Dihydrolipoamide dehydrogenase
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 7: 107.89 – 107.93 Mb | Chr 12: 31.38 – 31.4 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Dihydrolipoamide dehydrogenase (DLD), also known as dihydrolipoyl dehydrogenase, mitochondrial, is an enzyme that in humans is encoded by the DLD gene.[5][6][7][8] DLD is a flavoprotein enzyme that oxidizes dihydrolipoamide to lipoamide.
Dihydrolipoamide dehydrogenase (DLD) is a mitochondrial enzyme that plays a vital role in energy metabolism in eukaryotes. This enzyme is required for the complete reaction of at least five different multi-enzyme complexes.
Structure
The protein encoded by the DLD gene comes together with another protein to form a dimer in the
Function
The DLD homodimer functions as the E3 component of the
Moonlighting function
Certain DLD mutations can simultaneously induce the loss of a primary metabolic activity and the gain of a moonlighting proteolytic activity. The moonlighting proteolytic activity of DLD is revealed by conditions that destabilize the DLD homodimer and decrease its DLD activity.[9] Acidification of the mitochondrial matrix, as a result of ischemia-reperfusion injury, can disrupt the quaternary structure of DLD leading to decreased dehydrogenase activity and increased diaphorase activity.[20] The moonlighting proteolytic activity of DLD could also arise under pathological conditions. Proteolytic activity can further complicate the reduction in energy metabolism and an increase in oxidative damage as a result of decreased DLD activity and an increase in diaphorase activity respectively.[19] With its proteolytic function, DLD removes a functionally vital domain from the N-terminus of frataxin, a mitochondrial protein involved in iron metabolism and antioxidant protection.[21][22]
