DNA condensation

DNA condensation refers to the process of compacting
DNA diameter is about 2 nm, while the length of a stretched single molecule may be up to several dozens of centimetres depending on the organism. Many features of the DNA double helix contribute to its large stiffness, including the mechanical properties of the sugar-phosphate backbone, electrostatic repulsion between
In life
In viruses
In
In bacteria
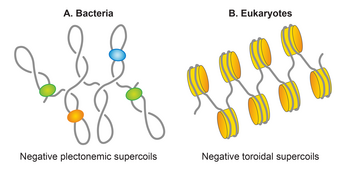
Bacterial DNA is packed with the help of
Sister chromosomes in the bacterium Escherichia coli are induced by stressful conditions to condense and undergo pairing.[6] Stress-induced condensation occurs by a non-random, zipper-like convergence of sister chromosomes. This convergence appears to depend on the ability of identical double-stranded DNA molecules to specifically identify each other, a process that culminates in the proximity of homologous sites along the paired chromosomes. Diverse stress conditions appear to prime bacteria to effectively cope with severe DNA damages such as double-strand breaks. The apposition of homologous sites associated with stress-induced chromosome condensation helps explain how repair of double-strand breaks and other damages occurs.[6]
In eukaryotes

Eukaryotic DNA with a typical length of dozens of centimeters should be orderly packed to be readily accessible inside the micrometer-size nucleus. In most eukaryotes, DNA is arranged in the cell nucleus with the help of histones. In this case, the basic level of DNA compaction is the nucleosome, where the double helix is wrapped around the histone octamer containing two copies of each
Chromosome scaffold has important role to hold the chromatin into compact chromosome. Chromosome scaffold is made of proteins including condensin, topoisomerase IIα and kinesin family member 4 (KIF4)[7]
In archaea
Depending on the organism, an archaeon may use a bacteria-like HU system or a eukaryote-like nucleosome system for packaging.[11]
In vitro
DNA condensation can be induced in vitro either by applying external force to bring the double helices together, or by inducing attractive interactions between the DNA segments. The former can be achieved e.g. with the help of the osmotic pressure exerted by crowding neutral polymers in the presence of monovalent salts. In this case, the forces pushing the double helices together are coming from entropic random collisions with the crowding polymers surrounding DNA condensates, and salt is required to neutralize DNA charges and decrease DNA-DNA repulsion. The second possibility can be realized by inducing attractive interactions between the DNA segments by multivalent cationic charged ligands (multivalent
Physics
Condensation of long double-helical DNAs is a sharp phase transition, which takes place within a narrow interval of condensing agent concentrations.[ref] Since the double helices come very closely to each other in the condensed phase, this leads to the restructuring of water molecules, which gives rise to the so-called hydration forces.[ref] To understand attraction between negatively charged DNA molecules, one also must account for correlations between counterions in the solution.[ref] DNA condensation by proteins can exhibit hysteresis, which can be explained using a modified Ising model.[12]
Role in gene regulation
Nowadays descriptions of gene regulation are based on the approximations of
See also
References
Further reading
- Gelbart W. M.; Bruinsma R.; Pincus P. A.; Parsegian V. A. (2000). "DNA-Inspired Electrostatics". Physics Today. 53 (9): 38. .
- Strey H. H.; Podgornik R.; Rau D. C.; Parsegian V. A. (1998). "DNA-DNA interactions". Current Opinion in Structural Biology. 8 (3): 309–313. PMID 9666326.
- Schiessel H (2003). "The physics of chromatin". J. Phys.: Condens. Matter. 15 (19): R699–R774. S2CID 250893202.
- Vijayanathan V.; Thomas T.; Thomas T. J. (2002). "DNA nanoparticles and development of DNA delivery vehicles for gene therapy". Biochemistry. 41 (48): 14085–14094. PMID 12450371.
- Yoshikawa K (2001). "Controlling the higher-order structure of giant DNA molecules". Advanced Drug Delivery Reviews. 52 (3): 235–244. PMID 11718948.
- Hud N. V.; Vilfan I. D. (2005). "Toroidal DNA condensates: unraveling the fine structure and the role of nucleation in determining size". Annu Rev Biophys Biomol Struct. 34: 295–318. PMID 15869392.
- Yoshikawa, K., and Y. Yoshikawa. 2002. Compaction and condensation of DNA. In Pharmaceutical perspectives of nucleic acid-based therapeutics. R. I. Mahato, and S. W. Kim, editors. Taylor & Francis. 137–163.
