Enzyme inhibitor
An enzyme inhibitor is a
An enzyme inhibitor stops ("inhibits") this process, either by binding to the enzyme's active site (thus preventing the substrate itself from binding) or by binding to another site on the enzyme such that the enzyme's
Enzyme inhibitors play an important role in all cells, since they are generally specific to one enzyme each and serve to control that enzyme's activity. For example, enzymes in a
Many
Structural classes
Enzyme inhibitors are a chemically diverse set of substances that range in size from organic small molecules to macromolecular proteins.
Small molecule inhibitors include essential primary metabolites that inhibit upstream enzymes that produce those metabolites. This provides a negative feedback loop that prevents over production of metabolites and thus maintains cellular homeostasis (steady internal conditions).[3][2] Small molecule enzyme inhibitors also include secondary metabolites, which are not essential to the organism that produces them, but provide the organism with an evolutionary advantage, in that they can be used to repel predators or competing organisms or immobilize prey.[4] In addition, many drugs are small molecule enzyme inhibitors that target either disease-modifying enzymes in the patient[1]: 5 or enzymes in pathogens which are required for the growth and reproduction of the pathogen.[5]
In addition to small molecules, some proteins act as enzyme inhibitors. The most prominent example are
The
Reversible inhibitors
Reversible inhibitors attach to enzymes with
In contrast to irreversible inhibitors, reversible inhibitors generally do not undergo chemical reactions when bound to the enzyme and can be easily removed by dilution or dialysis. A special case is covalent reversible inhibitors that form a chemical bond with the enzyme, but the bond can be cleaved so the inhibition is fully reversible.[13]
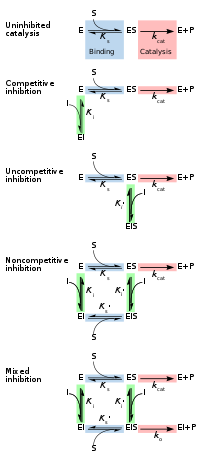
Reversible inhibitors are generally categorized into four types, as introduced by Cleland in 1963.[14] They are classified according to the effect of the inhibitor on the Vmax (maximum reaction rate catalysed by the enzyme) and Km (the concentration of substrate resulting in half maximal enzyme activity) as the concentration of the enzyme's substrate is varied.[15][16]
Competitive
In competitive inhibition the substrate and inhibitor cannot bind to the enzyme at the same time.[17]: 134 This usually results from the inhibitor having an affinity for the active site of an enzyme where the substrate also binds; the substrate and inhibitor compete for access to the enzyme's active site. This type of inhibition can be overcome by sufficiently high concentrations of substrate (Vmax remains constant), i.e., by out-competing the inhibitor.[17]: 134–135 However, the apparent Km will increase as it takes a higher concentration of the substrate to reach the Km point, or half the Vmax. Competitive inhibitors are often similar in structure to the real substrate (see for example the "methotrexate versus folate" figure in the "Drugs" section).[17]: 134
Uncompetitive
In uncompetitive inhibition the inhibitor binds only to the enzyme-substrate complex.[17]: 139 This type of inhibition causes Vmax to decrease (maximum velocity decreases as a result of removing activated complex) and Km to decrease (due to better binding efficiency as a result of Le Chatelier's principle and the effective elimination of the ES complex thus decreasing the Km which indicates a higher binding affinity).[18] Uncompetitive inhibition is rare.[17]: 139 [19]
Non-competitive
In
Mixed
In
These four types of inhibition can also be distinguished by the effect of increasing the substrate concentration [S] on the degree of inhibition caused by a given amount of inhibitor. For competitive inhibition the degree of inhibition is reduced by increasing [S], for noncompetitive inhibition the degree of inhibition is unchanged, and for uncompetitive (also called anticompetitive) inhibition the degree of inhibition increases with [S].[23]
Quantitative description
Reversible inhibition can be described quantitatively in terms of the inhibitor's
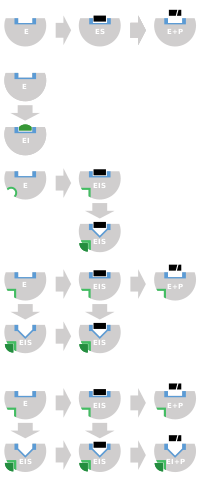
- Competitive inhibitors can bind to E, but not to ES. Competitive inhibition increases Km (i.e., the inhibitor interferes with substrate binding), but does not affect Vmax (the inhibitor does not hamper catalysis in ES because it cannot bind to ES).[24]: 102
- Uncompetitive inhibitors bind to ES. Uncompetitive inhibition decreases both Km and Vmax. The inhibitor affects substrate binding by increasing the enzyme's affinity for the substrate (decreasing Km) as well as hampering catalysis (decreases Vmax).[24]: 106
