Comparative genomics
Comparative genomics is a field of
Virtually started as soon as the whole genomes of two organisms became available (that is, the genomes of the bacteria
History
See also:
Comparative genomics has a root in the comparison of
The first complete genome sequence of a cellular organism, that of Haemophilus influenzae Rd, was published in 1995.[12] The second genome sequencing paper was of the small parasitic bacterium Mycoplasma genitalium published in the same year.[13] Starting from this paper, reports on new genomes inevitably became comparative-genomic studies.[8]
Microbial genomes. The first high-resolution whole genome comparison system of microbial genomes of 10-15kbp was developed in 1998 by Art Delcher, Simon Kasif and Steven Salzberg and applied to the comparison of entire highly related microbial organisms with their collaborators at the Institute for Genomic Research (TIGR). The system is called MUMMER and was described in a publication in Nucleic Acids Research in 1999. The system helps researchers to identify large rearrangements, single base mutations, reversals, tandem repeat expansions and other polymorphisms. In bacteria, MUMMER enables the identification of polymorphisms that are responsible for virulence, pathogenicity, and anti-biotic resistance. The system was also applied to the Minimal Organism Project at TIGR and subsequently to many other comparative genomics projects.
Eukaryote genomes.
With the publication of the large genomes of vertebrates in the 2000s, including
Evolutionary principles
One character of biology is evolution,
Similarity of related genomes is the basis of comparative genomics. If two creatures have a recent common ancestor, the differences between the two species genomes are evolved from the ancestors' genome. The closer the relationship between two organisms, the higher the similarities between their genomes. If there is close relationship between them, then their genome will display a linear behaviour (synteny), namely some or all of the genetic sequences are conserved. Thus, the genome sequences can be used to identify gene function, by analyzing their homology (sequence similarity) to genes of known function.

Orthologous sequences are related sequences in different species: a gene exists in the original species, the species divided into two species, so genes in new species are orthologous to the sequence in the original species. Paralogous sequences are separated by gene cloning (gene duplication): if a particular gene in the genome is copied, then the copy of the two sequences is paralogous to the original gene. A pair of orthologous sequences is called orthologous pairs (orthologs), a pair of paralogous sequence is called collateral pairs (paralogs). Orthologous pairs usually have the same or similar function, which is not necessarily the case for collateral pairs. In collateral pairs, the sequences tend to evolve into having different functions.
Comparative genomics exploits both similarities and differences in the
One of the important goals of the field is the identification of the mechanisms of eukaryotic genome evolution. It is however often complicated by the multiplicity of events that have taken place throughout the history of individual lineages, leaving only distorted and superimposed traces in the genome of each living organism. For this reason comparative genomics studies of small
Methods
Computational approaches are necessary for genome comparisons, given the large amount of data encoded in genomes. Many tools are now publicly available, ranging from whole genome comparisons to gene expression analysis.[22] This includes approaches from systems and control, information theory, string analysis and data mining.[23] Computational approaches will remain critical for research and teaching, especially when information science and genome biology is taught in conjunction.[24]

Comparative genomics starts with basic comparisons of genome size and gene density. For instance, genome size is important for coding capacity and possibly for regulatory reasons. High gene density facilitates
Sequence alignment
Alignments are used to capture information about similar sequences such as ancestry, common evolutionary descent, or common structure and function. Alignments can be done for both genetic and protein sequences.[26][27] Alignments consist of local or global pairwise alignments, and multiple sequence alignments. One way to find global alignments is to use a dynamic programming algorithm known as Needleman-Wunsch algorithm. This algorithm can be modified and used to find local alignments.

Phylogenetic reconstruction
Another computational method for comparative genomics is phylogenetic reconstruction. It is used to describe evolutionary relationships in terms of common ancestors. The relationships are usually represented in a tree called a phylogenetic tree. Similarly, coalescent theory is a retrospective model to trace alleles of a gene in a population to a single ancestral copy shared by members of the population. This is also known as the most recent common ancestor. Analysis based on coalescence theory tries predicting the amount of time between the introduction of a mutation and a particular allele or gene distribution in a population. This time period is equal to how long ago the most recent common ancestor existed. The inheritance relationships are visualized in a form similar to a phylogenetic tree. Coalescence (or the gene genealogy) can be visualized using dendrograms.[28]
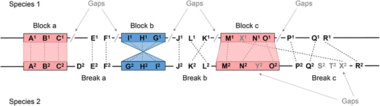
Genome maps
An additional method in comparative genomics is
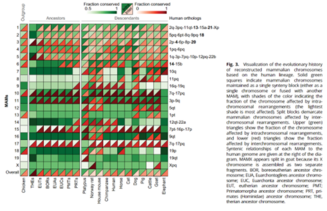

