Electron transport chain
An electron transport chain (ETC[1]) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. Many of the enzymes in the electron transport chain are embedded within the membrane.
The flow of electrons through the electron transport chain is an exergonic process. The energy from the redox reactions creates an electrochemical proton gradient that drives the synthesis of adenosine triphosphate (ATP). In aerobic respiration, the flow of electrons terminates with molecular oxygen as the final electron acceptor. In anaerobic respiration, other electron acceptors are used, such as sulfate.
In an electron transport chain, the redox reactions are driven by the difference in the
In
Mitochondrial electron transport chains
Most
Mitochondrial redox carriers
Energy associated with the transfer of electrons down the electron transport chain is used to pump protons from the
Four membrane-bound complexes have been identified in mitochondria. Each is an extremely complex
NADH, H+ → Complex I → Q → Complex III → cytochrome c → Complex IV → H2O
↑
Complex II
↑
Succinate
Complex I
In Complex I (NADH ubiquinone oxidoreductase, Type I NADH dehydrogenase, or mitochondrial complex I; EC 1.6.5.3), two electrons are removed from NADH and transferred to a lipid-soluble carrier, ubiquinone (Q). The reduced product, ubiquinol (QH2), freely diffuses within the membrane, and Complex I translocates four protons (H+) across the membrane, thus producing a proton gradient. Complex I is one of the main sites at which premature electron leakage to oxygen occurs, thus being one of the main sites of production of superoxide.[6]
The pathway of electrons is as follows:
Complex II
In
Complex III
In
When electron transfer is reduced (by a high membrane potential or respiratory inhibitors such as antimycin A), Complex III may leak electrons to molecular oxygen, resulting in superoxide formation.
This complex is inhibited by dimercaprol (British Anti-Lewisite, BAL), naphthoquinone and antimycin.
Complex IV
In
Coupling with oxidative phosphorylation
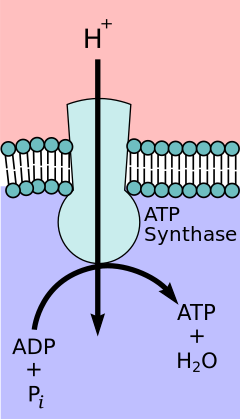
According to the
Coupling with oxidative phosphorylation is a key step for ATP production. However, in specific cases, uncoupling the two processes may be biologically useful. The uncoupling protein, thermogenin—present in the inner mitochondrial membrane of brown adipose tissue—provides for an alternative flow of protons back to the inner mitochondrial matrix. Thyroxine is also a natural uncoupler. This alternative flow results in thermogenesis rather than ATP production.[14]
Reverse electron flow
Prokaryotic electron transport chains
This section needs additional citations for verification. (December 2023) |
In eukaryotes, NADH is the most important electron donor. The associated electron transport chain is NADH → Complex I → Q → Complex III → cytochrome c → Complex IV → O2 where Complexes I, III and IV are proton pumps, while Q and cytochrome c are mobile electron carriers. The electron acceptor for this process is molecular oxygen.
In
) the situation is more complicated, because there are several different electron donors and several different electron acceptors. The generalized electron transport chain in bacteria is: Donor Donor Donor
↓ ↓ ↓
dehydrogenase → quinone → bc1 → cytochrome
↓ ↓
oxidase(reductase) oxidase(reductase)
↓ ↓
Acceptor Acceptor
Electrons can enter the chain at three levels: at the level of a dehydrogenase, at the level of the quinone pool, or at the level of a mobile cytochrome electron carrier. These levels correspond to successively more positive redox potentials, or to successively decreased potential differences relative to the terminal electron acceptor. In other words, they correspond to successively smaller Gibbs free energy changes for the overall redox reaction.
Individual bacteria use multiple electron transport chains, often simultaneously. Bacteria can use a number of different electron donors, a number of different dehydrogenases, a number of different oxidases and reductases, and a number of different electron acceptors. For example, E. coli (when growing aerobically using glucose and oxygen as an energy source) uses two different NADH dehydrogenases and two different quinol oxidases, for a total of four different electron transport chains operating simultaneously.
