DNA microarray
A DNA microarray (also commonly known as
Principle
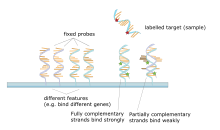
The core principle behind microarrays is hybridization between two DNA strands, the property of

Uses and types

Many types of arrays exist and the broadest distinction is whether they are spatially arranged on a surface or on coded beads:
- The traditional solid-phase array is a collection of orderly microscopic "spots", called features, each with thousands of identical and specific probes attached to a solid surface, such as glass, plastic or silicon biochip (commonly known as a genome chip, DNA chip or gene array). Thousands of these features can be placed in known locations on a single DNA microarray.
- The alternative bead array is a collection of microscopic polystyrene beads, each with a specific probe and a ratio of two or more dyes, which do not interfere with the fluorescent dyes used on the target sequence.
DNA microarrays can be used to detect DNA (as in
Applications include:
| Application or technology | Synopsis |
|---|---|
| Gene expression profiling | In an pathogens or other organisms by comparing gene expression in infected to that in uninfected cells or tissues.[2]
|
| Comparative genomic hybridization | Assessing genome content in different cells or closely related organisms, as originally described by Patrick Brown, Jonathan Pollack, Ash Alizadeh and colleagues at Stanford.[3][4] |
| GeneID | Small microarrays to check IDs of organisms in food and feed (like pathogens for disease detection, mostly combining PCR and microarray technology.
|
| Chromatin immunoprecipitation on Chip | DNA sequences bound to a particular protein can be isolated by transcription landscape .
|
| DamID | Analogously to DNA adenine methyltransferase .
|
| SNP detection | Identifying forensic analysis, measuring predisposition to disease, identifying drug-candidates, evaluating germline mutations in individuals or somatic mutations in cancers, assessing loss of heterozygosity, or genetic linkage analysis.
|
| Alternative splicing detection | An exon junction array design uses probes specific to the expected or potential splice sites of predicted exons for a gene. It is of intermediate density, or coverage, to a typical gene expression array (with 1–3 probes per gene) and a genomic tiling array (with hundreds or thousands of probes per gene). It is used to assay the expression of alternative splice forms of a gene. Exon arrays have a different design, employing probes designed to detect each individual exon for known or predicted genes, and can be used for detecting different splicing isoforms. |
| Fusion genes microarray | A Fusion gene microarray can detect fusion transcripts, e.g. from cancer specimens. The principle behind this is building on the alternative splicing microarrays. The oligo design strategy enables combined measurements of chimeric transcript junctions with exon-wise measurements of individual fusion partners. |
| Tiling array | Genome tiling arrays consist of overlapping probes designed to densely represent a genomic region of interest, sometimes as large as an entire human chromosome. The purpose is to empirically detect expression of transcripts or alternatively spliced forms which may not have been previously known or predicted.
|
| Double-stranded B-DNA microarrays | Right-handed double-stranded B-DNA microarrays can be used to characterize novel drugs and biologicals that can be employed to bind specific regions of immobilized, intact, double-stranded DNA. This approach can be used to inhibit gene expression.[6][7] They also allow for characterization of their structure under different environmental conditions. |
| Double-stranded Z-DNA microarrays | Left-handed double-stranded Z-DNA microarrays can be used to identify short sequences of the alternative Z-DNA structure located within longer stretches of right-handed B-DNA genes (e.g., transcriptional enhancement, recombination, RNA editing).[6][7] The microarrays also allow for characterization of their structure under different environmental conditions. |
| Multi-stranded DNA microarrays (triplex-DNA microarrays and quadruplex-DNA microarrays) | Multi-stranded DNA and RNA microarrays can be used to identify novel drugs that bind to these multi-stranded nucleic acid sequences. This approach can be used to discover new drugs and biologicals that have the ability to inhibit gene expression.[6][7][8][9] These microarrays also allow for characterization of their structure under different environmental conditions. |
Specialised arrays tailored to particular crops are becoming increasingly popular in molecular breeding applications. In the future they could be used to screen seedlings at early stages to lower the number of unneeded seedlings tried out in breeding operations.[10]
Fabrication
Microarrays can be manufactured in different ways, depending on the number of probes under examination, costs, customization requirements, and the type of scientific question being asked. Arrays from commercial vendors may have as few as 10 probes or as many as 5 million or more micrometre-scale probes.
Spotted vs. in situ synthesised arrays
Microarrays can be fabricated using a variety of technologies, including printing with fine-pointed pins onto glass slides, photolithography using pre-made masks, photolithography using dynamic micromirror devices, ink-jet printing,[11][12] or electrochemistry on microelectrode arrays.
