Molecular biology
| Part of a series on |
| Biology |
|---|
Molecular biology /məˈlɛkjʊlər/ is a branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions.[1][2][3]
Molecular biology was first described as an approach focused on the underpinnings of biological phenomena—uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology.[4]
The term molecular biology was first used in 1945 by physicist
The field of molecular biology includes techniques which enable scientists to learn about molecular processes.
History of molecular biology


Molecular biology sits at the intersection of biochemistry and genetics; as these scientific disciplines emerged and evolved in the 20th century, it became clear that they both sought to determine the molecular mechanisms which underlie vital cellular functions.[9] Advances in molecular biology have been closely related to the development of new technologies and their optimization.[10] Molecular biology has been elucidated by the work of many scientists, and thus the history of the field depends on an understanding of these scientists and their experiments.[citation needed]
The field of genetics arose as an attempt to understand the molecular mechanisms of
A major milestone in molecular biology was the discovery of the structure of DNA. This work began in 1869 by Friedrich Miescher, a Swiss biochemist who first proposed a structure called nuclein, which we now know to be (deoxyribonucleic acid), or DNA.[14] He discovered this unique substance by studying the components of pus-filled bandages, and noting the unique properties of the "phosphorus-containing substances".[15] Another notable contributor to the DNA model was Phoebus Levene, who proposed the "polynucleotide model" of DNA in 1919 as a result of his biochemical experiments on yeast.[16] In 1950, Erwin Chargaff expanded on the work of Levene and elucidated a few critical properties of nucleic acids: first, the sequence of nucleic acids varies across species.[17] Second, the total concentration of purines (adenine and guanine) is always equal to the total concentration of pyrimidines (cysteine and thymine).[14] This is now known as Chargaff's rule. In 1953, James Watson and Francis Crick published the double helical structure of DNA,[18] based on the X-ray crystallography work done by Rosalind Franklin which was conveyed to them by Maurice Wilkins and Max Perutz.[5] Watson and Crick described the structure of DNA and conjectured about the implications of this unique structure for possible mechanisms of DNA replication.[18] Watson and Crick were awarded the Nobel Prize in Physiology or Medicine in 1962, along with Wilkins, for proposing a model of the structure of DNA.[6]
In 1961, it was demonstrated that when a
Griffith's experiment

In 1928,
Griffith's experiment addressed the pneumococcus bacteria, which had two different strains, one virulent and smooth and one avirulent and rough. The smooth strain had glistering appearance owing to the presence of a type of specific polysaccharide – a polymer of glucose and glucuronic acid capsule. Due to this polysaccharide layer of bacteria, a host's immune system cannot recognize the bacteria and it kills the host. The other, avirulent, rough strain lacks this polysaccharide capsule and has a dull, rough appearance.[citation needed]
Presence or absence of capsule in the strain, is known to be genetically determined. Smooth and rough strains occur in several different type such as S-I, S-II, S-III, etc. and R-I, R-II, R-III, etc. respectively. All this subtypes of S and R bacteria differ with each other in antigen type they produce.[6]
Avery–MacLeod–McCarty experiment
The Avery–MacLeod–McCarty experiment was a landmark study conducted in 1944 that demonstrated that DNA, not protein as previously thought, carries genetic information in bacteria.
Hershey–Chase experiment

Confirmation that DNA is the genetic material which is cause of infection came from the Hershey–Chase experiment. They used E.coli and bacteriophage for the experiment. This experiment is also known as blender experiment, as kitchen blender was used as a major piece of apparatus. Alfred Hershey and Martha Chase demonstrated that the DNA injected by a phage particle into a bacterium contains all information required to synthesize progeny phage particles. They used radioactivity to tag the bacteriophage's protein coat with radioactive sulphur and DNA with radioactive phosphorus, into two different test tubes respectively. After mixing bacteriophage and E.coli into the test tube, the incubation period starts in which phage transforms the genetic material in the E.coli cells. Then the mixture is blended or agitated, which separates the phage from E.coli cells. The whole mixture is centrifuged and the pellet which contains E.coli cells was checked and the supernatant was discarded. The E.coli cells showed radioactive phosphorus, which indicated that the transformed material was DNA not the protein coat.
The transformed DNA gets attached to the DNA of E.coli and radioactivity is only seen onto the bacteriophage's DNA. This mutated DNA can be passed to the next generation and the theory of Transduction came into existence. Transduction is a process in which the bacterial DNA carry the fragment of bacteriophages and pass it on the next generation. This is also a type of horizontal gene transfer.[6]
Meselson–Stahl experiment

