Codon usage bias
Codon usage bias refers to differences in the frequency of occurrence of
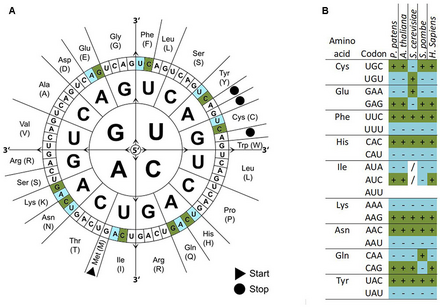
There are 64 different codons (61 codons encoding for amino acids and 3 stop codons) but only 20 different translated amino acids. The overabundance in the number of codons allows many amino acids to be encoded by more than one codon. Because of such redundancy it is said that the genetic code is degenerate. The genetic codes of different organisms are often biased towards using one of the several codons that encode the same amino acid over the others—that is, a greater frequency of one will be found than expected by chance. How such biases arise is a much debated area of molecular evolution. Codon usage tables detailing genomic codon usage bias for organisms in GenBank and RefSeq can be found in the HIVE-Codon Usage Tables (HIVE-CUTs) project,[1] which contains two distinct databases, CoCoPUTs and TissueCoCoPUTs. Together, these two databases provide comprehensive, up-to-date codon, codon pair and dinucleotide usage statistics for all organisms with available sequence information and 52 human tissues, respectively.[2][3]
It is generally acknowledged that codon biases reflect the contributions of 3 main factors,
The nature of the codon usage-tRNA optimization has been fiercely debated. It is not clear whether codon usage drives tRNA evolution or vice versa. At least one mathematical model has been developed where both codon usage and tRNA expression co-evolve in feedback fashion (i.e., codons already present in high frequencies drive up the expression of their corresponding tRNAs, and tRNAs normally expressed at high levels drive up the frequency of their corresponding codons). However, this model does not seem to yet have experimental confirmation. Another problem is that the evolution of tRNA genes has been a very inactive area of research.[citation needed]
Contributing factors
Different factors have been proposed to be related to codon usage bias, including gene expression level (reflecting selection for optimizing the translation process by tRNA abundance),
Evolutionary theories
Mutational bias versus selection
Although the mechanism of codon bias selection remains controversial, possible explanations for this bias fall into two general categories. One explanation revolves around the selectionist theory, in which codon bias contributes to the efficiency and/or accuracy of protein expression and therefore undergoes
The second explanation for codon usage can be explained by mutational bias, a theory which posits that codon bias exists because of nonrandomness in the mutational patterns. In other words, some codons can undergo more changes and therefore result in lower equilibrium frequencies, also known as “rare” codons. Different organisms also exhibit different mutational biases, and there is growing evidence that the level of genome-wide GC content is the most significant parameter in explaining codon bias differences between organisms. Additional studies have demonstrated that codon biases can be statistically predicted in
Mutation-selection-drift balance model
To reconcile the evidence from both
Consequences of codon composition
Effect on RNA secondary structure
Because
Effect on transcription or gene expression
Specialized codon bias is further seen in some
Effect on speed of translation elongation
Generally speaking for highly expressed genes, translation elongation rates are faster along transcripts with higher codon adaptation to tRNA pools, and slower along transcripts with rare codons. This correlation between codon translation rates and cognate tRNA concentrations provides additional modulation of translation elongation rates, which can provide several advantages to the organism. Specifically, codon usage can allow for global regulation of these rates, and rare codons may contribute to the accuracy of translation at the expense of speed.[26]
Effect on protein folding
Methods of analysis
In the field of
References
- PMID 28865429.
- S2CID 139104807.
- PMID 31982380.
- PMID 21646514.
- .
- hdl:20.500.12210/34500.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 8709146.
- S2CID 8582630.
- PMID 10570992.
- PMID 10784043.
- PMID 31174582.
- PMID 10858656.
- PMID 26504241.
- PMID 11719972.
- PMID 12364606.
- PMID 18397532.
- PMID 23637123.
- PMID 9724767.
- PMID 27842572.
- ^ S2CID 7085012.
- PMID 22921354.
- PMID 17017124.
- S2CID 202555575.
- PMID 29624661.
- ^ PMID 21102527.
- ^ PMID 24688635.
- S2CID 21862217.
- PMID 6175758.
- PMID 20453079.
- PMID 3547335.
- ^ Peden J (2005-04-15). "Codon usage indices". Correspondence Analysis of Codon Usage. SourceForge. Retrieved 2010-10-20.
- PMID 18940873.
External links
- Composition Analysis Toolkit Archived 2020-07-26 at the Wayback Machine: estimating codon usage bias and its statistical significance
- HIVE-Codon Usage Table database
- Codon Usage Database
- CodonW
- GCUA - General Codon Usage Analysis
- Graphical Codon Usage Analyser
- JCat - Java Codon Usage Adaptation Tool
- INCA - Interactive Codon Analysis software
- ACUA - Automated Codon Usage Analysis Tool Archived 2020-07-26 at the Wayback Machine
- OPTIMIZER - Codon usage optimization
- HEG-DB - Highly Expressed Genes Database
- E-CAI - Expected value of Codon Adaptation Index
- CAIcal -Set of tools to assess codon usage adaptation
- scRCA - Automatic determination of translational codon usage bias
- Online Synonymous Codon Usage Analyses with the ade4 and seqinR packages
- Genetic Algorithm Simulation for Codon Optimization
