Efflux pump
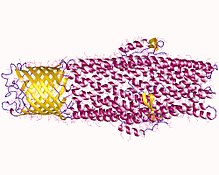

An efflux pump is an
Efflux systems function via an energy-dependent mechanism (active transport) to pump out unwanted toxic substances through specific efflux pumps. Some efflux systems are drug-specific, whereas others may accommodate multiple drugs with small multidrug resistance (SMR) transporters.[3][4]
Efflux pumps are
Bacterial
Bacterial efflux pumps are classified into five major superfamilies, based on their
- The major facilitator superfamily (MFS)[5]
- The ABC transporters[5]
- The small multidrug resistance family (SMR)[5]
- The resistance-nodulation-cell division superfamily (RND)[5]
- The multi antimicrobial extrusion protein family (MATE).[5]
Of these, only the ABC superfamily are primary transporters, the rest being
Structure
Efflux pumps generally consist of an outer membrane protein, a middle periplasmic protein, an inner membrane protein, and a transmembrane duct. The transmembrane duct is located in the outer membrane of the cell. The duct is also bound to two other proteins: a periplasmic membrane protein and an integral membrane transporter. The periplasmic membrane protein and the inner membrane protein of the system are coupled to control the opening and closing of the duct (channel). When a toxin binds to this inner membrane protein, the inner membrane proteins gives rise to a biochemical cascade that transmits signals to the periplasmic membrane protein and outer membrane protein to open the channel and move the toxin out of the cell. This mechanism uses an energy-dependent, protein-protein interaction that is generated by the transfer of the toxin for an H+ ion by the inner membrane transporter.[7] The fully assembled in vitro and in vivo structures of AcrAB-TolC pump have been solved by cryoEM and cryoET.[8][9]
Function
Although antibiotics are the most clinically important substrates of efflux systems, it is probable that most efflux pumps have other natural physiological functions. Examples include:
- The E. coli AcrAB efflux system, which has a physiologic role of pumping out bile acids and fatty acids to lower their toxicity.[10]
- The MFS family Ptr pump in Streptomyces pristinaespiralis appears to be an autoimmunity pump for this organism when it turns on production of pristinamycins I and II.[11]
- The AcrAB–TolC system in E. coli is suspected to have a role in the transport of the calcium-channel components in the E. coli membrane.[12]
- The MtrCDE system plays a protective role by providing resistance to faecal lipids in rectal isolates of Neisseria gonorrhoeae.[13]
- The AcrAB efflux system of
- The MexXY component of the MexXY-OprM multidrug efflux system of P. aeruginosa is inducible by antibiotics that target ribosomes via the PA5471 gene product.[15]
- Efflux pumps have also been shown to play a role in biofilm formation. However, the substrates for such pumps, and whether changes in their efflux activity affect biofilm formation directly or indirectly, remain to be determined.[16]
The ability of efflux systems to recognize a large number of compounds other than their natural substrates is probably because substrate recognition is based on
Impact on antimicrobial resistance
The impact of efflux mechanisms on antimicrobial resistance is large; this is usually attributed to the following:
- The transposons is also advantageous for the microorganisms as it allows for the easy spread of efflux genes between distant species.[17]
- Antibiotics can act as inducers and regulators of the expression of some efflux pumps.[15]
- Expression of several efflux pumps in a given bacterial species may lead to a broad spectrum of resistance when considering the shared substrates of some multi-drug efflux pumps, where one efflux pump may confer resistance to a wide range of antimicrobials.[2]
Eukaryotic
In eukaryotic cells, the existence of efflux pumps has been known since the discovery of P-glycoprotein in 1976 by Juliano and Ling.[18] Efflux pumps are one of the major causes of anticancer drug resistance in eukaryotic cells. They include monocarboxylate transporters (MCTs), multiple drug resistance proteins (MDRs)- also referred as P-glycoprotein, multidrug resistance-associated proteins (MRPs), peptide transporters (PEPTs), and Na+ phosphate transporters (NPTs). These transporters are distributed along particular portions of the renal proximal tubule, intestine, liver, blood–brain barrier, and other portions of the brain.
Inhibitors
Several trials are currently being conducted to develop drugs that can be co-administered with antibiotics to act as inhibitors for the efflux-mediated extrusion of antibiotics. As yet, no efflux inhibitor has been approved for therapeutic use, but some are being used to determine the prevalence of efflux pumps in clinical isolates and in
See also
- Antibiotic resistance
References
- PMID 31219077.
- ^ PMID 27681908.
- ISBN 978-3-319-39658-3.
- PMID 24878531.
- ^ PMID 24702006.
- PMID 17804667.
- PMID 23439914.
- PMID 28355133.
- PMID 31201302.
- PMID 8550435.
- PMID 19687245.
- PMID 24747401.
- PMID 10417654.
- PMID 24443882.
- ^ PMID 16484195.
- PMID 29506149.
- PMID 25788514.
- PMID 990323.
- ^ PMID 20645919.
- S2CID 205171789.
- PMID 20225250.