Tools
Computational tools for analyzing sequences and complete genomes are developing quickly due to the availability of large amount of genomic data. At the same time, comparative analysis tools are progressed and improved. In the challenges about these analyses, it is very important to visualize the comparative results.[31]
Visualization of sequence conservation is a tough task of comparative sequence analysis. As we know, it is highly inefficient to examine the alignment of long genomic regions manually. Internet-based genome browsers provide many useful tools for investigating genomic sequences due to integrating all sequence-based biological information on genomic regions. When we extract large amount of relevant biological data, they can be very easy to use and less time-consuming.[31]
- UCSC Browser: This site contains the reference sequence and working draft assemblies for a large collection of genomes.[32]
- Ensembl: The Ensembl project produces genome databases for vertebrates and other eukaryotic species, and makes this information freely available online.[33]
- MapView: The Map Viewer provides a wide variety of genome mapping and sequencing data.[34]
- VISTA is a comprehensive suite of programs and databases for comparative analysis of genomic sequences. It was built to visualize the results of comparative analysis based on DNA alignments. The presentation of comparative data generated by VISTA can easily suit both small and large scale of data.[35]
- BlueJay Genome Browser: a stand-alone visualization tool for the multi-scale viewing of annotated genomes and other genomic elements.[36]
An advantage of using online tools is that these websites are being developed and updated constantly. There are many new settings and content can be used online to improve efficiency.[31]
Selected applications
Agriculture
Medicine
Vaccine development
The medical field also benefits from the study of comparative genomics. In an approach known as
Mouse models in immunology
T-cell immune receptors are important in seeing the world of pathogens in the cellular immune system. One of the reasons for sequencing the human and mouse TCR loci was to match the orthologous gene family sequences and discover conserved areas using comparative genomics. These, it was thought, would reflect two sorts of biological information: (1) exons and (2) regulatory sequences. In fact, the majority of V, D, J, and C exons could be identified in this method. The variable regions are encoded by multiple unique DNA elements that are rearranged and connected during T cell (TCR) differentiation: variable (V), diversity (D), and joining (J) elements for the and polypeptides; and V and J elements for the and polypeptides.[Figure 1] However, several short noncoding conserved blocks of the genome had been shown. Both human and mouse motifs are largely clustered in the 200 bp [Figure 2], the known 3′ enhancers in the TCR/ were identified, and a conserved region of 100 bp in the mouse J intron was subsequently shown to have a regulatory function.

Comparisons of the genomic sequences within each physical site or location of a specific gene on a chromosome (locs) and across species allow for research on other mechanisms and other regulatory signals. Some suggest new hypotheses about the evolution of TCRs, to be tested (and improved) by comparison to the TCR gene complement of other vertebrate species. A comparative genomic investigation of humans and mice will obviously allow for the discovery and annotation of many other genes, as well as identifying in other species for regulatory sequences.[43]
Research
Comparative genomics also opens up new avenues in other areas of research. As DNA sequencing technology has become more accessible, the number of
A notable case of this increased potency is found in recent primate research. Comparative genomic methods have allowed researchers to gather information about genetic variation, differential gene expression, and evolutionary dynamics in primates that were indiscernible using previous data and methods.[44]
Great Ape Genome Project
The Great Ape Genome Project used comparative genomic methods to investigate genetic variation with reference to the six great ape species, finding healthy levels of variation in their gene pool despite shrinking population size.[45] Another study showed that patterns of DNA methylation, which are a known regulation mechanism for gene expression, differ in the prefrontal cortex of humans versus chimps, and implicated this difference in the evolutionary divergence of the two species.[46]
See also
- Data mining
- Molecular evolution
- Comparative anatomy
- Homology
- Sequence mining
- Alignment-free sequence analysis
References
- PMID 18650965.

- ^ a b c Touchman J (2010). "Comparative Genomics". Nature Education Knowledge. 3 (10): 13.
- ^ S2CID 5491782.
- ^ a b Russel PJ, Hertz PE, McMillan B (2011). Biology: The Dynamic Science (2nd ed.). Belmont, CA: Brooks/Cole. pp. 409–410.
- ISBN 9781405101202.
- PMID 14624258.

- S2CID 43171654.
- ^ a b c d Koonin EV, Galperin MY (2003). Sequence - Evolution - Function: Computational approaches in comparative genomics. Dordrecht: Springer Science+Business Media.
- ^ PMID 22199376.
- PMID 6384934.
- PMID 3012465.
- PMID 7542800.
- S2CID 29825758.
- S2CID 16763139.
- PMID 9851916.
- PMID 10731132.
- PMID 10731134.
- PMID 10899144.

