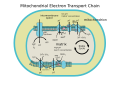
Cytochrome c oxidase
| Cytochrome c oxidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Cytochrome c oxidase | |
|---|---|
 | |
| Identifiers | |
| Symbol | Cytochrome c oxidase |
| OPM superfamily | 4 |
| OPM protein | 2dyr |
| Membranome | 257 |
The
It is the last enzyme in the respiratory electron transport chain of cells located in the membrane. It receives an electron from each of four cytochrome c molecules and transfers them to one oxygen molecule and four protons, producing two molecules of water. In addition to binding the four protons from the inner aqueous phase, it transports another four protons across the membrane, increasing the transmembrane difference of proton electrochemical potential, which the ATP synthase then uses to synthesize ATP.
Structure
The complex
The complex is a large
Crystallographic studies of cytochrome c oxidase show an unusual post-translational modification, linking C6 of Tyr(244) and the ε-N of His(240) (bovine enzyme numbering). It plays a vital role in enabling the cytochrome a3- CuB binuclear center to accept four electrons in reducing molecular oxygen and four protons to water. The mechanism of reduction was formerly thought to involve a peroxide intermediate, which was believed to lead to superoxide production. However, the currently accepted mechanism involves a rapid four-electron reduction involving immediate oxygen–oxygen bond cleavage, avoiding any intermediate likely to form superoxide.[4]: 865–866
The conserved subunits
| No. | Subunit name | Human protein | Protein description from UniProt | Pfam family with Human protein |
|---|---|---|---|---|
| 1 | Cox1 | COX1_HUMAN |
Cytochrome c oxidase subunit 1 | Pfam PF00115 |
| 2 | Cox2 | COX2_HUMAN |
Cytochrome c oxidase subunit 2 | Pfam PF02790, Pfam PF00116 |
| 3 | Cox3 | COX3_HUMAN |
Cytochrome c oxidase subunit 3 | Pfam PF00510 |
| 4 | Cox4i1 | COX41_HUMAN | Cytochrome c oxidase subunit 4 isoform 1, mitochondrial | Pfam PF02936 |
| 5 | Cox4a2 | COX42_HUMAN | Cytochrome c oxidase subunit 4 isoform 2, mitochondrial | Pfam PF02936 |
| 6 | Cox5a | COX5A_HUMAN | Cytochrome c oxidase subunit 5A, mitochondrial | Pfam PF02284 |
| 7 | Cox5b | COX5B_HUMAN | Cytochrome c oxidase subunit 5B, mitochondrial | Pfam PF01215 |
| 8 | Cox6a1 | CX6A1_HUMAN | Cytochrome c oxidase subunit 6A1, mitochondrial | Pfam PF02046 |
| 9 | Cox6a2 | CX6A2_HUMAN | Cytochrome c oxidase subunit 6A2, mitochondrial | Pfam PF02046 |
| 10 | Cox6b1 | CX6B1_HUMAN | Cytochrome c oxidase subunit 6B1 | Pfam PF02297 |
| 11 | Cox6b2 | CX6B2_HUMAN | Cytochrome c oxidase subunit 6B2 | Pfam PF02297 |
| 12 | Cox6c | COX6C_HUMAN | Cytochrome c oxidase subunit 6C | Pfam PF02937 |
| 13 | Cox7a1 | CX7A1_HUMAN | Cytochrome c oxidase subunit 7A1, mitochondrial | Pfam PF02238 |
| 14 | Cox7a2 | CX7A2_HUMAN | Cytochrome c oxidase subunit 7A2, mitochondrial | Pfam PF02238 |
| 15 | Cox7a3 | COX7S_HUMAN | Putative cytochrome c oxidase subunit 7A3, mitochondrial | Pfam PF02238 |
| 16 | Cox7b | COX7B_HUMAN | Cytochrome c oxidase subunit 7B, mitochondrial | Pfam PF05392 |
| 17 | Cox7c | COX7C_HUMAN | Cytochrome c oxidase subunit 7C, mitochondrial | Pfam PF02935 |
| 18 | Cox7r | COX7R_HUMAN | Cytochrome c oxidase subunit 7A-related protein, mitochondrial | Pfam PF02238 |
| 19 | Cox8a | COX8A_HUMAN | Cytochrome c oxidase subunit 8A, mitochondrial P | Pfam PF02285 |
| 20 | Cox8c | COX8C_HUMAN | Cytochrome c oxidase subunit 8C, mitochondrial | Pfam PF02285 |
| Assembly subunits[7][8][9] | ||||
| 1 | Coa1 | COA1_HUMAN | Cytochrome c oxidase assembly factor 1 homolog | Pfam PF08695 |
| 2 | Coa3 | COA3_HUMAN | Cytochrome c oxidase assembly factor 3 homolog, mitochondrial | Pfam PF09813 |
| 3 | Coa4 | COA4_HUMAN | Cytochrome c oxidase assembly factor 4 homolog, mitochondrial | Pfam PF06747 |
| 4 | Coa5 | COA5_HUMAN | Cytochrome c oxidase assembly factor 5 | Pfam PF10203 |
| 5 | Coa6 | COA6_HUMAN | Cytochrome c oxidase assembly factor 6 homolog | Pfam PF02297 |
| 6 | Coa7 | COA7_HUMAN | Cytochrome c oxidase assembly factor 7, | Pfam PF08238 |
| 7 | Cox11 | COX11_HUMAN | Cytochrome c oxidase assembly protein COX11 mitochondrial | Pfam PF04442 |
| 8 | Cox14 | COX14_HUMAN | Cytochrome c oxidase assembly protein | Pfam PF14880 |
| 9 | Cox15 | COX15_HUMAN | Cytochrome c oxidase assembly protein COX15 homolog | Pfam PF02628 |
