Phylogenetics and diversity
ΦX174 is closely related to other microviridae, especially the NC phage (e.g. NC1, NC7, NC11, NC16, NC37, NC5, NC41, NC56, NC51, etc.) and more distantly related to the G4-like phages and even more distantly related to the α3-like phage. Rokyta et al. 2006 presented a phylogenetic tree of their relationships.[23]
Uses
Experimental evolution
ΦX174 has been used as a model organism in many evolution experiments.[24]
Biotechnology
ΦX174 is regularly used as a
ΦX174 is also used to test the resistance of personal protective equipment to bloodborne viruses.[26]
ΦX174 has also been modified to enable peptide display (phage display) from the viral capsid G protein.[27]
Synthetic Biology
The ΦX174 genome was the first phage to be cloned in yeast,[9] which provides a convenient drydock for genome modifications.[28] ΦX174 was also the first genome to be fully decompressed, having all gene overlaps removed.[13] The effect of these changes resulted in significantly reduced host attachment, protein expression dysregulation, and heat sensitivity.[14]
See also
References
- PMID 33177747.
- ^ S2CID 4206886.
- PMID 13945085.
- ^ National Library of Medicine Profiles in Science. The Arthur Kornberg Papers. "Creating Life in the Test Tube," 1959-1970. link[non-primary source needed]
- PMID 4873588.
- PMID 4610569.
- PMID 14657399.
- PMID 21840317.
- ^ PMID 23079106.
- ^ a b c Enterobacteria phage phiX174 sensu lato, complete genome. "Complete genome: accession NC_001422", National Center for Biotechnology Information. Retrieved on 30 January 2016.
- S2CID 24174007.
- PMID 28334756.
- ^ PMID 34611352.
- ^ S2CID 222300240.
- PMID 31719208.
- S2CID 219459208.
- ISBN 978-0195148503.
- PMID 1094681.
- PMID 11590105.
- PMID 21227478.
- PMID 19640994.
- PMID 1370343.
- PMID 16428417.
- PMID 20643739.
- ^ "Using a PhiX Control for HiSeq® Sequencing Runs". Illumina. Archived from the original on 9 January 2019. Retrieved 8 January 2019.
- ^ "PPE-Info – Standard Details". wwwn.cdc.gov. Retrieved 8 February 2019.
- PMID 26655242.
- PMID 26973885.
External links
- Goodsell D (February 2000). "Bacteriophage phiX174". Molecule of the Month. RCSB-PDB.
