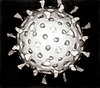
Capsid


A capsid is the protein shell of a
Capsids are broadly classified according to their structure. The majority of the viruses have capsids with either
Some viruses are enveloped, meaning that the capsid is coated with a lipid membrane known as the viral envelope. The envelope is acquired by the capsid from an intracellular membrane in the virus' host; examples include the inner nuclear membrane, the Golgi membrane, and the cell's outer membrane.[7]
Once the virus has infected a cell and begins replicating itself, new capsid subunits are synthesized using the protein biosynthesis mechanism of the cell. In some viruses, including those with helical capsids and especially those with RNA genomes, the capsid proteins co-assemble with their genomes. In other viruses, especially more complex viruses with double-stranded DNA genomes, the capsid proteins assemble into empty precursor procapsids that include a specialized portal structure at one vertex. Through this portal, viral DNA is translocated into the capsid.[8]
Structural analyses of major capsid protein (MCP) architectures have been used to categorise viruses into lineages. For example, the bacteriophage PRD1, the algal virus Paramecium bursaria Chlorella virus-1 (PBCV-1), mimivirus and the mammalian adenovirus have been placed in the same lineage, whereas tailed, double-stranded DNA bacteriophages (Caudovirales) and herpesvirus belong to a second lineage.[9][10][11][12]
Specific shapes
Icosahedral
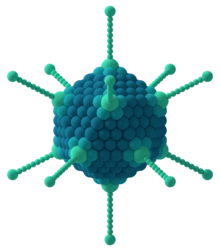


The icosahedral structure is extremely common among viruses. The
In this scheme, icosahedral capsids contain 12 pentamers plus 10(T − 1) hexamers.[14][15] The T-number is representative of the size and complexity of the capsids.[16] Geometric examples for many values of h, k, and T can be found at List of geodesic polyhedra and Goldberg polyhedra.
Many exceptions to this rule exist: For example, the
T-numbers can be represented in different ways, for example T = 1 can only be represented as an icosahedron or a dodecahedron and, depending on the type of quasi-symmetry, T = 3 can be presented as a truncated dodecahedron, an icosidodecahedron, or a truncated icosahedron and their respective duals a triakis icosahedron, a rhombic triacontahedron, or a pentakis dodecahedron.[18][clarification needed]
Prolate
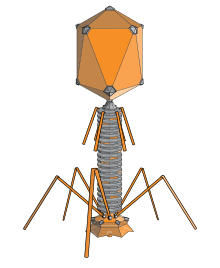
An elongated icosahedron is a common shape for the heads of bacteriophages. Such a structure is composed of a cylinder with a cap at either end. The cylinder is composed of 10 elongated triangular faces. The Q number (or Tmid), which can be any positive integer,[19] specifies the number of triangles, composed of asymmetric subunits, that make up the 10 triangles of the cylinder. The caps are classified by the T (or Tend) number.[20]
The bacterium E. coli is the host for bacteriophage T4 that has a prolate head structure. The bacteriophage encoded gp31 protein appears to be functionally homologous to E. coli chaperone protein GroES and able to substitute for it in the assembly of bacteriophage T4 virions during infection.[21] Like GroES, gp31 forms a stable complex with GroEL chaperonin that is absolutely necessary for the folding and assembly in vivo of the bacteriophage T4 major capsid protein gp23.[21]
Helical

Many rod-shaped and filamentous plant viruses have capsids with helical symmetry.[22] The helical structure can be described as a set of n 1-D molecular helices related by an n-fold axial symmetry.[23] The helical transformation are classified into two categories: one-dimensional and two-dimensional helical systems.[23] Creating an entire helical structure relies on a set of translational and rotational matrices which are coded in the protein data bank.[23] Helical symmetry is given by the formula P = μ x ρ, where μ is the number of structural units per turn of the helix, ρ is the axial rise per unit and P is the pitch of the helix. The structure is said to be open due to the characteristic that any volume can be enclosed by varying the length of the helix.[24] The most understood helical virus is the tobacco mosaic virus.[22] The virus is a single molecule of (+) strand RNA. Each coat protein on the interior of the helix bind three nucleotides of the RNA genome. Influenza A viruses differ by comprising multiple ribonucleoproteins, the viral NP protein organizes the RNA into a helical structure. The size is also different; the tobacco mosaic virus has a 16.33 protein subunits per helical turn,[22] while the influenza A virus has a 28 amino acid tail loop.[25]
Functions
The functions of the capsid are to:
- protect the genome,
- deliver the genome, and
- interact with the host.
The virus must assemble a stable, protective protein shell to protect the genome from lethal chemical and physical agents. These include extremes of pH or temperature and proteolytic and nucleolytic enzymes. For non-enveloped viruses, the capsid itself may be involved in interaction with receptors on the host cell, leading to penetration of the host cell membrane and internalization of the capsid. Delivery of the genome occurs by subsequent uncoating or disassembly of the capsid and release of the genome into the cytoplasm, or by ejection of the genome through a specialized portal structure directly into the host cell nucleus.
Origin and evolution
It has been suggested that many viral capsid proteins have evolved on multiple occasions from functionally diverse cellular proteins.
A computational model (2015) has shown that capsids may have originated before viruses and that they served as a means of
See also
References
- S2CID 16706951.
- S2CID 6023873.
- PMID 18003933.
- PMID 21368184.
- ISBN 978-0-8153-0270-4.
- ^ "Virus Structure (web-books.com)". Archived from the original on 2021-02-07. Retrieved 2007-07-10.
- Watson JD (1994). Molecular Biology of the Cell (4th ed.). p. 280.
- PMID 16051846.
- S2CID 31542714.
- PMID 16505372.
- PMID 16357204.
- PMID 16338410.
- PMID 14019094.
- PMID 18981051. Archived from the originalon 2018-02-11. Retrieved 2011-03-18.
- ISBN 978-0-12-375146-1.
- PMID 20209096.
- ^ Sgro JY. "Virusworld". Institute for Molecular Virology. University of Wisconsin-Madison.
- PMID 12460573.
- PMID 20550912.
- ISBN 978-0-12-375146-1.
- ^ a b Marusich EI, Kurochkina LP, Mesyanzhinov VV. Chaperones in bacteriophage T4 assembly. Biochemistry (Mosc). 1998;63(4):399-406
- ^ PMID 3629216.
- ^ S2CID 33065417.
- ISBN 978-1-55581-479-3.
- PMID 23269829.
- ^ PMID 28265094.
- ^ S2CID 169035711.
- PMID 25955384.
Further reading
- ISBN 978-0-486-23729-9.
- Pugh A (1 September 1976). Polyhedra: A Visual Approach. University of California Press. Chapter 6. The Geodesic Polyhedra of R. Buckminster Fuller and Related Polyhedra. ISBN 978-0-520-02926-2.
- Almansour I, Alhagri M, Alfares R, Alshehri M, Bakhashwain R, Maarouf A (January 2019). "IRAM: virus capsid database and analysis resource". Database: The Journal of Biological Databases and Curation. 2019. PMID 31318422.
External links
- IRAM-Virus Capsid Database and Analysis Resource Archived 2019-10-23 at the Wayback Machine