Coronavirus spike protein
| Coronavirus spike glycoprotein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
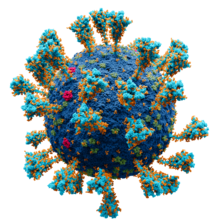 Model of the external structure of the SARS-CoV-2 virion.[1]
● Blue: envelope ● Turquoise: spike glycoprotein (S) ● Red: envelope proteins (E) ● Green: membrane proteins (M) ● Orange: glycan | |||||||||
| Identifiers | |||||||||
| Symbol | CoV_S1 | ||||||||
| Pfam | PF01600 | ||||||||
| InterPro | IPR002551 | ||||||||
| |||||||||
Spike (S) glycoprotein (sometimes also called spike protein,
The function of the spike
Spike glycoprotein is highly
Structure
The spike protein is very large, often 1200 to 1400
Spike glycoprotein forms
S1
| Betacoronavirus spike glycoprotein S1, receptor binding | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | bCoV_S1_RBD | ||||||||
| Pfam | PF09408 | ||||||||
| InterPro | IPR018548 | ||||||||
| |||||||||
| Betacoronavirus-like spike glycoprotein S1, N-terminal | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | bCoV_S1_N | ||||||||
| Pfam | PF16451 | ||||||||
| InterPro | IPR032500 | ||||||||
| |||||||||
The S1 region of the spike glycoprotein is responsible for interacting with receptor molecules on the surface of the host cell in the first step of
The CTD is responsible for the interactions of
Within coronavirus lineages, as well as across the four major coronavirus subgroups, the S1 region is less well
S2
| Coronavirus spike glycoprotein S2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | CoV_S2 | ||||||||
| Pfam | PF01601 | ||||||||
| InterPro | IPR002552 | ||||||||
| |||||||||
The S2 region of spike glycoprotein is responsible for
Relative to S1, the S2 region is very well
Post-translational modifications

Spike glycoprotein is heavily
Spike proteins are activated through proteolytic cleavage. They are cleaved by host cell proteases at the S1-S2 boundary and later at what is known as the S2' site at the N-terminus of the fusion peptide.[4][5][7][8]
Conformational change
Like other
Expression and localization
Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2, indicating the location of the S gene | |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser (UCSC) | |
The
The spike protein is not required for viral assembly or the formation of
Function
The spike protein is responsible for
Attachment
The interaction of the receptor-binding domain in the S1 region with its target receptor on the cell surface initiates the process of viral entry. Different coronaviruses target different cell-surface receptors, sometimes using sugar molecules such as
| Species | Genus | Receptor | Reference |
|---|---|---|---|
| Human coronavirus 229E | Alphacoronavirus | Aminopeptidase N
|
[4][30] |
| Human coronavirus NL63 | Alphacoronavirus | Angiotensin-converting enzyme 2 | [4][31] |
| Human coronavirus HKU1 | Betacoronavirus | N-acetyl-9-O-acetylneuraminic acid | [28][32] |
| Human coronavirus OC43 | Betacoronavirus | N-acetyl-9-O-acetylneuraminic acid | [4][33] |
Middle East respiratory syndrome–related coronavirus
|
Betacoronavirus | Dipeptidyl peptidase-4 | [4][34] |
Severe acute respiratory syndrome coronavirus
|
Betacoronavirus | Angiotensin-converting enzyme 2 | [4][35] |
Severe acute respiratory syndrome coronavirus 2
|
Betacoronavirus | N-acetylneuraminic acid
|
[5][9][36] |
Proteolytic cleavage
Membrane fusion

Like other
In addition to fusion of viral and host cell membranes, some coronavirus spike proteins can initiate membrane fusion between infected cells and neighboring cells, forming
Immunogenicity
Because it is exposed on the surface of the virus, the spike protein is a major
COVID-19 response
Vaccines
In response to the
According to a study published in January 2023, markedly elevated levels of full-length spike protein unbound by antibodies were found in people who developed postvaccine myocarditis (vs. controls that remained healthy). However, these results do not alter the risk-benefit ratio favoring vaccination against COVID-19 to prevent severe clinical outcomes.[54][non-primary source needed]
Monoclonal antibodies
SARS-CoV-2 variants
Throughout the
Spike protein mutations raise concern because they may affect
The SARS-CoV-2 Omicron variant is notable for having an unusually high number of mutations in the spike protein.[77] The SARS CoV-2 spike gene (S gene, S-gene) mutation 69–70del (Δ69-70) causes a TaqPath PCR test probe to not bind to its S gene target, leading to S gene target failure (SGTF) in SARS CoV-2 positive samples. This effect was used as a marker to monitor the propagation of the Alpha variant[78][79] and the Omicron variant.[80]
Additional Key Role in Illness
In 2021, Circulation Research and Salk had a new study that proves COVID-19 can be also a vascular disease, not only respiratory disease. The scientists created an “pseudovirus”, surrounded by SARS-CoV-2 spike proteins but without any actual virus. And pseudovirus resulted in damaging lungs and arteries of animal models. It shows SARS-CoV-2 spike protein alone can cause vascular disease and could explain some covid-19 patients who suffered from strokes, or other vascular problems in other parts of human body at the same time. The team replicated the process by removing replicating capabilities of virus and showed the same damaging effect on vascular cells again.[81][82]
Misinformation
During the
Evolution, conservation and recombination
The
Within the S1 region, the N-terminal domain (NTD) is more conserved than the C-terminal domain (CTD).
References
- ^ Solodovnikov, Alexey; Arkhipova, Valeria (29 July 2021). "Достоверно красиво: как мы сделали 3D-модель SARS-CoV-2" [Truly beautiful: how we made the SARS-CoV-2 3D model] (in Russian). N+1. Archived from the original on 30 July 2021. Retrieved 30 July 2021.