A common feature of all electron transport chains is the presence of a proton pump to create an electrochemical gradient over a membrane. Bacterial electron transport chains may contain as many as three proton pumps, like mitochondria, or they may contain two or at least one.
Electron donors
In the current biosphere, the most common electron donors are organic molecules. Organisms that use organic molecules as an electron source are called
Some prokaryotes can use inorganic matter as an electron source. Such an organism is called a
The use of inorganic electron donors such as hydrogen as an energy source is of particular interest in the study of evolution. This type of metabolism must logically have preceded the use of organic molecules and oxygen as an energy source.
Dehydrogenases: equivalants to complexes I and II
Bacteria can use several different electron donors. When organic matter is the electron source, the donor may be NADH or succinate, in which case electrons enter the electron transport chain via NADH dehydrogenase (similar to Complex I in mitochondria) or succinate dehydrogenase (similar to Complex II). Other dehydrogenases may be used to process different energy sources: formate dehydrogenase, lactate dehydrogenase, glyceraldehyde-3-phosphate dehydrogenase, H2 dehydrogenase (
Quinone carriers
Proton pumps
A
Some dehydrogenases are proton pumps, while others are not. Most oxidases and reductases are proton pumps, but some are not. Cytochrome bc1 is a proton pump found in many, but not all, bacteria (not in E. coli). As the name implies, bacterial bc1 is similar to mitochondrial bc1 (Complex III).
Cytochrome electron carriers
Some cytochromes are water-soluble carriers that shuttle electrons to and from large, immobile macromolecular structures imbedded in the membrane. The mobile cytochrome electron carrier in mitochondria is cytochrome c. Bacteria use a number of different mobile cytochrome electron carriers.
Other cytochromes are found within macromolecules such as Complex III and Complex IV. They also function as electron carriers, but in a very different, intramolecular, solid-state environment.
Electrons may enter an electron transport chain at the level of a mobile cytochrome or quinone carrier. For example, electrons from inorganic electron donors (nitrite, ferrous iron, electron transport chain) enter the electron transport chain at the cytochrome level. When electrons enter at a redox level greater than NADH, the electron transport chain must operate in reverse to produce this necessary, higher-energy molecule.
Electron acceptors and terminal oxidase/reductase
This section may require cleanup to meet Wikipedia's quality standards. The specific problem is: We talk as if oxidases are not also reductases, and as if reductases are not also oxidizing something. That's messed up. (December 2023) |
As there are a number of different electron donors (organic matter in organotrophs, inorganic matter in lithotrophs), there are a number of different electron acceptors, both organic and inorganic. As with other steps of the ETC, an enzyme is required to help with the process.
If oxygen is available, it is most often used as the terminal electron acceptor in aerobic bacteria and facultative anaerobes. An
Bacterial terminal oxidases can be split into classes according to the molecules act as terminal electron acceptors. Class I oxidases are cytochrome oxidases and use oxygen as the terminal electron acceptor. Class II oxidases are quinol oxidases and can use a variety of terminal electron acceptors. Both of these classes can be subdivided into categories based on what redox-active components they contain. E.g. Heme aa3 Class 1 terminal oxidases are much more efficient than Class 2 terminal oxidases.[2]
Mostly in anaerobic environments different electron acceptors are used, including nitrate, nitrite, ferric iron, sulfate, carbon dioxide, and small organic molecules such as fumarate. When bacteria grow in anaerobic environments, the terminal electron acceptor is reduced by an enzyme called a reductase. E. coli can use fumarate reductase, nitrate reductase, nitrite reductase, DMSO reductase, or trimethylamine-N-oxide reductase, depending on the availability of these acceptors in the environment.
Most terminal oxidases and reductases are inducible. They are synthesized by the organism as needed, in response to specific environmental conditions.