| 10 | Cox16 | COX16_HUMAN | Cytochrome c oxidase assembly protein COX16 homolog mitochondrial | Pfam PF14138 |
| 11 | Cox17 | COX17_HUMAN | Cytochrome c oxidase copper chaperone | Pfam PF05051 |
| 12 | Cox18[10] | COX18_HUMAN | Mitochondrial inner membrane protein (Cytochrome c oxidase assembly protein 18) | Pfam PF02096 |
| 13 | Cox19 | COX19_HUMAN | Cytochrome c oxidase assembly protein | Pfam PF06747 |
| 14 | Cox20 | COX20_HUMAN | Cytochrome c oxidase protein 20 homolog | Pfam PF12597 |
Assembly
COX assembly in
Cofactors, including hemes, are inserted into subunits I & II. The two heme molecules reside in subunit I, helping with transport to subunit II where two copper molecules aid with the continued transfer of electrons.[13] Subunits I and IV initiate assembly. Different subunits may associate to form sub-complex intermediates that later bind to other subunits to form the COX complex.[11] In post-assembly modifications, COX will form a homodimer. This is required for activity. Dimers are connected by a cardiolipin molecule,[11][14][15] which has been found to play a key role in stabilization of the holoenzyme complex. The dissociation of subunits VIIa and III in conjunction with the removal of cardiolipin results in total loss of enzyme activity.[15] Subunits encoded in the nuclear genome are known to play a role in enzyme dimerization and stability. Mutations to these subunits eliminate COX function.[11]
Assembly is known to occur in at least three distinct rate-determining steps. The products of these steps have been found, though specific subunit compositions have not been determined.[11]
Synthesis and assembly of COX subunits I, II, and III are facilitated by translational activators, which interact with the 5’ untranslated regions of mitochondrial mRNA transcripts. Translational activators are encoded in the nucleus. They can operate through either direct or indirect interaction with other components of translation machinery, but exact molecular mechanisms are unclear due to difficulties associated with synthesizing translation machinery in-vitro.[16][17] Though the interactions between subunits I, II, and III encoded within the mitochondrial genome make a lesser contribution to enzyme stability than interactions between bigenomic subunits, these subunits are more conserved, indicating potential unexplored roles for enzyme activity.[18]
Biochemistry
doi:10.1038/s41467-021-27174-y . (December 2021) |
The overall reaction is
- 4 Fe2+ – cytochrome c + 4 H+ + O2 → 4 Fe3+ – cytochrome c + 2 H2O ΔGo' = - 218 kJ/mol, Eo' = +565 mV
Two electrons are passed from two cytochrome c's, through the CuA and cytochrome a sites to the cytochrome a3–CuB binuclear center, reducing the metals to the Fe2+ form and Cu+. The hydroxide ligand is protonated and lost as water, creating a void between the metals that is filled by O2. The oxygen is rapidly reduced, with two electrons coming from the Fe2+-cytochrome a3, which is converted to the ferryl oxo form (Fe4+=O). The oxygen atom close to CuB picks up one electron from Cu+, and a second electron and a proton from the
Inhibition
COX exists in three conformational states: fully oxidized (pulsed), partially reduced, and fully reduced. Each inhibitor has a high affinity to a different state. In the pulsed state, both the heme a3 and the CuB nuclear centers are oxidized; this is the conformation of the enzyme that has the highest activity. A two-electron reduction initiates a conformational change that allows oxygen to bind at the active site to the partially-reduced enzyme. Four electrons bind to COX to fully reduce the enzyme. Its fully reduced state, which consists of a reduced Fe2+ at the cytochrome a3 heme group and a reduced CuB+ binuclear center, is considered the inactive or resting state of the enzyme.[19]
Cyanide is a non-competitive inhibitor for COX,[22][23] binding with high affinity to the partially-reduced state of the enzyme and hindering further reduction of the enzyme. In the pulsed state, cyanide binds slowly, but with high affinity. The ligand is posited to electrostatically stabilize both metals at once by positioning itself between them. A high nitric oxide concentration, such as one added exogenously to the enzyme, reverses cyanide inhibition of COX.[24]