Clinical significance
In humans, mutations in DLD are linked to a severe disorder of infancy with failure to thrive, hypotonia, and metabolic acidosis. [23] DLD deficiency manifests itself in a great degree of variability, which has been attributed to varying effects of different DLD mutations on the stability of the protein and its ability to dimerize or interact with other components of the three α-ketoacid dehydrogenase complexes.[23] With its proteolytic function, DLD causes a deficiency in frataxin, which leads to the neurodegenerative and cardiac disease, Friedreich's ataxia.[24]
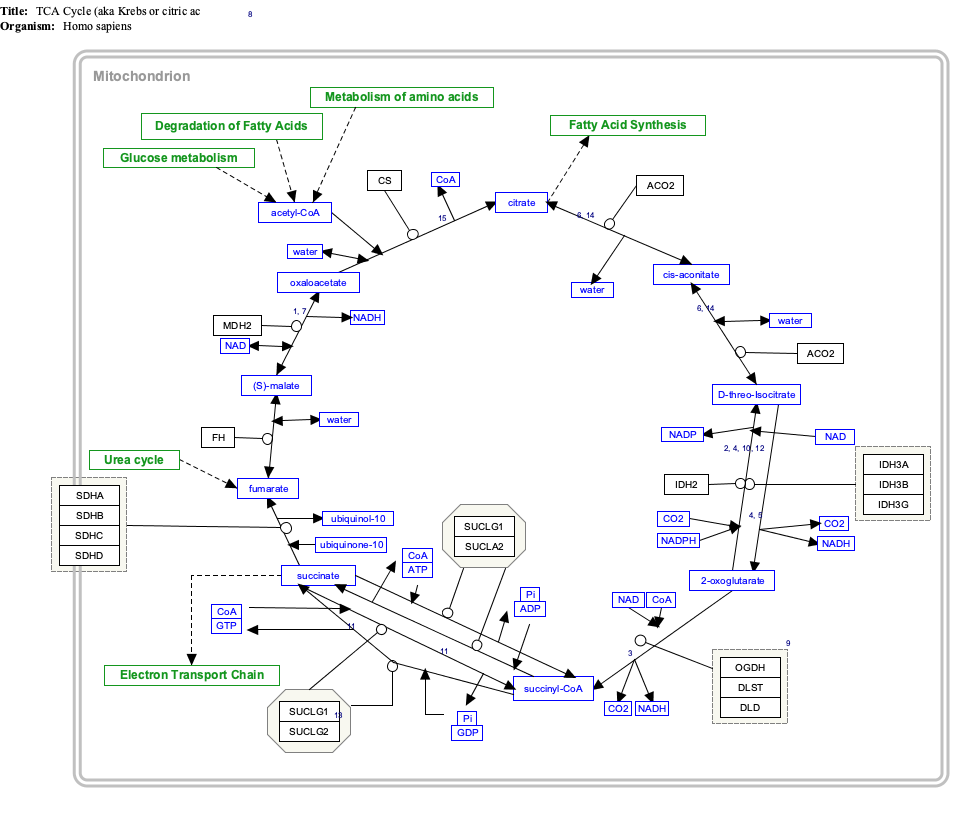
Interactive pathway map
| Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
TCACycle_WP78 edit
|
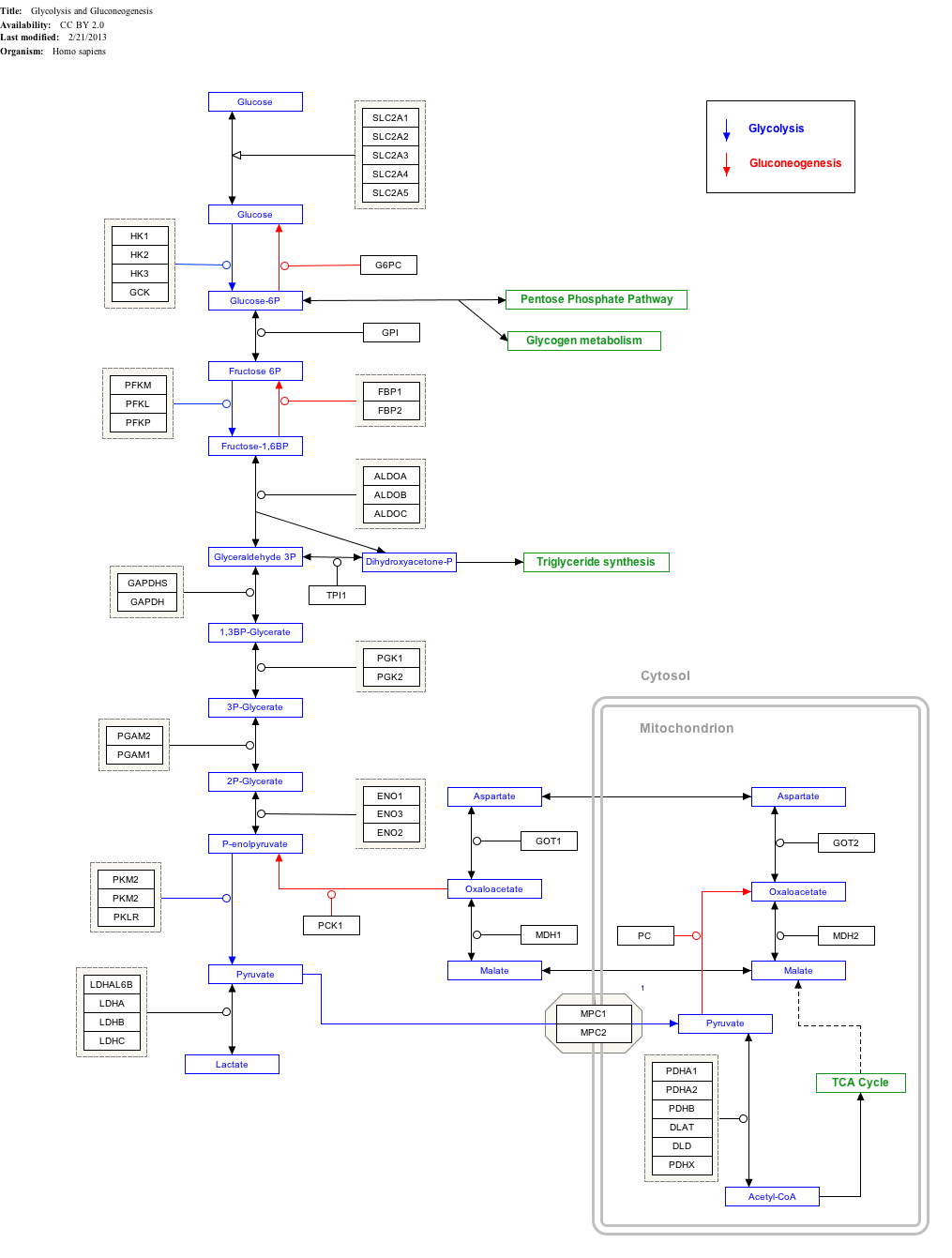
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
Glycolysis and Gluconeogenesis edit
|
Enzyme regulation
This protein may use the morpheein model of allosteric regulation.[25]
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000091140 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000020664 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: dihydrolipoamide dehydrogenase".
- PMID 3693355.
- PMID 3278312.
- PMID 2055113.
- ^ PMID 17404228.
- PMID 16263718.
- PMID 15826505.
- PMID 17960497.
- PMID 15946682.
- PMID 2643922.
- PMID 12963736.
- PMID 25202086.
- S2CID 20180110.
- S2CID 34370150.
- ^ PMID 11231302.
- PMID 15710613.
- PMID 16469498.
- PMID 15641778.
- ^ PMID 25032271.
- S2CID 38777180.
- PMID 22182754.
Further reading
- Silverberg MS, Cho JH, Rioux JD, McGovern DP, Wu J, Annese V, et al. (February 2009). "Ulcerative colitis-risk loci on chromosomes 1p36 and 12q15 found by genome-wide association study". Nature Genetics. 41 (2): 216–20. PMID 19122664.