- Non-competitive inhibitors have identical affinities for E and ES (Ki = Ki'). Non-competitive inhibition does not change Km (i.e., it does not affect substrate binding) but decreases Vmax (i.e., inhibitor binding hampers catalysis).[24]: 97
- Mixed-type inhibitors bind to both E and ES, but their affinities for these two forms of the enzyme are different (Ki ≠ Ki'). Thus, mixed-type inhibitors affect substrate binding (increase or decrease Km) and hamper catalysis in the ES complex (decrease Vmax).[25]: 63–64
When an enzyme has multiple substrates, inhibitors can show different types of inhibition depending on which substrate is considered. This results from the active site containing two different binding sites within the active site, one for each substrate. For example, an inhibitor might compete with substrate A for the first binding site, but be a non-competitive inhibitor with respect to substrate B in the second binding site.[26]
Traditionally reversible enzyme inhibitors have been classified as competitive, uncompetitive, or non-competitive, according to their effects on Km and Vmax.[14] These three types of inhibition result respectively from the inhibitor binding only to the enzyme E in the absence of substrate S, to the enzyme–substrate complex ES, or to both. The division of these classes arises from a problem in their derivation and results in the need to use two different binding constants for one binding event.[27] It is further assumed that binding of the inhibitor to the enzyme results in 100% inhibition and fails to consider the possibility of partial inhibition.[27] The common form of the inhibitory term also obscures the relationship between the inhibitor binding to the enzyme and its relationship to any other binding term be it the Michaelis–Menten equation or a dose response curve associated with ligand receptor binding. To demonstrate the relationship the following rearrangement can be made:[28]
This rearrangement demonstrates that similar to the Michaelis–Menten equation, the maximal rate of reaction depends on the proportion of the enzyme population interacting with its substrate.
fraction of the enzyme population bound by substrate
fraction of the enzyme population bound by inhibitor
the effect of the inhibitor is a result of the percent of the enzyme population interacting with inhibitor. The only problem with this equation in its present form is that it assumes absolute inhibition of the enzyme with inhibitor binding, when in fact there can be a wide range of effects anywhere from 100% inhibition of substrate turn over to no inhibition. To account for this the equation can be easily modified to allow for different degrees of inhibition by including a delta Vmax term.[29]: 361
or
This term can then define the residual enzymatic activity present when the inhibitor is interacting with individual enzymes in the population. However the inclusion of this term has the added value of allowing for the possibility of activation if the secondary Vmax term turns out to be higher than the initial term. To account for the possibly of activation as well the notation can then be rewritten replacing the inhibitor "I" with a modifier term (stimulator or inhibitor) denoted here as "X".[28]: eq 13
While this terminology results in a simplified way of dealing with kinetic effects relating to the maximum velocity of the Michaelis–Menten equation, it highlights potential problems with the term used to describe effects relating to the Km. The Km relating to the affinity of the enzyme for the substrate should in most cases relate to potential changes in the binding site of the enzyme which would directly result from enzyme inhibitor interactions. As such a term similar to the delta Vmax term proposed above to modulate Vmax should be appropriate in most situations:[28]: eq 14
Dissociation constants
![2D plots of 1/[S] concentration (x-axis) and 1/V (y-axis) demonstrating that as inhibitor concentration is changed, competitive inhibitor lines intersect at a single point on the y-axis, non-competitive inhibitors intersect at the x-axis, and mixed inhibitors intersect a point that is on neither axis](http://upload.wikimedia.org/wikipedia/commons/thumb/b/ba/Inhibition_diagrams-1-.png/220px-Inhibition_diagrams-1-.png)
An enzyme inhibitor is characterised by its
where the modifying factors α and α' are defined by the inhibitor concentration and its two dissociation constants
Thus, in the presence of the inhibitor, the enzyme's effective Km and Vmax become (α/α')Km and (1/α')Vmax, respectively. However, the modified Michaelis-Menten equation assumes that binding of the inhibitor to the enzyme has reached equilibrium, which may be a very slow process for inhibitors with sub-nanomolar dissociation constants. In these cases the inhibition becomes effectively irreversible, hence it is more practical to treat such tight-binding inhibitors as irreversible (see below).
