Chemical biology
Chemical biology is a scientific discipline between the fields of chemistry and biology. The discipline involves the application of chemical techniques, analysis, and often small molecules produced through synthetic chemistry, to the study and manipulation of biological systems.[1] Although often confused with biochemistry, which studies the chemistry of biomolecules and regulation of biochemical pathways within and between cells, chemical biology remains distinct by focusing on the application of chemical tools to address biological questions.[2]
History
Although considered a relatively new scientific field,[2] the term "chemical biology" has been in use since the early 20th century,[3] and has roots in scientific discovery from the early 19th century. The term 'chemical biology' can be traced back to an early appearance in a book published by Alonzo E. Taylor in 1907 titled "On Fermentation",[4] and was subsequently used in John B. Leathes' 1930 article titled "The Harveian Oration on The Birth of Chemical Biology".[5] However, it is unclear when the term was first used.[3]
Friedrich Wöhler's 1828 synthesis of urea is an early example of the application of synthetic chemistry to advance biology.[6] It showed that biological compounds could be synthesized with inorganic starting materials and weakened the previous notion of vitalism, or that a 'living' source was required to produce organic compounds.[7][8] Wöhler's work is often considered to be instrumental in the development of organic chemistry and natural product synthesis, both of which play a large part in modern chemical biology.[9]
Friedrich Miescher's work during the late 19th century investigating the cellular contents of human leukocytes led to the discovery of 'nuclein', which would later be renamed DNA.[6] After isolating the nuclein from the nucleus of leukocytes through protease digestion, Miescher used chemical techniques such as elemental analysis and solubility tests to determine the composition of nuclein.[10] This work would lay the foundations for Watson and Crick's discovery of the double-helix structure of DNA.[10][11]
The rising interest into chemical biology has led to the creation of multiple journals dedicated to the field. Nature Chemical Biology, created in 2005,[12] and ACS Chemical Biology, created in 2006,[13] are two of the most well-known journals in this field, with impact factors of 14.8[14] and 4.0[15] respectively.
Nobel laureates in chemical biology
| Laureate | Year | Discipline | Contribution |
|---|---|---|---|
| Paul Berg | 1980 | Chemistry | Recombinant DNA[16] |
| Walter Gilbert | 1980 | Chemistry | Genome sequencing[16] |
| Kary Mullis | 1993 | Chemistry | Polymerase chain reaction[17] |
| Michael Smith | 1993 | Chemistry | Site-directed mutagenesis[17] |
| Venkatraman Ramakrishnan | 2009 | Chemistry | Elucidation of ribosome structure and function[18] |
| Robert J. Lefkowitz | 2012 | Chemistry | G-protein-coupled receptors[19] |
| Frances H. Arnold | 2018 | Chemistry | Enzyme development through directed evolution[20] |
| Emmanuelle Charpentier | 2020 | Chemistry | CRISPR/Cas9 genetic scissors[21] |
| Barry Sharpless | 2022 | Chemistry | Click chemistry[22] |
| Carolyn Bertozzi | 2022 | Chemistry | Applications of click chemistry in living organisms[22] |
Research areas
Glycobiology

Combinatorial chemistry
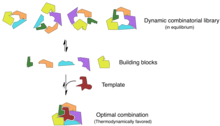
Combinatorial chemistry involves simultaneously synthesizing a large number of related compounds for high-throughput analysis.[27] Chemical biologists are able to use principles from combinatorial chemistry in synthesizing active drug compounds and maximizing screening efficiency.[28] Similarly, these principles can be used in areas of agriculture and food research, specifically in the syntheses of unnatural products and in generating novel enzyme inhibitors.[29]
Peptide synthesis

Chemical synthesis of proteins is a valuable tool in chemical biology as it allows for the introduction of non-natural amino acids as well as residue specific incorporation of "
To make protein-sized polypeptide chains with the small peptide fragments made by synthesis, chemical biologists can use the process of
Enrichment techniques for proteomics
Chemical biologists work to improve
Enzyme probes
To investigate enzymatic activity as opposed to total protein, activity-based reagents have been developed to label the enzymatically active form of proteins (see
Employing biology