Nitric oxide can reversibly[25] bind to either metal ion in the binuclear center to be oxidized to nitrite. NO and CN− will compete with oxygen to bind at the site, reducing the rate of cellular respiration. Endogenous NO, however, which is produced at lower levels, augments CN− inhibition. Higher levels of NO, which correlate with the existence of more enzyme in the reduced state, lead to a greater inhibition of cyanide.[19] At these basal concentrations, NO inhibition of Complex IV is known to have beneficial effects, such as increasing oxygen levels in blood vessel tissues. The inability of the enzyme to reduce oxygen to water results in a buildup of oxygen, which can diffuse deeper into surrounding tissues.[25] NO inhibition of Complex IV has a larger effect at lower oxygen concentrations, increasing its utility as a vasodilator in tissues of need.[25]
Hydrogen sulfide will bind COX in a noncompetitive fashion at a regulatory site on the enzyme, similar to carbon monoxide. Sulfide has the highest affinity to either the pulsed or partially reduced states of the enzyme, and is capable of partially reducing the enzyme at the heme a3 center. It is unclear whether endogenous H2S levels are sufficient to inhibit the enzyme. There is no interaction between hydrogen sulfide and the fully reduced conformation of COX.[21]
Extramitochondrial and subcellular localizations
Cytochrome c oxidase has 3 subunits which are encoded by
Genetic defects and disorders
Defects involving genetic mutations altering cytochrome c oxidase (COX) functionality or structure can result in severe, often fatal
The vast majority of COX disorders are linked to mutations in nuclear-encoded proteins referred to as assembly factors, or assembly proteins. These assembly factors contribute to COX structure and functionality, and are involved in several essential processes, including transcription and translation of mitochondrion-encoded subunits, processing of preproteins and membrane insertion, and cofactor biosynthesis and incorporation.[32]
Currently, mutations have been identified in seven COX assembly factors:
Histochemistry
The increased reliance of neurons on oxidative phosphorylation for energy[33] facilitates the use of COX histochemistry in mapping regional brain metabolism in animals, since it establishes a direct and positive correlation between enzyme activity and neuronal activity.[34] This can be seen in the correlation between COX enzyme amount and activity, which indicates the regulation of COX at the level of gene expression. COX distribution is inconsistent across different regions of the animal brain, but its pattern of its distribution is consistent across animals. This pattern has been observed in the monkey, mouse, and calf brain. One isozyme of COX has been consistently detected in histochemical analysis of the brain.[35] Such brain mapping has been accomplished in spontaneous mutant mice with cerebellar disease such as reeler[36] and a transgenic model of Alzheimer's disease.[37] This technique has also been used to map learning activity in the animal brain.[38]
Additional images
-
ETC
-
Complex IV
See also
- Cytochrome c oxidase subunit I
- Cytochrome c oxidase subunit II
- Cytochrome c oxidase subunit III
- Heme a
References
- PMID 8013452.
- PMID 22902835.
- S2CID 27210776.
- ^ ISBN 978-0-470-57095-1.
- S2CID 4380033.
- PMID 21211513.
- PMID 22356826.
- PMID 23260140.
- PMID 24333015.
- PMID 16911509.
- ^ PMID 16760263.
- PMID 10854440.
- ^ Crofts A (1996). "Cytochrome oxidase: Complex IV". University of Illinois at Urbana-Champaign. Archived from the original on 2018-01-23. Retrieved 2018-01-28.
- PMID 16199211.
- ^ PMID 26284624.
- PMID 22450032.
- PMID 21958598.
- PMID 25359921.
- ^ PMID 17906319.
- PMID 12969439.
- ^ S2CID 11554252.
- ISBN 9780174387329. Archivedfrom the original on 2022-02-24. Retrieved 2020-10-25.
- ISBN 9780174480198. Archivedfrom the original on 2022-02-24. Retrieved 2020-10-25.
- PMID 6098268.
- ^ PMID 19461104.
- PMID 9363790.
- ^ S2CID 24440427.
- )
- ^ PMID 10494626.
- PMID 10322429.
- (PDF) from the original on 2011-07-18. Retrieved 2010-11-17.
- PMID 17215873.
- PMID 24032355.
- S2CID 42996304.
- PMID 2555458.
- S2CID 45787322.
- S2CID 9366458.
- S2CID 24271956.
External links
- The Cytochrome Oxidase home page at Rice University
- Interactive Molecular model of cytochrome c oxidase (Requires MDL Chime)
- UMich Orientation of Proteins in Membranes families/superfamily-4
- Cytochrome-c+Oxidase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)