COVID Moonshot
This article needs to be updated. (April 2022) |
| External videos | |
|---|---|
Oxford University, November 17, 2020 | |
COVID Moonshot is a collaborative
COVID Moonshot researchers are targeting theCOVID Moonshot may be the first open-science community effort for the development of an antiviral drug.[2] Hundreds of scientists around the world, from academic and industrial organizations, have shared their expertise, resources, data, and results to more rapidly identify, screen, and test candidate compounds for the treatment of COVID-19.[4]
Project history
Development of antiviral drugs is a complicated and time-consuming multistage process.[5] The public sharing of information in the early stages of genome identification and protein structure identification has accelerated the process of searching for COVID-19 treatments and established a basis for the COVID Moonshot initiative.[6][7]
Genome identification
On January 3, 2020, Chinese virologist
Protein structures
With that information, structural biologists world-wide began examining its protein structures. Investigators from the Center for Structural Genomics of Infectious Diseases (CSGID) and other groups began working to characterize the 3D structure of the proteins, sharing their results via the Protein Data Bank (PDB).[6][7][11]

Scientists were able to identify a key protein in the virus: 3C-like protease (Mpro).[6][7] Crucial early
Approaches to accelerating drug development have been suggested, but identification of proteins and drug development commonly take years.
Possible targets
Identifying and recreating viral proteins in the lab is a first step to developing drugs to attack them and vaccines to protect against them.[6] The COVID Moonshot initiative follows an approach to structure-based drug design in which researchers attempt to find a molecule that will bind tightly to a drug target and prevent it from carrying out its normal activities.[7][2]
In the case of SARS-CoV-2, the coronavirus enters the body and then replicates its genomic
Fragment screening
In collaboration with the
Researchers examined thousands of possible fragments from diverse screening libraries and identified at least 71 possible protein–ligand crystal structures, chemical fragments which might have the potential to bind to Mpro.[19][15] These results were immediately made available online.[4][15]
Designing candidates
The open release of the data and its announcement on Twitter on March 7, 2020, mark a critical point in the formation of COVID Moonshot. The scientists shared their information and challenged chemists worldwide to use that information to design potential openly available antiviral drug candidates.[7][6][9] They expected a couple of hundred submissions. By May 2020 more than 4,600 design submissions for potential inhibitors were received.[6] By January 2021, the number of unique compound designs had risen to 14,000.[7] In response, those involved began to shift from a spontaneous virtual collaboration to a larger and more organized network of partners with specialized skills and well-articulated goals.[20]

The design submissions were stored in Collaborative Drug Discovery's CDD Vault, a database used for large-scale management of chemical structures, experimental protocols and experimental results.[4] Alpha Lee and Matt Robinson brought computational expertise from PostEra to the project. PostEra used techniques from artificial intelligence and machine learning to develop analysis tools for computational drug discovery, chemical synthesis and biochemical assays. When COVID Moonshot's appeal resulted in not hundreds but thousands of responses, they built a platform capable of triaging large numbers of compounds and designing routes for their synthetic formation.[4]
Supercomputer access was provided through the COVID-19 High Performance Computing (HPC) Consortium, accelerating the speed at which designs could be examined and compared.[21][22] The distributed supercomputing initiative Folding@home has carried out multiple sprints to model novel protein structures and target desirable structures as a part of COVID Moonshot.[23][24][25]
Many of the criteria for selecting drug candidates were determined by the group's goals. An ideal drug candidate would be effective in treating COVID-19. It also would be easily and cheaply made, so that as many countries and companies as possible could produce and distribute it. The ingredients to make it should be easy to obtain, and the processes involved should be as simple as possible. A drug shouldn't require special handling (like refrigeration) and it should be easy to administer (a pill rather than an injection).[4][20]
In a matter of months, researchers were able to identify more than 200 promising crystal structure designs and to begin creating and testing them in the lab.[26][27] Chris Schofield at the University of Oxford synthesized and tested 4 of the most promising of the novel designed peptides to demonstrate their ability to block and inhibit Mpro.[27] Freely available data from COVID Moonshot has also been used to assess the predictive ability of docking scores in suggesting the potency of SARS-CoV-2 M-pro inhibitors.[28]
To go beyond the design phase, possible drug candidates must be created and tested for both effectiveness and safety in animal and human trials.