Many research programs are also focused on employing natural biomolecules to perform biological tasks or to support a new chemical method. In this regard, chemical biology researchers have shown that DNA can serve as a template for synthetic chemistry, self-assembling proteins can serve as a structural scaffold for new materials, and RNA can be evolved in vitro to produce new catalytic function. Additionally, heterobifunctional (two-sided) synthetic small molecules such as dimerizers or PROTACs bring two proteins together inside cells, which can synthetically induce important new biological functions such as targeted protein degradation.[43]
Directed evolution
A primary goal of
Several methods exist for creating large libraries of sequence variants. Among the most widely used are subjecting
Once useful variants are found, their DNA sequence is amplified and subjected to further rounds of diversification and selection.The development of directed evolution methods was honored in 2018 with the awarding of the Nobel Prize in Chemistry to Frances Arnold for evolution of enzymes, and George Smith and Gregory Winter for phage display.[52]
Bioorthogonal reactions
Successful labeling of a molecule of interest requires specific functionalization of that molecule to react chemospecifically with an optical probe. For a labeling experiment to be considered robust, that functionalization must minimally perturb the system. Unfortunately, these requirements are often hard to meet. Many of the reactions normally available to organic chemists in the laboratory are unavailable in living systems.[53] Water- and redox- sensitive reactions would not proceed, reagents prone to nucleophilic attack would offer no chemospecificity, and any reactions with large kinetic barriers would not find enough energy in the relatively low-heat environment of a living cell.[54] Thus, chemists have recently developed a panel of bioorthogonal chemistry that proceed chemospecifically, despite the milieu of distracting reactive materials in vivo.
The coupling of a probe to a molecule of interest must occur within a reasonably short time frame;
Discovery of biomolecules through metagenomics
The advances in modern sequencing technologies in the late 1990s allowed scientists to investigate DNA of communities of organisms in their natural environments ("eDNA"), without culturing individual species in the lab. This

Functional metagenomic studies enable the discovery of novel genes that encode biologically active molecules. These assays include top agar overlay assays where antibiotics generate zones of growth inhibition against test microbes, and pH assays that can screen for pH change due to newly synthesized molecules using pH indicator on an
Kinases
Through the use of small molecule modulators of protein kinases, chemical biologists have gained a better understanding of the effects of protein phosphorylation. For example, nonselective and selective kinase inhibitors, such as a class of pyridinylimidazole compounds [63] are potent inhibitors useful in the dissection of MAP kinase signaling pathways. These pyridinylimidazole compounds function by targeting the ATP binding pocket. Although this approach, as well as related approaches,[64][65] with slight modifications, has proven effective in a number of cases, these compounds lack adequate specificity for more general applications. Another class of compounds, mechanism-based inhibitors, combines knowledge of the kinase enzymology with previously utilized inhibition motifs. For example, a "bisubstrate analog" inhibits kinase action by binding both the conserved ATP binding pocket and a protein/peptide recognition site on the specific kinase.[66] Research groups also utilized ATP analogs as chemical probes to study kinases and identify their substrates.[67][68][69]
The development of novel chemical means of incorporating
Advances in chemical biology have also improved upon classical techniques of imaging kinase action. For example, the development of peptide biosensors—peptides containing incorporated fluorophores improved temporal resolution of in vitro binding assays.[73] One of the most useful techniques to study kinase action is Fluorescence Resonance Energy Transfer (FRET). To utilize FRET for phosphorylation studies, fluorescent proteins are coupled to both a phosphoamino acid binding domain and a peptide that can by phosphorylated. Upon phosphorylation or dephosphorylation of a substrate peptide, a conformational change occurs that results in a change in fluorescence.[74] FRET has also been used in tandem with Fluorescence Lifetime Imaging Microscopy (FLIM)[75] or fluorescently conjugated antibodies and flow cytometry[76] to provide quantitative results with excellent temporal and spatial resolution.