In spotted microarrays, the probes are
In oligonucleotide microarrays, the probes are short sequences designed to match parts of the sequence of known or predicted
Two-channel vs. one-channel detection
Two-color microarrays or two-channel microarrays are typically
Oligonucleotide microarrays often carry control probes designed to hybridize with
In single-channel microarrays or one-color microarrays, the arrays provide intensity data for each probe or probe set indicating a relative level of hybridization with the labeled target. However, they do not truly indicate abundance levels of a gene but rather relative abundance when compared to other samples or conditions when processed in the same experiment. Each RNA molecule encounters protocol and batch-specific bias during amplification, labeling, and hybridization phases of the experiment making comparisons between genes for the same microarray uninformative. The comparison of two conditions for the same gene requires two separate single-dye hybridizations. Several popular single-channel systems are the Affymetrix "Gene Chip", Illumina "Bead Chip", Agilent single-channel arrays, the Applied Microarrays "CodeLink" arrays, and the Eppendorf "DualChip & Silverquant". One strength of the single-dye system lies in the fact that an aberrant sample cannot affect the raw data derived from other samples, because each array chip is exposed to only one sample (as opposed to a two-color system in which a single low-quality sample may drastically impinge on overall data precision even if the other sample was of high quality). Another benefit is that data are more easily compared to arrays from different experiments as long as batch effects have been accounted for.
One channel microarray may be the only choice in some situations. Suppose samples need to be compared: then the number of experiments required using the two channel arrays quickly becomes unfeasible, unless a sample is used as a reference.
| number of samples | one-channel microarray | two channel microarray |
two channel microarray (with reference) |
|---|---|---|---|
| 1 | 1 | 1 | 1 |
| 2 | 2 | 1 | 1 |
| 3 | 3 | 3 | 2 |
| 4 | 4 | 6 | 3 |
A typical protocol

This is an example of a DNA microarray experiment which includes details for a particular case to better explain DNA microarray experiments, while listing modifications for RNA or other alternative experiments.
- The two samples to be compared (pairwise comparison) are grown/acquired. In this example treated sample (control).
- The Guanidinium thiocyanate-phenol-chloroform extraction (e.g. Trizol) which isolates most RNA (whereas column methods have a cut off of 200 nucleotides) and if done correctly has a better purity.
- The purified RNA is analysed for quality (by μg, although the required amount varies by microarray platform), the experiment can proceed.
- The labeled product is generated via rRNA). miRNAmicroarrays ligate an oligonucleotide to the purified small RNA (isolated with a fractionator), which is then reverse transcribed and amplified.
- The label is added either during the reverse transcription step, or following amplification if it is performed. The cDNAis antisense and the microarray probe is sense, except in the case of negative controls.
- The label is typically radiolabels.
- The labeling can be direct (not used) or indirect (requires a coupling stage). For two-channel arrays, the coupling stage occurs before hybridization, using NHS amino-reactive dyes (such as cyanine dyes); for single-channel arrays, the coupling stage occurs after hybridization, using biotin and labeled streptavidin. The modified nucleotides (usually in a ratio of 1 aaUTP: 4 TTP (thymidine triphosphate)) are added enzymatically in a low ratio to normal nucleotides, typically resulting in 1 every 60 bases. The aaDNA is then purified with a column (using a phosphate buffer solution, as Triscontains amine groups). The aminoallyl group is an amine group on a long linker attached to the nucleobase, which reacts with a reactive dye.
- A form of replicate known as a dye flip can be performed to control for dye aminoallyl-UTP is present in the reverse-transcribed mixture.
- A form of replicate known as a dye flip can be performed to control for dye
- The label is added either during the reverse transcription step, or following amplification if it is performed. The
- The labeled samples are then mixed with a proprietary PolyA, or PolyT), Denhardt's solution, or formamine.
- The mixture is denatured and added to the pinholes of the microarray. The holes are sealed and the microarray hybridized, either in a hyb oven, where the microarray is mixed by rotation, or in a mixer, where the microarray is mixed by alternating pressure at the pinholes.
- After an overnight hybridization, all nonspecific binding is washed off (SDS and SSC).
- The microarray is dried and scanned by a machine that uses a laser to excite the dye and measures the emission levels with a detector.
- The image is gridded with a template and the intensities of each feature (composed of several pixels) is quantified.
- The raw data is normalized; the simplest normalization method is to subtract background intensity and scale so that the total intensities of the features of the two channels are equal, or to use the intensity of a reference gene to calculate the z-ratio, loess and lowess regressionand RMA (robust multichip analysis) for Affymetrix chips (single-channel, silicon chip, in situ synthesized short oligonucleotides).