The Meselson-Stahl experiment was a landmark experiment in molecular biology that provided evidence for the semiconservative replication of DNA. Conducted in 1958 by Matthew Meselson and Franklin Stahl, the experiment involved growing E. coli bacteria in a medium containing heavy isotope of nitrogen (15N) for several generations. This caused all the newly synthesized bacterial DNA to be incorporated with the heavy isotope.
After allowing the bacteria to replicate in a medium containing normal nitrogen (14N), samples were taken at various time points. These samples were then subjected to centrifugation in a density gradient, which separated the DNA molecules based on their density.
The results showed that after one generation of replication in the 14N medium, the DNA formed a band of intermediate density between that of pure 15N DNA and pure 14N DNA. This supported the semiconservative DNA replication proposed by Watson and Crick, where each strand of the parental DNA molecule serves as a template for the synthesis of a new complementary strand, resulting in two daughter DNA molecules, each consisting of one parental and one newly synthesized strand.
The Meselson-Stahl experiment provided compelling evidence for the semiconservative replication of DNA, which is fundamental to the understanding of genetics and molecular biology.
Modern molecular biology
In the early 2020s, molecular biology entered a golden age defined by both vertical and horizontal technical development. Vertically, novel technologies are allowing for real-time monitoring of biological processes at the atomic level.[22] Molecular biologists today have access to increasingly affordable sequencing data at increasingly higher depths, facilitating the development of novel genetic manipulation methods in new non-model organisms. Likewise, synthetic molecular biologists will drive the industrial production of small and macro molecules through the introduction of exogenous metabolic pathways in various prokaryotic and eukaryotic cell lines.[23]
Horizontally, sequencing data is becoming more affordable and used in many different scientific fields. This will drive the development of industries in developing nations and increase accessibility to individual researchers. Likewise, CRISPR-Cas9 gene editing experiments can now be conceived and implemented by individuals for under $10,000 in novel organisms, which will drive the development of industrial and medical applications.[24]
Relationship to other biological sciences

The following list describes a viewpoint on the interdisciplinary relationships between molecular biology and other related fields.[25]
- Molecular biology is the study of the molecular underpinnings of the biological phenomena, focusing on molecular synthesis, modification, mechanisms and interactions.
- Biochemistry is the study of the chemical substances and vital processes occurring in living organisms. Biochemists focus heavily on the role, function, and structure of biomolecules such as proteins, lipids, carbohydrates and nucleic acids.[26]
- Genetics is the study of how genetic differences affect organisms. Genetics attempts to predict how mutations, individual genes and genetic interactions can affect the expression of a phenotype[27]
While researchers practice techniques specific to molecular biology, it is common to combine these with methods from genetics and biochemistry. Much of molecular biology is quantitative, and recently a significant amount of work has been done using computer science techniques such as bioinformatics and computational biology. Molecular genetics, the study of gene structure and function, has been among the most prominent sub-fields of molecular biology since the early 2000s. Other branches of biology are informed by molecular biology, by either directly studying the interactions of molecules in their own right such as in cell biology and developmental biology, or indirectly, where molecular techniques are used to infer historical attributes of populations or species, as in fields in evolutionary biology such as population genetics and phylogenetics. There is also a long tradition of studying biomolecules "from the ground up", or molecularly, in biophysics.[28]
Techniques of molecular biology

Molecular cloning

Molecular cloning is used to isolate and then transfer a DNA sequence of interest into a plasmid vector.
This plasmid can be inserted into either bacterial or animal cells. Introducing DNA into bacterial cells can be done by
DNA coding for a protein of interest is now inside a cell, and the
Polymerase chain reaction
Polymerase chain reaction (PCR) is an extremely versatile technique for copying DNA. In brief, PCR allows a specific

Gel electrophoresis

Gel electrophoresis is a technique which separates molecules by their size using an agarose or polyacrylamide gel.[37] This technique is one of the principal tools of molecular biology. The basic principle is that DNA fragments can be separated by applying an electric current across the gel - because the DNA backbone contains negatively charged phosphate groups, the DNA will migrate through the agarose gel towards the positive end of the current.[37] Proteins can also be separated on the basis of size using an SDS-PAGE gel, or on the basis of size and their electric charge by using what is known as a 2D gel electrophoresis.[38]

The Bradford protein assay
The
This method was developed in 1975 by Marion M. Bradford, and has enabled significantly faster, more accurate protein quantitation compared to previous methods: the Lowry procedure and the biuret assay.[39] Unlike the previous methods, the Bradford assay is not susceptible to interference by several non-protein molecules, including ethanol, sodium chloride, and magnesium chloride.[39] However, it is susceptible to influence by strong alkaline buffering agents, such as sodium dodecyl sulfate (SDS).[39]
Macromolecule blotting and probing
The terms northern, western and eastern blotting are derived from what initially was a molecular biology joke that played on the term Southern blotting, after the technique described by Edwin Southern for the hybridisation of blotted DNA. Patricia Thomas, developer of the RNA blot which then became known as the northern blot, actually did not use the term.[41]
Southern blotting
Named after its inventor, biologist
Northern blotting