- S2CID 2037634.
- PMID 14624247.

- PMC 261884.

- ISBN 978-0-521-67191-0.
- PMID 25984837.
- PMID 22046119.

- ^ PMID 36161960.
- PMID 29206392. Retrieved 2022-12-18.
- S2CID 226247797.
- S2CID 2041895.
- PMID 29382321.
- PMID 19347649.
- ^ PMID 21250292.
- ^ "UCSC Browser".
- ^ "Ensembl Genome Browser". Archived from the original on 2013-10-21.
- ^ "Map Viewer".
- ^ "VISTA tools".
- S2CID 34553139.
- S2CID 439442.

- S2CID 13358998.

- PMID 22882709.

- PMID 15994562.

- PMID 18676672.

- ^ "Group a Streptococcus Vaccine Target Candidates Identified from Global Genome Set". 28 May 2019.
- ^ PMID 11567625.
- PMID 24709753.

- PMID 23823723.

- PMID 22922032.

Further reading
- Kellis M, Patterson N, Endrizzi M, Birren B, Lander ES (May 2003). "Sequencing and comparison of yeast species to identify genes and regulatory elements". Nature. 423 (6937): 241–254. S2CID 1530261.
- Cliften P, Sudarsanam P, Desikan A, Fulton L, Fulton B, Majors J, et al. (July 2003). "Finding functional features in Saccharomyces genomes by phylogenetic footprinting". Science. 301 (5629): 71–76. S2CID 1305166.
- Boffelli D, McAuliffe J, Ovcharenko D, Lewis KD, Ovcharenko I, Pachter L, Rubin EM (February 2003). "Phylogenetic shadowing of primate sequences to find functional regions of the human genome". Science. 299 (5611): 1391–1394. S2CID 17217612.
- Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, Lafontaine I, et al. (July 2004). "Genome evolution in yeasts". Nature. 430 (6995): 35–44. S2CID 4399964.
- Filipski A, Kumar S (2005). "Comparative genomics in eukaryotes". In Gregory TR (ed.). The Evolution of the Genome. San Diego: Elsevier. pp. 521–583.
- Gregory TR, DeSalle R (2005). "Comparative genomics in prokaryotes". In Gregory TR (ed.). The Evolution of the Genome. San Diego: Elsevier. pp. 585–675.
- Xie X, Lu J, Kulbokas EJ, Golub TR, Mootha V, Lindblad-Toh K, et al. (March 2005). "Systematic discovery of regulatory motifs in human promoters and 3' UTRs by comparison of several mammals". Nature. 434 (7031): 338–345. PMID 15735639.
- Champ PC, Binnewies TT, Nielsen N, Zinman G, Kiil K, Wu H, et al. (March 2006). "Genome update: purine strand bias in 280 bacterial genomes". Microbiology. 152 (Pt 3): 579–583. PMID 16514138.
- Kumar L, Breakspear A, Kistler C, Ma LJ, Xie X (March 2010). "Systematic discovery of regulatory motifs in Fusarium graminearum by comparing four Fusarium genomes". BMC Genomics. 11: 208. PMID 20346147.

- Batzoglou S, Pachter L, Mesirov JP, Berger B, Lander ES (July 2000). "Human and mouse gene structure: comparative analysis and application to exon prediction". Genome Research. 10 (7): 950–958. PMID 10899144.

External links
This article's use of external links may not follow Wikipedia's policies or guidelines. (February 2017) |
- Genomes OnLine Database (GOLD)
- Genome News Network
- JCVI Comprehensive Microbial Resource
- Pathema: A Clade Specific Bioinformatics Resource Center
- CBS Genome Atlas Database Archived 2016-05-16 at the Portuguese Web Archive
- The UCSC Genome Browser
- The U.S. National Human Genome Research Institute
- Ensembl The EnsemblGenome Browser
- Genolevures, comparative genomics of the Hemiascomycetous yeasts
- Phylogenetically Inferred Groups (PhIGs), a recently developed method incorporates phylogenetic signals in building gene clusters for use in comparative genomics.
- Metazome Archived 2006-08-10 at the Wayback Machine, a resource for the phylogenomic exploration and analysis of Metazoan gene families.
- IMG The Integrated Microbial Genomes system, for comparative genome analysis by the DOE-JGI.
- Dcode.org Dcode.org Comparative Genomics Center.
- SUPERFAMILY Protein annotations for all completely sequenced organisms
- Comparative Genomics
- Blastology and Open Source: Needs and Deeds
- Alignment-free comparative Genomics tool