The effects of different types of reversible enzyme inhibitors on enzymatic activity can be visualised using graphical representations of the Michaelis–Menten equation, such as
Special cases
Partially competitive
The mechanism of partially competitive inhibition is similar to that of non-competitive, except that the EIS complex has catalytic activity, which may be lower or even higher (partially competitive activation) than that of the enzyme–substrate (ES) complex. This inhibition typically displays a lower Vmax, but an unaffected Km value.[18]
Substrate or product
Substrate or product inhibition is where either an enzymes substrate or product also act as an inhibitor. This inhibition may follow the competitive, uncompetitive or mixed patterns. In substrate inhibition there is a progressive decrease in activity at high substrate concentrations, potentially from an enzyme having two competing substrate-binding sites. At low substrate, the high-affinity site is occupied and normal kinetics are followed. However, at higher concentrations, the second inhibitory site becomes occupied, inhibiting the enzyme.[34] Product inhibition (either the enzyme's own product, or a product to an enzyme downstream in its metabolic pathway) is often a regulatory feature in metabolism and can be a form of negative feedback.[2]
Slow-tight
Slow-tight inhibition occurs when the initial enzyme–inhibitor complex EI undergoes
Multi-substrate analogues
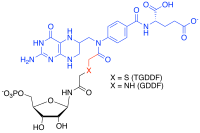
Multi-substrate analogue inhibitors are high affinity selective inhibitors that can be prepared for enzymes that catalyse reactions with more than one substrate by capturing the binding energy of each of those substrate into one molecule.
Examples
As enzymes have evolved to bind their substrates tightly, and most reversible inhibitors bind in the active site of enzymes, it is unsurprising that some of these inhibitors are strikingly similar in structure to the substrates of their targets. Inhibitors of
Enzyme inhibitors are often designed to mimic the transition state or intermediate of an enzyme-catalysed reaction.[46] This ensures that the inhibitor exploits the transition state stabilising effect of the enzyme, resulting in a better binding affinity (lower Ki) than substrate-based designs. An example of such a transition state inhibitor is the antiviral drug oseltamivir; this drug mimics the planar nature of the ring oxonium ion in the reaction of the viral enzyme neuraminidase.[47]
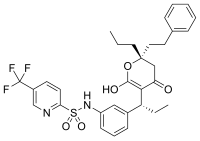
However, not all inhibitors are based on the structures of substrates. For example, the structure of another HIV protease inhibitor
In drug design it is important to consider the concentrations of substrates to which the target enzymes are exposed. For example, some protein kinase inhibitors have chemical structures that are similar to ATP, one of the substrates of these enzymes.[49] However, drugs that are simple competitive inhibitors will have to compete with the high concentrations of ATP in the cell. Protein kinases can also be inhibited by competition at the binding sites where the kinases interact with their substrate proteins, and most proteins are present inside cells at concentrations much lower than the concentration of ATP. As a consequence, if two protein kinase inhibitors both bind in the active site with similar affinity, but only one has to compete with ATP, then the competitive inhibitor at the protein-binding site will inhibit the enzyme more effectively.[50]
Irreversible inhibitors
Types
Irreversible inhibitors
Irreversible inhibition is different from irreversible enzyme inactivation.[54] Irreversible inhibitors are generally specific for one class of enzyme and do not inactivate all proteins; they do not function by destroying protein structure but by specifically altering the active site of their target. For example, extremes of pH or temperature usually cause denaturation of all protein structure, but this is a non-specific effect. Similarly, some non-specific chemical treatments destroy protein structure: for example, heating in concentrated hydrochloric acid will hydrolyse the peptide bonds holding proteins together, releasing free amino acids.[55]
Irreversible inhibitors display time-dependent inhibition and their potency therefore cannot be characterised by an IC50 value. This is because the amount of active enzyme at a given concentration of irreversible inhibitor will be different depending on how long the inhibitor is pre-incubated with the enzyme. Instead, kobs/[I] values are used,[56] where kobs is the observed pseudo-first order rate of inactivation (obtained by plotting the log of % activity versus time) and [I] is the concentration of inhibitor. The kobs/[I] parameter is valid as long as the inhibitor does not saturate binding with the enzyme (in which case kobs = kinact) where kinact is the rate of inactivation.
Measuring
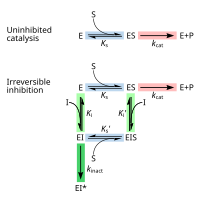
Irreversible inhibitors first form a reversible non-covalent complex with the enzyme (EI or ESI). Subsequently, a chemical reaction occurs between the enzyme and inhibitor to produce the covalently modified "dead-end complex" EI* (an irreversible covalent complex). The rate at which EI* is formed is called the inactivation rate or kinact.[13] Since formation of EI may compete with ES, binding of irreversible inhibitors can be prevented by competition either with substrate or with a second, reversible inhibitor. This protection effect is good evidence of a specific reaction of the irreversible inhibitor with the active site.