Photosynthetic
In oxidative phosphorylation, electrons are transferred from an electron donor such as NADH to an acceptor such as O2 through an electron transport chain, releasing energy. In photophosphorylation, the energy of sunlight is used to create a high-energy electron donor which can subsequently reduce oxidized components and couple to ATP synthesis via proton translocation by the electron transport chain.[9]
Photosynthetic electron transport chains, like the mitochondrial chain, can be considered as a special case of the bacterial systems. They use mobile, lipid-soluble quinone carriers (
See also
- Charge-transfer complex
- CoRR hypothesis
- Electron equivalent
- Hydrogen hypothesis
- Respirasome
- Electric bacteria
References
- ISBN 978-0-443-10281-3.
- ^ PMID 3052268.
- PMID 26124754. – This source shows four ETCs (Geobacter, Shewanella, Moorella , Acetobacterium) in figures 1 and 2.
- ISSN 0001-6101.
- PMID 28711444.
- ^ Lauren, Biochemistry, Johnson/Cole, 2010, pp 598-611
- ^ Garrett & Grisham, Biochemistry, Brooks/Cole, 2010, pp 598-611
- ISBN 978-1-305-57720-6.
- ^ OCLC 785100491.
- PMID 21874297.
- ^ ISBN 978-1-133-10629-6.
- S2CID 38896804.
- ^ Berg JM, Tymoczko JL, Stryer L (2002-01-01). "A Proton Gradient Powers the Synthesis of ATP".
{{cite journal}}: Cite journal requires|journal=(help) - PMID 14715917.
- ISBN 978-0-511-79046-1.
- PMID 27667687.
- ^ EC 1.3.5.1
- PMID 6387427.
Further reading
- Fenchel T, King GM, Blackburn TH (September 2006). Bacterial Biogeochemistry: The Ecophysiology of Mineral Cycling (2nd ed.). Elsevier. ISBN 978-0-12-103455-9.
- Lengeler JW (January 1999). Drews G; Schlegel HG (eds.). Biology of the Prokaryotes. Blackwell Science. ISBN 978-0-632-05357-5.
- Nelson DL, Cox MM (April 2005). Lehninger Principles of Biochemistry (4th ed.). W. H. Freeman. ISBN 978-0-7167-4339-2.
- Nicholls DG, Ferguson SJ (July 2002). Bioenergetics 3. Academic Press. ISBN 978-0-12-518121-1.
- Stumm W; Morgan JJ (1996). Aquatic Chemistry (3rd ed.). ISBN 978-0-471-51185-4.
- Thauer RK, Jungermann K, Decker K (March 1977). "Energy conservation in chemotrophic anaerobic bacteria". Bacteriological Reviews. 41 (1): 100–80. PMID 860983.
- White D (September 1999). The Physiology and Biochemistry of Prokaryotes (2nd ed.). ISBN 978-0-19-512579-5.
- Voet D, Voet JG (March 2004). Biochemistry. Vol. 28 (3rd ed.). )
- Kim HS, Patel K, Muldoon-Jacobs K, Bisht KS, Aykin-Burns N, Pennington JD, et al. (January 2010). "SIRT3 is a mitochondria-localized tumor suppressor required for maintenance of mitochondrial integrity and metabolism during stress". Cancer Cell. 17 (1): 41–52. PMID 20129246.
- Raimondi V, Ciccarese F, Ciminale V (January 2020). "Oncogenic pathways and the electron transport chain: a dangeROS liaison". Br J Cancer. 122 (2): 168–181. PMID 31819197.
- Reguera, Gemma (29 May 2018). "Biological electron transport goes the extra mile". Proceedings of the National Academy of Sciences. 115 (22): 5632–5634. PMID 29769327. – Editorial commentary mentioning two unusual ETCs: that of Geobacter sulfurreducens and that of cable bacteria. Also has schematic of E. coli ETC.
External links
- Electron+Transport+Chain+Complex+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Khan Academy, video lecture
- KEGG pathway: Oxidative phosphorylation, overlaid with genes found in Pseudomonas fluorescens Pf0-1. Click "help" for a how-to.