Potential for antiviral treatments
COVID Moonshot anticipates that they will select three pre-clinical candidates by March 2022,[needs update] to be followed by preclinical safety and toxicology testing and identification of needed chemistry, manufacturing and control (CMC) steps. Based on that data, the most promising candidate will be chosen. Phase-1 clinical trials, the first stage of testing in human subjects, are projected to begin by June 2023.[31][20]
Unlike a
Mpro is present in other
Mpro does not mutate easily, so it is less likely that variants of the virus will adapt that can avoid the effects of such a drug.[2]
Open science
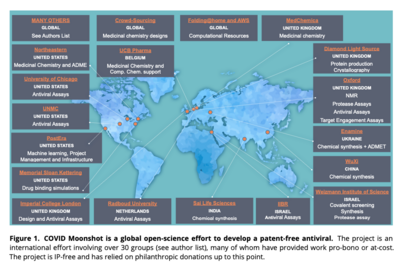
Among the many participants in the COVID Moonshot project are the University of Oxford, University of Cambridge, Diamond Light Source,
Temple University,[4] Memorial Sloan Kettering Cancer Center, PostEra, University of Johannesburg, and theBecause COVID Moonshot is based in open science and shared open data, any drug that the project develops can be manufactured and sold by whoever wishes to produce it, worldwide. Countries that are unable to buy or manufacture expensive licensed drugs would therefore have the opportunity to produce their own supplies, and competition between suppliers is likely to result in greater availability and reduced prices for consumers.[4]
This would circumvent issues around the time needed to vaccinate people worldwide. As of July 2021, it was estimated that at current rates, this was likely to take several years. Inequities in distribution will increase both the spreading of the virus and the risk that new and more dangerous variants will emerge.[35][36]
Supporters of the COVID Moonshot initiative have argued that open-science drug discovery is an essential model for combating both current and future pandemics, and that the prevention of the spread of pandemic diseases is an essential public service.[4]
References
- ^ Whipple, Tom (October 23, 2021). "Moonshot is the spanner in the Covid-19 works the country needs". The Times. Retrieved 5 November 2021.
- ^ S2CID 244170138. Retrieved 1 November 2021.
- S2CID 233922849. Retrieved 1 November 2021.
- ^ a b c d e f g h i j k Thomas, Uduak Grace (12 August 2020). "The COVID-19 antiviral race: we're all in this together". Drug Discovery World. Vol. 21, no. 3. Retrieved 1 November 2021.
- ^ PMID 27483339. Retrieved 2 November 2021.
- ^ a b c d e f g h i j Scudellari, Megan (19 May 2020). "The sprint to solve coronavirus protein structures — and disarm them with drugs". Nature. Retrieved 2 November 2021.
- ^ PMID 33248689.
- ^ a b Cyranoski, David (14 December 2020). "Nature's 10: ten people who helped shape science in 2020 : Zhang Yongzhen: Genome sharer". Nature. Retrieved 2 November 2021.
- ^ a b Howes, Laura (May 2, 2020). "How structural biologists revealed the new coronavirus's structure so quickly". Chemical & Engineering News. 98 (17). Retrieved 5 November 2021.
- ^ Holmes, Edward (11 January 2020). "Novel 2019 coronavirus genome". virological.org. Retrieved 2 November 2021.
- ^ "Structural Genomics Centers for Infectious Diseases". National Institute of Allergy and Infectious Diseases (NIAID). 7 May 2021. Retrieved 2 November 2021.
- ^ PMID 33889562.
- ^ Ahuja, Anjana (7 April 2020). "How to marshal a crowd to launch a moonshot against Covid-19". Financial Times. Retrieved 5 November 2021.
- S2CID 230551125.
- ^ PMID 33028810. Retrieved 3 November 2021.
- PMID 22222452. Retrieved 3 November 2021.
- PMID 28986384. Retrieved 3 November 2021.
- PMID 33856612.
- ^ S2CID 219906197.
- ^ a b c d e f "COVID Moonshot funded by COVID-19 Therapeutics Accelerator to rapidly develop a safe, globally accessible and affordable antiviral pill". Drugs for Neglected Diseases Initiative (DNDi). 27 September 2021. Retrieved 3 November 2021.
- ^ Wiggers, Kyle (28 May 2020). "COVID-19 HPC Consortium pours 437 petaflops of compute power toward virus research". VentureBeat. Retrieved 5 November 2021.
- ^ Morris, Aaron. "PostEra: COVID MoonShot". COVID-19 HPC Consortium. Retrieved 5 November 2021.
- ^ Masterson, Victoria (22 December 2020). "Your computer can help scientists find a cure for COVID-19. Here's how". World Economic Forum. Retrieved 5 November 2021.
- ^ "Together We Are Powerful". FOLDINGATHOME. Retrieved 5 November 2021.
- ^ S2CID 235438395.
- ^ PMID 34760153.
- S2CID 239888873. Retrieved 5 November 2021.
- ^ Scudellari, Megan (4 December 2020). "COVID Moonshot Effort Generates "Elite" Antivirals". IEEE Spectrum. Retrieved 5 November 2021.
- ^ "Moonshot initiative to develop affordable COVID-19 antivirals gets funding boost". University of Oxford. 28 September 2021. Retrieved 5 November 2021.
- ^ von Delft, Annette; Mowbray, Charles; Nyaoke, Borna (24 December 2021). "The Moonshot: Crowdsourcing To Develop The First Open-Source, Generic COVID-19 Antiviral Pill - Health Policy Watch". Health Policy Watch. Retrieved 17 January 2022.
- ^ "Israeli scientists to participate in int'l drive to find COVID-curing pills". The Jerusalem Post. September 29, 2021. Retrieved 5 November 2021.
- ^ "COVID Moonshot consortium receives funding from Wellcome". Diamond Light. 2020. Retrieved 5 November 2021.
- ^ "PostEra and LifeArc collaborate on novel open science initiative to develop new antiviral for COVID-19". LifeArc. 20 May 2021. Retrieved 5 November 2021.
- ^ Safi, Michael (27 January 2021). "Most poor nations 'will take until 2024 to achieve mass Covid-19 immunisation'". The Guardian. Retrieved 5 November 2021.
- S2CID 235744340.
