Chromosomal crossover

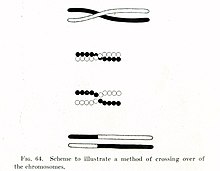
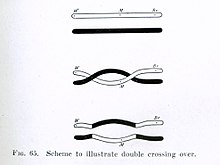
Chromosomal crossover, or crossing over, is the exchange of genetic material during
Crossing over was described, in theory, by Thomas Hunt Morgan; the term crossover was coined by Morgan and Eleth Cattell.[3] Hunt relied on the discovery of Frans Alfons Janssens who described the phenomenon in 1909 and had called it "chiasmatypie".[4] The term chiasma is linked, if not identical, to chromosomal crossover. Morgan immediately saw the great importance of Janssens' cytological interpretation of chiasmata to the experimental results of his research on the heredity of Drosophila. The physical basis of crossing over was first demonstrated by Harriet Creighton and Barbara McClintock in 1931.[5]
The linked frequency of crossing over between two
When Hotta et al. in 1977 compared meiotic crossing-over (recombination) in lily and mouse they concluded that diverse eukaryotes share a common pattern.[8] This finding suggested that chromosomal crossing over is a general characteristic of eukaryotic meiosis.
Origins
There are two popular and overlapping theories that explain the origins of crossing-over, coming from the different theories on the origin of
In 1931, Barbara McClintock discovered a triploid maize plant. She made key findings regarding corn's karyotype, including the size and shape of the chromosomes. McClintock used the prophase and metaphase stages of mitosis to describe the morphology of corn's chromosomes, and later showed the first ever cytological demonstration of crossing over in meiosis. Working with student Harriet Creighton, McClintock also made significant contributions to the early understanding of codependency of linked genes.
DNA repair theory
Crossing over and DNA repair are very similar processes, which utilize many of the same protein complexes.[10][11] In her report, "The Significance of Responses of the Genome to Challenge", McClintock studied corn to show how corn's genome would change itself to overcome threats to its survival. She used 450 self-pollinated plants that received from each parent a chromosome with a ruptured end. She used modified patterns of gene expression on different sectors of leaves of her corn plants to show that transposable elements ("controlling elements") hide in the genome, and their mobility allows them to alter the action of genes at different loci. These elements can also restructure the genome, anywhere from a few nucleotides to whole segments of chromosome. Recombinases and primases lay a foundation of nucleotides along the DNA sequence. One such particular protein complex that is conserved between processes is RAD51, a well conserved recombinase protein that has been shown to be crucial in DNA repair as well as cross over.[12] Several other genes in D. melanogaster have been linked as well to both processes, by showing that mutants at these specific loci cannot undergo DNA repair or crossing over. Such genes include mei-41, mei-9, hdm, spnA, and brca2.[citation needed] This large group of conserved genes between processes supports the theory of a close evolutionary relationship. Furthermore, DNA repair and crossover have been found to favor similar regions on chromosomes. In an experiment using radiation hybrid mapping on wheat's (Triticum aestivum L.) 3B chromosome, crossing over and DNA repair were found to occur predominantly in the same regions.[13] Furthermore, crossing over has been correlated to occur in response to stressful, and likely DNA damaging, conditions [14][15]
Links to bacterial transformation
The process of bacterial transformation also shares many similarities with chromosomal cross over, particularly in the formation of overhangs on the sides of the broken DNA strand, allowing for the annealing of a new strand. Bacterial transformation itself has been linked to DNA repair many times.[ Thus, this evidence suggests that it is a question of whether cross over is linked to DNA repair or bacterial transformation, as the two do not appear to be mutually exclusive. It is likely that crossing over may have evolved from bacterial transformation, which in turn developed from DNA repair, thus explaining the links between all three processes.
Chemistry
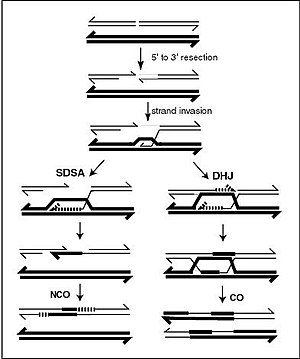
Meiotic recombination may be initiated by double-stranded breaks that are introduced into the DNA by exposure to DNA damaging agents,
MSH4 and MSH5
The MSH4 and MSH5 proteins form a hetero-oligomeric structure (heterodimer) in yeast and humans.[19][20][21] In the yeast Saccharomyces cerevisiae MSH4 and MSH5 act specifically to facilitate crossovers between homologous chromosomes during meiosis.[19] The MSH4/MSH5 complex binds and stabilizes double Holliday junctions and promotes their resolution into crossover products. An MSH4 hypomorphic (partially functional) mutant of S. cerevisiae showed a 30% genome wide reduction in crossover numbers, and a large number of meioses with non exchange chromosomes.[22] Nevertheless, this mutant gave rise to spore viability patterns suggesting that segregation of non-exchange chromosomes occurred efficiently. Thus in S. cerevisiae proper segregation apparently does not entirely depend on crossovers between homologous pairs.