- ^ ISBN 9780128145166.
- ^ PMID 16877062.
- ^ PMID 32833200.
- ^ PMID 34125455.
- PMC 7086490.
- ^ PMID 27578435.
- ^ PMID 29275820.
- ^ PMID 33116300.
- ^ PMID 33123165.
- ^ S2CID 221503034.
- ^ PMID 33619260.
- ^ PMID 32075877.
- ^ PMID 27712627.
- ^ https://doi.org/10.1126/science.abm3125
- ^ https://doi.org/10.1016/S0021-9258(18)63732-9
- ^ Zimmer, Carl (9 October 2020). "The Coronavirus Unveiled". The New York Times. Retrieved 12 August 2021.
- ^ PMID 33140034.
- PMID 32363391.
- ^ PMID 25855243.
- PMID 32155444.
- ^ PMID 33208793.
- PMID 32694201.
- ^ PMID 33229438.
- PMID 32760062.
- ^ PMID 21130884.
- PMID 32805734.
- ^ PMID 28933406.
- .
- PMID 1350662.
- PMID 15897467.
- PMID 25926653.
- PMID 8317096.
- PMID 23486063.
- PMID 14647384.
- ^ https://doi.org/10.1126/science.abm3125
- ^ PMID 34611326.
- PMID 30712865.
- PMID 32681106.
- ^ PMID 26935856.
- PMID 22816037.
- ^ PMID 33051876.
- PMID 33879858.
- PMID 33827113.
- PMID 33981020.
- ^ PMID 32566896.
- PMID 33215063.
- PMID 32527802.
- ^ PMID 34075212.
- PMID 35476528.
- S2CID 258717083.
- S2CID 233186026.
- PMID 34133867.
- S2CID 255475007.
Extensive antibody profiling and T-cell responses in the individuals who developed postvaccine myocarditis were essentially indistinguishable from those of vaccinated control subjects, [...] A notable finding was that markedly elevated levels of full-length spike protein (33.9±22.4 pg/mL), unbound by antibodies, were detected in the plasma of individuals with postvaccine myocarditis, [...] (unpaired t test; P<0.0001).
- PMID 32540901.
- ^ a b "Therapeutic Management of Nonhospitalized Adults With COVID-19". Covid-19 Treatment Guidelines. National Institutes of Health. Archived from the original on 4 December 2021. Retrieved 11 August 2021.
- ^ "etesevimab". IUPHAR/BPS Guide to Pharmacology. Retrieved 10 February 2021.
- ^ "Lilly announces agreement with U.S. government to supply 300,000 vials of investigational neutralizing antibody bamlanivimab (LY-CoV555) in an effort to fight COVID-19". Eli Lilly and Company (Press release). 28 October 2020.
- ^ "Casirivimab injection, solution, concentrate Imdevimab injection, solution, concentrate REGEN-COV- casirivimab and imdevimab kit". DailyMed. Retrieved 18 March 2021.
- ^ "Sotrovimab injection, solution, concentrate". DailyMed. Retrieved 15 June 2021.
- ^ PMID 32742035.
- ^ PMID 34071984.
- PMID 33042039.
- PMID 32238584.
- PMID 32820179.
- PMID 32697968.
- PMID 33243994.
- PMID 33220921.
- PMID 33083031.
- S2CID 244647259.
- PMID 33761203.
- PMID 35022734.
- PMID 34435953.
- PMID 35235558.
- S2CID 237254466.
- S2CID 235249387.
- ^ "Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of Concern". World Health Organization. 26 November 2021. Archived from the original on 26 November 2021. Retrieved 26 November 2021.
- PMID 33830171.
- ^ Methods for the detection and identification of SARS-CoV-2 variants (Technical report). Stockholm and Copenhagen: European Centre for Disease Prevention and Control/World Health Organization Regional Office for Europe. 3 March 2021. Diagnostic screening assays of known VOCs.
- ^ SARS-CoV-2 variants of concern and variants under investigation in England Variant of concern: Omicron, VOC21NOV-01 (B.1.1.529), technical briefing 30 (PDF) (Briefing). Public Health England. 3 December 2021. GOV-10547. Archived (PDF) from the original on 11 December 2021. Retrieved 15 December 2021.
- ^ "The novel coronavirus' spike protein plays additional key role in illness". Salk researchers. 30 April 2021. Archived from the original on 1 December 2022.
- S2CID 232430540.
- ^ "COVID-19 vaccines are not 'cytotoxic'" (Fact check). Reuters. 18 June 2021.
- ^ Gorski DH (24 May 2021). "The 'deadly' coronavirus spike protein (according to antivaxxers)". Science-Based Medicine.
- PMID 35579205.
- ^ McCarthy B (5 May 2021). "Debunking the anti-vaccine hoax about 'vaccine shedding'". PolitiFact. Retrieved 11 May 2021.
- ^ Fiore K (29 April 2021). "The Latest Anti-Vax Myth: 'Vaccine Shedding'". MedPage Today. Retrieved 11 May 2021.
- PMID 32634411.
- PMID 22205743.
- PMID 29684066.
External links
- Scudellari, Megan (28 July 2021). "How the coronavirus infects cells — and why Delta is so dangerous". Nature. Retrieved 15 August 2021.
- Iwasa, Janet; Meyer, Miriah; Lex, Alexander; Rogers, Jen; Liu, Ann (Hui); Riggi, Margot. "Building a visual consensus model of the SARS-CoV-2 life cycle". Animation Lab. University of Utah. Retrieved 15 August 2021.