- Brautigam CA, Chuang JL, Tomchick DR, Machius M, Chuang DT (July 2005). "Crystal structure of human dihydrolipoamide dehydrogenase: NAD+/NADH binding and the structural basis of disease-causing mutations". Journal of Molecular Biology. 350 (3): 543–52. PMID 15946682.
- Reed LJ, Hackert ML (June 1990). "Structure-function relationships in dihydrolipoamide acyltransferases". The Journal of Biological Chemistry. 265 (16): 8971–4. PMID 2188967.
- Ciszak EM, Makal A, Hong YS, Vettaikkorumakankauv AK, Korotchkina LG, Patel MS (January 2006). "How dihydrolipoamide dehydrogenase-binding protein binds dihydrolipoamide dehydrogenase in the human pyruvate dehydrogenase complex". The Journal of Biological Chemistry. 281 (1): 648–55. PMID 16263718.
- Asano K, Matsushita T, Umeno J, Hosono N, Takahashi A, Kawaguchi T, et al. (December 2009). "A genome-wide association study identifies three new susceptibility loci for ulcerative colitis in the Japanese population". Nature Genetics. 41 (12): 1325–9. S2CID 20507558.
- Odièvre MH, Chretien D, Munnich A, Robinson BH, Dumoulin R, Masmoudi S, et al. (March 2005). "A novel mutation in the dihydrolipoamide dehydrogenase E3 subunit gene (DLD) resulting in an atypical form of alpha-ketoglutarate dehydrogenase deficiency". Human Mutation. 25 (3): 323–4. S2CID 19929944.
- Brautigam CA, Wynn RM, Chuang JL, Machius M, Tomchick DR, Chuang DT (March 2006). "Structural insight into interactions between dihydrolipoamide dehydrogenase (E3) and E3 binding protein of human pyruvate dehydrogenase complex". Structure. 14 (3): 611–21. PMID 16442803.
- Kim H (March 2006). "Activity of human dihydrolipoamide dehydrogenase is largely reduced by mutation at isoleucine-51 to alanine". Journal of Biochemistry and Molecular Biology. 39 (2): 223–7. PMID 16584639.
- Sugden MC, Holness MJ (May 2003). "Recent advances in mechanisms regulating glucose oxidation at the level of the pyruvate dehydrogenase complex by PDKs". American Journal of Physiology. Endocrinology and Metabolism. 284 (5): E855-62. PMID 12676647.
- Wang YC, Wang ST, Li C, Chen LY, Liu WH, Chen PR, et al. (January 2008). "The role of amino acids T148 and R281 in human dihydrolipoamide dehydrogenase". Journal of Biomedical Science. 15 (1): 37–46. PMID 17960497.
- Brown AM, Gordon D, Lee H, Caudy M, Hardy J, Haroutunian V, Blass JP (November 2004). "Association of the dihydrolipoamide dehydrogenase gene with Alzheimer's disease in an Ashkenazi Jewish population". American Journal of Medical Genetics. Part B, Neuropsychiatric Genetics. 131B (1): 60–6. S2CID 26098296.
- Wang YC, Wang ST, Li C, Liu WH, Chen PR, Chen LY, Liu TC (March 2007). "The role of N286 and D320 in the reaction mechanism of human dihydrolipoamide dehydrogenase (E3) center domain". Journal of Biomedical Science. 14 (2): 203–10. PMID 17171578.
- Foster LJ, Rudich A, Talior I, Patel N, Huang X, Furtado LM, et al. (January 2006). "Insulin-dependent interactions of proteins with GLUT4 revealed through stable isotope labeling by amino acids in cell culture (SILAC)". Journal of Proteome Research. 5 (1): 64–75. PMID 16396496.
- Kim H (March 2005). "Asparagine-473 residue is important to the efficient function of human dihydrolipoamide dehydrogenase". Journal of Biochemistry and Molecular Biology. 38 (2): 248–52. PMID 15826505.
- Hiromasa Y, Fujisawa T, Aso Y, Roche TE (February 2004). "Organization of the cores of the mammalian pyruvate dehydrogenase complex formed by E2 and E2 plus the E3-binding protein and their capacities to bind the E1 and E3 components". The Journal of Biological Chemistry. 279 (8): 6921–33. PMID 14638692.
- Wynn RM, Kato M, Machius M, Chuang JL, Li J, Tomchick DR, Chuang DT (December 2004). "Molecular mechanism for regulation of the human mitochondrial branched-chain alpha-ketoacid dehydrogenase complex by phosphorylation". Structure. 12 (12): 2185–96. PMID 15576032.
- Martins-de-Souza D, Gattaz WF, Schmitt A, Novello JC, Marangoni S, Turck CW, Dias-Neto E (April 2009). "Proteome analysis of schizophrenia patients Wernicke's area reveals an energy metabolism dysregulation". BMC Psychiatry. 9: 17. PMID 19405953.
External links
- Dihydrolipoamide+dehydrogenase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the United States National Library of Medicine, which is in the public domain.