Chiasma
The grasshopper Melanoplus femur-rubrum was exposed to an acute dose of X-rays during each individual stage of meiosis, and chiasma frequency was measured.[23] Irradiation during the leptotene-zygotene stages of meiosis (that is, prior to the pachytene period in which crossover recombination occurs) was found to increase subsequent chiasma frequency. Similarly, in the grasshopper Chorthippus brunneus, exposure to X-irradiation during the zygotene-early pachytene stages caused a significant increase in mean cell chiasma frequency.[24] Chiasma frequency was scored at the later diplotene-diakinesis stages of meiosis. These results suggest that X-rays induce DNA damages that are repaired by a crossover pathway leading to chiasma formation.
Class I and class II crossovers
Double strand breaks (DSBs) are repaired by two pathways to generate crossovers in eukaryotes.[25] The majority of them are repaired by MutL homologs MLH1 and MLH3, which defines the class I crossovers. The remaining are the result of the class II pathway, which is regulated by MUS81 endonuclease and FANCM translocase. There are interconnections between these two pathways—class I crossovers can compensate for the loss of class II pathway. In MUS81 knockout mice, class I crossovers are elevated, while total crossover counts at chiasmata are normal. However, the mechanisms underlining this crosstalk are not well understood. A recent study suggests that a scaffold protein called SLX4 may participate in this regulation.[26] Specifically, SLX4 knockout mice largely phenocopies the MUS81 knockout—once again, an elevated class I crossovers while normal chiasmata count. In FANCM knockout mice, the class II pathway is hyperactivated, resulting in increased numbers of crossovers that are independent of the MLH1/MLH3 pathway.[27]
Consequences

In most
Recombination results in a new arrangement of maternal and paternal alleles on the same chromosome. Although the same genes appear in the same order, some alleles are different. In this way, it is theoretically possible to have any combination of parental alleles in an offspring, and the fact that two alleles appear together in one offspring does not have any influence on the
Non-homologous crossover
Crossovers typically occur between homologous regions of matching chromosomes, but similarities in sequence and other factors can result in mismatched alignments. Most DNA is composed of base pair sequences repeated very large numbers of times.[29] These repetitious segments, often referred to as satellites, are fairly homogeneous among a species.[29] During DNA replication, each strand of DNA is used as a template for the creation of new strands using a partially-conserved mechanism; proper functioning of this process results in two identical, paired chromosomes, often called sisters. Sister chromatid crossover events are known to occur at a rate of several crossover events per cell per division in eukaryotes.[29] Most of these events involve an exchange of equal amounts of genetic information, but unequal exchanges may occur due to sequence mismatch. These are referred to by a variety of names, including non-homologous crossover, unequal crossover, and unbalanced recombination, and result in an insertion or deletion of genetic information into the chromosome. While rare compared to homologous crossover events, these mutations are drastic, affecting many loci at the same time. They are considered the main driver behind the generation of gene duplications and are a general source of mutation within the genome.[30]
The specific causes of non-homologous crossover events are unknown, but several influential factors are known to increase the likelihood of an unequal crossover. One common vector leading to unbalanced recombination is the repair of
The presence of transposable elements is another influential element of non-homologous crossover. Repetitive regions of code characterize transposable elements; complementary but non-homologous regions are ubiquitous within transposons. Because chromosomal regions composed of transposons have large quantities of identical, repetitious code in a condensed space, it is thought that transposon regions undergoing a crossover event are more prone to erroneous complementary match-up;[33] that is to say, a section of a chromosome containing a lot of identical sequences, should it undergo a crossover event, is less certain to match up with a perfectly homologous section of complementary code and more prone to binding with a section of code on a slightly different part of the chromosome. This results in unbalanced recombination, as genetic information may be either inserted or deleted into the new chromosome, depending on where the recombination occurred.
While the motivating factors behind unequal recombination remain obscure, elements of the physical mechanism have been elucidated.
See also
- Unequal crossing over
- Coefficient of coincidence
- Genetic distance
- Independent assortment
- Mitotic crossover
- Recombinant frequency
References
- ^ Griffiths AJ, Gelbart WM, Miller JH, et al. (1999). "Mitotic Crossing-Over". Modern Genetic Analysis. New York: W. H. Freeman.
- PMID 25590558.
- .
- PMID 22701051.
- PMID 16587654. (Original paper)
- ISBN 978-3-540-07668-1.
- ISBN 0-19-509442-5.
- S2CID 4268089.
- ^ S2CID 86663600.
- PMID 24635147.
- PMID 20195513.
- PMID 360220.
- PMID 22827734.
- PMID 20223252.
- PMID 15306305.
- PMID 21169481.
- S2CID 8294596.
- PMID 15899844.
- ^ PMID 9374523.
- PMID 9787078.
- PMID 10029069.
- PMID 25467183.
- PMID 5797806.
- PMID 5289295.
- PMID 18787696.
- PMID 21655083.
- PMID 37601968.
- ^ a b Darling D. "Genetic recombination".
- ^ PMID 1251186.
- ISBN 9780878932665.
- ^ PMID 15557293.
- PMID 2060774.
- PMID 23212949.
- ^ PMID 15952900.
- S2CID 19219813.