Biological fluorescence
Chemical biologists often study the functions of biological macromolecules using
Fluorescent techniques have been used assess a number of protein dynamics including protein tracking, conformational changes, protein–protein interactions, protein synthesis and turnover, and enzyme activity, among others. Three general approaches for measuring protein net redistribution and diffusion are single-particle tracking,
Education in chemical biology
Undergraduate education
Despite an increase in biological research within chemistry departments, attempts at integrating chemical biology into undergraduate curricula are lacking.[80] For example, although the American Chemical Society (ACS) requires for foundational courses in a Chemistry Bachelor's degree to include biochemistry, no other biology-related chemistry course is required.[81]
Although a chemical biology course is often not required for an undergraduate degree in Chemistry, many universities now provide introductory chemical biology courses for their undergraduate students. The University of British Columbia, for example, offers a fourth-year course in synthetic chemical biology.[82]
See also
References
- S2CID 14399359.
- ^ ISBN 978-0-470-84530-1.
- ^ PMID 36835407.
- ^ Taylor AE (1907). On fermentation. 8. Vol. 1. University Press.
- PMID 20775787.
- ^ S2CID 8427286.
- S2CID 44613876.
- S2CID 71727190.
- PMID 25043880.
- ^ S2CID 915930.
- PMID 15680349.
- ^ "LC Catalog - Item Information (Full Record)". catalog.loc.gov. Retrieved 6 November 2023.
- ^ "LC Catalog - Item Information (Full Record)". catalog.loc.gov. Retrieved 6 November 2023.
- ^ "Journal Metrics | Nature Chemical Biology". www.nature.com. Retrieved 6 November 2023.
- ^ "About the Journal". ACS Publications. Retrieved 5 November 2023.
- ^ a b "The Nobel Prize in Chemistry 1980". NobelPrize.org. Retrieved 6 November 2023.
- ^ a b "The Nobel Prize in Chemistry 1993". NobelPrize.org. Retrieved 6 November 2023.
- ^ "The Nobel Prize in Chemistry 2009". NobelPrize.org. Retrieved 6 November 2023.
- ^ "The Nobel Prize in Chemistry 2012". NobelPrize.org. Retrieved 6 November 2023.
- ^ "The Nobel Prize in Chemistry 2018". NobelPrize.org. Retrieved 6 November 2023.
- ^ "The Nobel Prize in Chemistry 2020". NobelPrize.org. Retrieved 6 November 2023.
- ^ a b "The Nobel Prize in Chemistry 2022". NobelPrize.org. Retrieved 6 November 2023.
- PMID 11848770.
- PMID 26626628.
- PMID 21503322.
- ^ "Bertozzi Group". Bertozzi Group. Retrieved 4 December 2023.
- PMID 28494316.
- PMID 18220367.
- PMID 15563194.
- PMID 33826699.
- PMID 35498070.
- PMID 23584057.
- PMID 30645048.
- ISBN 978-0-12-382239-0.
- PMID 9618476.
- PMID 28971554.
- PMID 15869385.
- PMID 19743430.
- S2CID 7586786.
- PMID 11182321.
- PMID 11813306.
- S2CID 25869680.
- PMID 30082609.
- S2CID 196001607. Retrieved 6 December 2023.
- PMID 31417196.
- PMID 18573077.
- PMID 11535813.
- PMID 15118093.
- PMID 15811807.
- PMID 11274392.
- S2CID 25527137.
- ^ "The Nobel Prize in Chemistry 2018". NobelPrize.org. Retrieved 3 October 2018.
- PMID 28951470.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "Catalysis and the Use of Energy by Cells". Molecular Biology of the Cell (4th ed.). Garland Science. Retrieved 6 December 2023.
- PMID 20388106.
- S2CID 11512358.
- PMID 9818143.
- ^ PMID 20884282.
- ^ S2CID 32604394.
- PMID 20834152.
- PMID 20479271.
- PMID 19489734.
- PMID 9224565.
- S2CID 22680843.
- S2CID 957274.
- S2CID 12994600.
- PMID 26119262.
- PMID 26672511.
- ISBN 978-3-527-68303-1.
- PMID 9618476.
- PMID 2649980.
- PMID 16689635.
- PMID 17881302.
- PMID 12782683.
- PMID 11090353.
- PMID 10447744.
- S2CID 1288600.
- PMID 15681376.