Microarrays and bioinformatics

The advent of inexpensive microarray experiments created several specific bioinformatics challenges:
Experimental design
Due to the biological complexity of gene expression, the considerations of experimental design that are discussed in the
There are three main elements to consider when designing a microarray experiment. First, replication of the biological samples is essential for drawing conclusions from the experiment. Second, technical replicates (e.g. two RNA samples obtained from each experimental unit) may help to quantitate precision. The biological replicates include independent RNA extractions. Technical replicates may be two aliquots of the same extraction. Third, spots of each cDNA clone or oligonucleotide are present as replicates (at least duplicates) on the microarray slide, to provide a measure of technical precision in each hybridization. It is critical that information about the sample preparation and handling is discussed, in order to help identify the independent units in the experiment and to avoid inflated estimates of statistical significance.[20]
Standardization
Microarray data is difficult to exchange due to the lack of standardization in platform fabrication, assay protocols, and analysis methods. This presents an
For example, the "Minimum Information About a Microarray Experiment" (
Data analysis

Microarray data sets are commonly very large, and analytical precision is influenced by a number of variables. Statistical challenges include taking into account effects of background noise and appropriate normalization of the data. Normalization methods may be suited to specific platforms and, in the case of commercial platforms, the analysis may be proprietary.[22] Algorithms that affect statistical analysis include:
- Image analysis: gridding, spot recognition of the scanned image (segmentation algorithm), removal or marking of poor-quality and low-intensity features (called flagging).
- Data processing: background subtraction (based on global or local background), determination of spot intensities and intensity ratios, visualisation of data (e.g. see MA plot), and log-transformation of ratios, global or local normalization of intensity ratios, and segmentation into different copy number regions using step detection algorithms.[23]
- Class discovery analysis: This analytic approach, sometimes called unsupervised classification or knowledge discovery, tries to identify whether microarrays (objects, patients, mice, etc.) or genes cluster together in groups. Identifying naturally existing groups of objects (microarrays or genes) which cluster together can enable the discovery of new groups that otherwise were not previously known to exist. During knowledge discovery analysis, various unsupervised classification techniques can be employed with DNA microarray data to identify novel clusters (classes) of arrays.[24] This type of approach is not hypothesis-driven, but rather is based on iterative pattern recognition or statistical learning methods to find an "optimal" number of clusters in the data. Examples of unsupervised analyses methods include self-organizing maps, neural gas, k-means cluster analyses,[25] hierarchical cluster analysis, Genomic Signal Processing based clustering and model-based cluster analysis. For some of these methods the user also has to define a distance measure between pairs of objects. Although the Pearson correlation coefficient is usually employed, several other measures have been proposed and evaluated in the literature.[26] The input data used in class discovery analyses are commonly based on lists of genes having high informativeness (low noise) based on low values of the coefficient of variation or high values of Shannon entropy, etc. The determination of the most likely or optimal number of clusters obtained from an unsupervised analysis is called cluster validity. Some commonly used metrics for cluster validity are the silhouette index, Davies-Bouldin index,[27] Dunn's index, or Hubert's statistic.
- Class prediction analysis: This approach, called supervised classification, establishes the basis for developing a predictive model into which future unknown test objects can be input in order to predict the most likely class membership of the test objects. Supervised analysisant colony optimization. Input data for class prediction are usually based on filtered lists of genes which are predictive of class, determined using classical hypothesis tests (next section), Gini diversity index, or information gain (entropy).
- Hypothesis-driven statistical analysis: Identification of statistically significant changes in gene expression are commonly identified using the multiple comparisons[29] or cluster analysis.[30] These methods assess statistical power based on the variation present in the data and the number of experimental replicates, and can help minimize Type I and type II errors in the analyses.[31]
- Dimensional reduction: Analysts often reduce the number of dimensions (genes) prior to data analysis.[24] This may involve linear approaches such as principal components analysis (PCA), or non-linear manifold learning (distance metric learning) using kernel PCA, diffusion maps, Laplacian eigenmaps, local linear embedding, locally preserving projections, and Sammon's mapping.
- Network-based methods: Statistical methods that take the underlying structure of gene networks into account, representing either associative or causative interactions or dependencies among gene products.[32] Weighted gene co-expression network analysis is widely used for identifying co-expression modules and intramodular hub genes. Modules may corresponds to cell types or pathways. Highly connected intramodular hubs best represent their respective modules.