The northern blot is used to study the presence of specific RNA molecules as relative comparison among a set of different samples of RNA. It is essentially a combination of
Western blotting
A western blot is a technique by which specific proteins can be detected from a mixture of proteins.
Eastern blotting
The eastern blotting technique is used to detect post-translational modification of proteins. Proteins blotted on to the PVDF or nitrocellulose membrane are probed for modifications using specific substrates.[48]
Microarrays
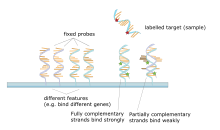
A DNA microarray is a collection of spots attached to a solid support such as a
Allele-specific oligonucleotide
Allele-specific oligonucleotide (ASO) is a technique that allows detection of single base mutations without the need for PCR or gel electrophoresis. Short (20–25 nucleotides in length), labeled probes are exposed to the non-fragmented target DNA, hybridization occurs with high specificity due to the short length of the probes and even a single base change will hinder hybridization. The target DNA is then washed and the labeled probes that did not hybridize are removed. The target DNA is then analyzed for the presence of the probe via radioactivity or fluorescence. In this experiment, as in most molecular biology techniques, a control must be used to ensure successful experimentation.[53][54]
In molecular biology, procedures and technologies are continually being developed and older technologies abandoned. For example, before the advent of DNA
See also
- Astrobiology
- Central dogma of molecular biology
- Genetic code
- Geniom RT Analyzer, diagnostic testing instrument
- Genome
- Molecular biology institutes
- Molecular engineering
- Molecular modeling
- Protein interaction prediction
- Protein structure prediction
- Proteome
- Cell biology
References
- ISBN 978-1-317-56375-4.
- PMID 11839687.
- ^ "Molecular biology – Latest research and news | Nature". nature.com. Retrieved 2021-11-07.
- S2CID 4172248.
- ^ a b "Rosalind Franklin: A Crucial Contribution". nature.com.
- ^ OCLC 1045495545.
- ISBN 9780470016176, retrieved 2021-11-07
- S2CID 29581630.
- OCLC 1184123419.
- PMID 11517346.
- PMID 21775188.
- ^ "12.3C: Mendel's Law of Segregation". Biology LibreTexts. 2018-07-12. Retrieved 2021-11-18.
- ^ "Mendelian Inheritance". Genome.gov. Retrieved 2021-11-18.
- ^ a b "Discovery of DNA Double Helix: Watson and Crick | Learn Science at Scitable". www.nature.com. Retrieved 2021-11-25.
- OCLC 907773747.
- ISSN 0021-9258.
- S2CID 2522535.
- ^ S2CID 4253007.
- S2CID 4276146.
- ISSN 0091-7451.
- PMID 15514035.
- PMID 34221652.
- ^ van Warmerdam, T. "Molecular Biology Laboratory Resource". Yourbiohelper.com.
- ^ van Warmerdam, T. "Molecular biology laboratory resource". Yourbiohelper.com.
- ISBN 978-0-7167-3136-8.
- OCLC 48055706.
- ^ Reference, Genetics Home. "Help Me Understand Genetics". Genetics Home Reference. Retrieved 31 December 2016.
- ^ ISBN 9783642343032. Retrieved 2019-07-08.
- ^ "Foundations of Molecular Cloning - Past, Present and Future | NEB". www.neb.com. Retrieved 2021-11-25.
- ^ "Foundations of Molecular Cloning - Past, Present and Future | NEB". www.neb.com. Retrieved 2021-11-04.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Isolating, Cloning, and Sequencing DNA. Retrieved 31 December 2016.
- PMID 24011038.
- ISBN 9788131239872. Retrieved 2019-07-08.
- PMID 7580841.
- ^ "Polymerase Chain Reaction (PCR)". National Center for Biotechnology Information. U.S. National Library of Medicine. Retrieved 31 December 2016.
- ^ "Polymerase Chain Reaction (PCR) Fact Sheet". National Human Genome Research Institute (NHGRI). Retrieved 31 December 2016.
- ^ PMID 22546956.
- PMID 22546956.
- ^ S2CID 4359292.
- ^ a b c "Protein determination by the Bradford method". www.ruf.rice.edu. Retrieved 2021-11-08.
- PMID 6159641.
- S2CID 20686993.
- PMID 21125485.
- PMID 24034315.
- ^ PMID 23050259.
- ^ "Western blot | Learn Science at Scitable". www.nature.com. Retrieved 2021-11-25.
- PMID 16483794. – via ScienceDirect(Subscription may be required or content may be available in libraries.)
- PMID 19493209.
- ^ "Microarrays". National Center for Biotechnology Information. U.S. National Library of Medicine. Retrieved 31 December 2016.
- PMID 23288464.
- PMID 23066278.
- PMID 16890548.
- ISBN 978-1-59745-405-6. Retrieved 31 December 2016.
- ISBN 978-3-319-19674-9. Retrieved 31 December 2016.
- ISBN 9783642343032. Retrieved 2019-07-08.
Further reading
- Cohen SN, Chang AC, Boyer HW, Helling RB (November 1973). "Construction of biologically functional bacterial plasmids in vitro". Proceedings of the National Academy of Sciences of the United States of America. 70 (11): 3240–4. PMID 4594039.
- Rodgers M (June 1975). "The Pandora's box congress". Rolling Stone. Vol. 189. pp. 37–77.
- Roberts K, Raff M, Alberts B, Walter P, Lewis J, Johnson A (2002). Molecular Biology of the Cell. Garland Science. ISBN 978-0-8153-3218-3.
External links
 Media related to Molecular biology at Wikimedia Commons
Media related to Molecular biology at Wikimedia Commons- Biochemistry and Molecular Biology at Curlie