- PMID 18771748.
- S2CID 30591672.
- ^ "Coursework". American Chemical Society. Retrieved 2 December 2023.
- ^ "Chemistry 461: Synthetic Chemical Biology | UBC Chemistry". www.chem.ubc.ca. Retrieved 2 December 2023.
Further reading
- Dertinger SKW, Chiu DT, Jeon NL, Whitesides GM (2001). "Generation of gradients having complex shapes using microfluidic networks". Analytical Chemistry. 73 (6): 1240–1246. .
- Greif D, Pobigaylo N, Frage B, Becker A, Regtmeier J, Anselmetti D (September 2010). "Space- and time-resolved protein dynamics in single bacterial cells observed on a chip". Journal of Biotechnology. 149 (4): 280–288. PMID 20599571.
- Li L, Ismagilov RF (2010). "Protein crystallization using microfluidic technologies based on valves, droplets, and SlipChip". Annual Review of Biophysics. 39: 139–158. PMID 20192773.
- Lucchetta EM, Lee JH, Fu LA, Patel NH, Ismagilov RF (April 2005). "Dynamics of Drosophila embryonic patterning network perturbed in space and time using microfluidics". Nature. 434 (7037): 1134–1138. PMID 15858575.
- Melin J, Quake SR (2007). "Microfluidic large-scale integration: the evolution of design rules for biological automation". Annual Review of Biophysics and Biomolecular Structure. 36: 213–231. PMID 17269901.
- Shen F, Du W, Kreutz JE, Fok A, Ismagilov RF (October 2010). "Digital PCR on a SlipChip". Lab on a Chip. 10 (20): 2666–2672. PMID 20596567.
- Song H, Chen DL, Ismagilov RF (November 2006). "Reactions in droplets in microfluidic channels". Angewandte Chemie. 45 (44): 7336–7356. PMID 17086584.
- Spiller DG, Wood CD, Rand DA, White MR (June 2010). "Measurement of single-cell dynamics". Nature. 465 (7299): 736–745. S2CID 4426105.
- Tice JD, Song H, Lyon AD, Ismagilov RF (2003). "Formation of droplets and mixing in multiphase microfluidics at low values of the Reynolds and the capillary numbers". Langmuir. 19 (22): 9127–9133. .
- Vincent ME, Liu W, Haney EB, Ismagilov RF (March 2010). "Microfluidic stochastic confinement enhances analysis of rare cells by isolating cells and creating high density environments for control of diffusible signals". Chemical Society Reviews. 39 (3): 974–984. PMID 20179819.
- Weibel DB, Whitesides GM (December 2006). "Applications of microfluidics in chemical biology". Current Opinion in Chemical Biology. 10 (6): 584–591. PMID 17056296.
- Whitesides GM (July 2006). "The origins and the future of microfluidics". Nature. 442 (7101): 368–373. S2CID 205210989.
- Young EW, Beebe DJ (March 2010). "Fundamentals of microfluidic cell culture in controlled microenvironments". Chemical Society Reviews. 39 (3): 1036–1048. PMID 20179823.
Journals
- ACS Chemical Biology – The new Chemical Biology journal from the American Chemical Society.
- Bioorganic & Medicinal Chemistry – The Tetrahedron Journal for Research at the Interface of Chemistry and Biology
- ChemBioChem – A European Journal of Chemical Biology
- Chemical Biology– A point of access to chemical biology news and research from across RSC Publishing
- Cell Chemical Biology – An interdisciplinary journal that publishes papers of exceptional interest in all areas at the interface between chemistry and biology. chembiol.com
- Journal of Chemical Biology – A new journal publishing novel work and reviews at the interface between biology and the physical sciences, published by Springer. link
- Journal of the Royal Society Interface – A cross-disciplinary publication promoting research at the interface between the physical and life sciences
- Molecular BioSystems– Chemical biology journal with a particular focus on the interface between chemistry and the -omic sciences and systems biology.
- Nature Chemical Biology – A monthly multidisciplinary journal providing an international forum for the timely publication of significant new research at the interface between chemistry and biology.
- Wiley Encyclopedia of Chemical Biology link

