Enolase
| phosphopyruvate hydratase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Enolase, N-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
| Enolase | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Enolase | ||||||||||
| Pfam | PF00113 | ||||||||||
| InterPro | IPR000941 | ||||||||||
| PROSITE | PDOC00148 | ||||||||||
| |||||||||||
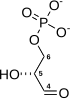
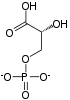
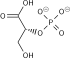
Phosphopyruvate hydratase, usually known as enolase, is a
- 2-phospho-D-glycerate phosphoenolpyruvate + H2O
Phosphopyruvate hydratase belongs to the family of lyases, specifically the hydro-lyases, which cleave carbon-oxygen bonds. The systematic name of this enzyme is 2-phospho-D-glycerate hydro-lyase (phosphoenolpyruvate-forming).
The reaction is reversible, depending on environmental concentrations of substrates.
Isozymes
In humans there are three subunits of enolase,
Three of these isoenzymes (all homodimers) are more commonly found in adult human cells than the others:- αα or non-neuronal enolase (NNE). Also known as enolase 1. Found in a variety of tissues, including liver, brain, kidney, spleen, adipose. It is present at some level in all normal human cells.
- ββ or muscle-specific enolase (MSE). Also known as enolase 3. This enzyme is largely restricted to muscle where it is present at very high levels in muscle.
- γγ or neuron-specific enolase (NSE). Also known as enolase 2. Expressed at very high levels in neurons and neural tissues, where it can account for as much as 3% of total soluble protein. It is expressed at much lower levels in most mammalian cells.
When present in the same cell, different isozymes readily form
Structure
Enolase is a member of the large
Enolase is a highly conserved enzyme with five active-site residues being especially important for activity. When compared to wild-type enolase, a mutant enolase that differs at either the Glu168, Glu211, Lys345, or Lys396 residue has an activity level that is cut by a factor of 105.[3] Also, changes affecting His159 leave the mutant with only 0.01% of its catalytic activity.[3] An integral part of enolase are two Mg2+ cofactors in the active site, which serve to stabilize negative charges in the substrate.[3][6]
Recently,
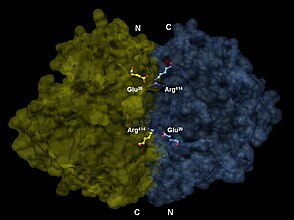
-
3-D depiction of enolase dimer in antiparallel orientation. One dimer’s N-terminal Glu20 forms an ionic bond with the other’s C-terminal Arg414 to stabilize the enzyme’s quaternary structure.
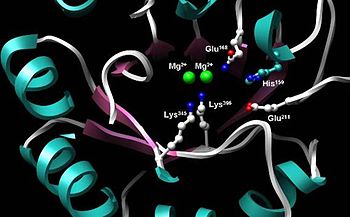
-
Active site of enolase in the middle of the C-terminal domain’s barrel. Depicted are two Mg2+ cofactors and five highly conserved residues imperative for proper catalytic function: His159, Glu168, Glu211, Lys345, Lys396.
Mechanism

Using isotopic probes, the overall mechanism for converting 2-PG to PEP is proposed to be an
Additionally, conformational changes occur within the enzyme that aid catalysis. In human α-enolase, the substrate is rotated into position upon binding to the enzyme due to interactions with the two catalytic magnesium ions,
Diagnostic uses
In recent medical experiments, enolase concentrations have been sampled in an attempt to diagnose certain conditions and their severity. For example, higher concentrations of enolase in
The same study showed that the fastest rate of tumor growth occurred in patients with the highest levels of CSF enolase. Increased levels of enolase have also been identified in patients who have suffered a recentInhibitors
Small-molecule inhibitors of enolase have been synthesized as chemical probes (substrate-analogues) of the catalytic mechanism of the enzyme and more recently, have been investigated as potential treatments for cancer and infectious diseases.
Active site transition state analogue Enolase inhibitors have been explored pre-clinically for the treatment of various microbial pathogens, as well as in precision oncology for tumors with 1p36 homozygous deletions, that lack ENO1.[31][35][36][37][38][39][40]
Fluoride is a known competitor of enolase's substrate 2-PG. Fluoride can form a complex with magnesium and phosphate, which binds in the active site instead of 2-PG.[4] One study found that fluoride could inhibit bacterial enolase in vitro.[41] The Enolase inhibitory activity of Fluoride anion may contribute to the anti-cavity effect of fluoride toothpaste, by limiting lactic acid (a product of glycolysis, which requires Enolase) production.[medical citation needed]
References
- PMID 9376357.
- ^ PDB: 2XSX; Vollmar M, Krysztofinska E, Chaikuad A, Krojer T, Cocking R, Vondelft F, Bountra C, Arrowsmith CH, Weigelt J, Edwards A, Yue WW, Oppermann U (2010). "Crystal structure of human beta enolase ENOB". Protein Data Bank.
- ^ S2CID 9191423.
- ^ S2CID 86699159.
- ^ Lohman, K; Meyerhof, O (1934). "Über die enzymatische umwandlung von phosphoglyzerinsäure in brenztraubensäure und phosphorsäure" [Enzymatic transformation of phosphoglyceric acid into pyruvic and phosphoric acid]. Biochemische Zeitschrift (in German). 273: 60–72.
- ^ PMID 1840492.
- PMID 15476816.
- S2CID 9976031.
- .
- PMID 8634301.
- PMID 8994873.
- PMID 7703246.
- PMID 8049235.
- PMID 8605183.
- PMID 7547999.
- PMID 7334408.
- PMID 2742544.
- PMID 6747647.
- PMID 18821669.
- S2CID 43249019.
- PMID 6380574.
- ^ PMID 23547795.
- PMID 1322695.
- ^ PMID 8193144.
- PMID 22895339.
- ^ Watanabe H, Yoshida J, Tanaka E, Ito M, Miyadoh S, Shomura T (1986). "Studies on a new phosphonic acid antibiotic, SF-2312". Sci Rep Meiji Seika Kaisha. 25: 12–17.
- PMID 27723749.
- PMID 31745118.
- PMID 31324042.
- ^ PMID 33230295.
- PMID 28030597.
- ^ PMID 28272459.
- S2CID 85416928.
- PMID 34604112.
- PMID 34279224.
- S2CID 226052354.
- PMID 33086062.
- PMID 31745118.
- PMID 31324042.
- PMID 2318530.
Further reading
- Holt A, Wold F (December 1961). "The isolation and characterization of rabbit muscle enolase". The Journal of Biological Chemistry. 236 (12): 3227–3231. PMID 13908561.
- Boyer, P.D., Lardy, H. and Myrback, K. (Eds.), The Enzymes, 2nd ed., vol. 5, Academic Press, New York, 1961, p. 471-494.
- Westhead EW, Mclain G (August 1964). "A Purification of Brewers' and Bakers' Yeast Enolase Yielding a Single Active Component". The Journal of Biological Chemistry. 239 (8): 2464–2468. PMID 14235523.
External links
- Enolase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)