Microarray data may require further processing aimed at reducing the dimensionality of the data to aid comprehension and more focused analysis.[33] Other methods permit analysis of data consisting of a low number of biological or technical replicates; for example, the Local Pooled Error (LPE) test pools standard deviations of genes with similar expression levels in an effort to compensate for insufficient replication.[34]
Annotation
The relation between a probe and the
Data warehousing
Microarray data was found to be more useful when compared to other similar datasets. The sheer volume of data, specialized formats (such as
Alternative technologies
Advances in massively parallel sequencing has led to the development of RNA-Seq technology, that enables a whole transcriptome shotgun approach to characterize and quantify gene expression.[36][37] Unlike microarrays, which need a reference genome and transcriptome to be available before the microarray itself can be designed, RNA-Seq can also be used for new model organisms whose genome has not been sequenced yet.[37]
Glossary
- An array or slide is a collection of featuresspatially arranged in a two dimensional grid, arranged in columns and rows.
- Block or subarray: a group of spots, typically made in one print round; several subarrays/ blocks form an array.
- Case/control: an experimental design paradigm especially suited to the two-colour array system, in which a condition chosen as control (such as healthy tissue or state) is compared to an altered condition (such as a diseased tissue or state).
- Channel: the fluorescence output recorded in the scanner for an individual fluorophore and can even be ultraviolet.
- Dye flip or dye swap or fluor reversal: reciprocal labelling of DNA targets with the two dyes to account for dye bias in experiments.
- Scanner: an instrument used to detect and quantify the intensity of fluorescence of spots on a microarray slide, by selectively exciting fluorophores with a filter (optics) photomultipliersystem.
- Spot or feature: a small area on an array slide that contains picomoles of specific DNA samples.
- For other relevant terms see:
- Glossary of gene expression terms
- Protocol (natural sciences)
See also
- Transcriptomics technologies
- MAGIChip
- Microarray analysis techniques
- Microarray databases
- fluorophoreswith microarrays
- Gene chip analysis
- Significance analysis of microarrays
- Methylation specific oligonucleotide microarray
- lab-on-chip
- Pathogenomics
- Phenotype microarray
- Systems biology
- Whole genome sequencing
References
- PMID 6198132.
- PMID 18381269.
- S2CID 997032.
- PMID 15470115.
- S2CID 41718227.
- ^ PMID 19450135.
- ^ S2CID 41959328.
- PMID 21501652.
- PMID 26350216.
- S2CID 33780984.
- ^ J Biochem Biophys Methods. 2000 Mar 16;42(3):105–10. DNA-printing: utilization of a standard inkjet printer for the transfer of nucleic acids to solid supports. Goldmann T, Gonzalez JS.
- PMID 15287980.
- S2CID 195368323.
- PMID 8197176.
- PMID 12421762.
- PMID 8796352.
- PMID 17698492.
- ISSN 2002-4436.
- ISSN 1535-6108.
- S2CID 15412245. Archived from the original(PDF) on 8 May 2005. Retrieved 12 December 2013.
- ^ NCTR Center for Toxicoinformatics – MAQC Project
- ^ "Prosigna | Prosigna algorithm". prosigna.com. Retrieved 22 June 2017.
- PMID 22003312.
- ^ ISBN 978-0-470-17081-6.
- ^ De Souto M et al. (2008) Clustering cancer gene expression data: a comparative study, BMC Bioinformatics, 9(497).
- PMID 24564555.
- ^ Bolshakova N, Azuaje F (2003) Cluster validation techniques for genome expression data, Signal Processing, Vol. 83, pp. 825–833.
- PMID 15797905.
- ^ Yuk Fai Leung and Duccio Cavalieri, Fundamentals of cDNA microarray data analysis. Trends in Genetics Vol.19 No.11 November 2003.
- PMID 17397530.
- PMID 15533245.
- ISBN 978-3-527-31822-3.
- S2CID 16248921.
- PMID 14555628.
- PMID 19923232.
- S2CID 205418589.
- ^ PMID 19015660.
External links
- Gene Expression at Curlie
- Micro Scale Products and Services for Biochemistry and Molecular Biology at Curlie
- Products and Services for Gene Expression at Curlie
- Online Services for Gene Expression Analysis at Curlie
- Microarray Animation 1Lec.com
- PLoS Biology Primer: Microarray Analysis
- Rundown of microarray technology
- ArrayMining.net – a free web-server for online microarray analysis
- Microarray – How does it work?
- PNAS Commentary: Discovery of Principles of Nature from Mathematical Modeling of DNA Microarray Data
- DNA microarray virtual experiment