The binding and inactivation steps of this reaction are investigated by incubating the enzyme with inhibitor and assaying the amount of activity remaining over time. The activity will be decreased in a time-dependent manner, usually following exponential decay. Fitting these data to a rate equation gives the rate of inactivation at this concentration of inhibitor. This is done at several different concentrations of inhibitor. If a reversible EI complex is involved the inactivation rate will be saturable and fitting this curve will give kinact and Ki.[57]
Another method that is widely used in these analyses is
Slow binding
Not all irreversible inhibitors form covalent adducts with their enzyme targets. Some reversible inhibitors bind so tightly to their target enzyme that they are essentially irreversible. These tight-binding inhibitors may show kinetics similar to covalent irreversible inhibitors. In these cases some of these inhibitors rapidly bind to the enzyme in a low-affinity EI complex and this then undergoes a slower rearrangement to a very tightly bound EI* complex (see the "irreversible inhibition mechanism" diagram). This kinetic behaviour is called slow-binding.
Some examples

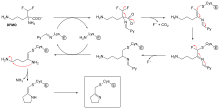
Suicide inhibition is an unusual type of irreversible inhibition where the enzyme converts the inhibitor into a reactive form in its active site.[69] An example is the inhibitor of polyamine biosynthesis, α-difluoromethylornithine (DFMO), which is an analogue of the amino acid ornithine, and is used to treat African trypanosomiasis (sleeping sickness). Ornithine decarboxylase can catalyse the decarboxylation of DFMO instead of ornithine (see the "DFMO inhibitor mechanism" diagram). However, this decarboxylation reaction is followed by the elimination of a fluorine atom, which converts this catalytic intermediate into a conjugated imine, a highly electrophilic species. This reactive form of DFMO then reacts with either a cysteine or lysine residue in the active site to irreversibly inactivate the enzyme.[61]
Since irreversible inhibition often involves the initial formation of a non-covalent enzyme inhibitor (EI) complex,[13] it is sometimes possible for an inhibitor to bind to an enzyme in more than one way. For example, in the figure showing trypanothione reductase from the human protozoan parasite Trypanosoma cruzi, two molecules of an inhibitor called quinacrine mustard are bound in its active site. The top molecule is bound reversibly, but the lower one is bound covalently as it has reacted with an amino acid residue through its nitrogen mustard group.[70]
Applications
Enzyme inhibitors are found in nature
Metabolic regulation
Enzyme inhibition is a common feature of
Physiological enzyme inhibition can also be produced by specific protein inhibitors. This mechanism occurs in the pancreas, which synthesises many digestive precursor enzymes known as zymogens. Many of these are activated by the trypsin protease, so it is important to inhibit the activity of trypsin in the pancreas to prevent the organ from digesting itself. One way in which the activity of trypsin is controlled is the production of a specific and potent trypsin inhibitor protein in the pancreas. This inhibitor binds tightly to trypsin, preventing the trypsin activity that would otherwise be detrimental to the organ.[78] Although the trypsin inhibitor is a protein, it avoids being hydrolysed as a substrate by the protease by excluding water from trypsin's active site and destabilising the transition state.[79] Other examples of physiological enzyme inhibitor proteins include the barstar inhibitor of the bacterial ribonuclease barnase.[80]
Natural poisons

Animals and plants have evolved to synthesise a vast array of poisonous products including secondary metabolites,[81] peptides and proteins[82] that can act as inhibitors. Natural toxins are usually small organic molecules and are so diverse that there are probably natural inhibitors for most metabolic processes.[83] The metabolic processes targeted by natural poisons encompass more than enzymes in metabolic pathways and can also include the inhibition of receptor, channel and structural protein functions in a cell. For example, paclitaxel (taxol), an organic molecule found in the Pacific yew tree, binds tightly to tubulin dimers and inhibits their assembly into microtubules in the cytoskeleton.[84]
Many natural poisons act as
Although many natural toxins are secondary metabolites, these poisons also include peptides and proteins. An example of a toxic peptide is
Proteins can also be natural poisons or
Drugs

The most common uses for enzyme inhibitors are as drugs to treat disease. Many of these inhibitors target a human enzyme and aim to correct a pathological condition. For instance,
As of 2017,[update] an estimated 29% of approved drugs are enzyme inhibitors
An example of the structural similarity of some inhibitors to the substrates of the enzymes they target is seen in the figure comparing the drug
A common treatment for
Antibiotics

Drugs are also used to inhibit enzymes needed for the survival of pathogens. For example, bacteria are surrounded by a thick cell wall made of a net-like polymer called peptidoglycan. Many antibiotics such as penicillin and vancomycin inhibit the enzymes that produce and then cross-link the strands of this polymer together.[102][103] This causes the cell wall to lose strength and the bacteria to burst. In the figure, a molecule of penicillin (shown in a ball-and-stick form) is shown bound to its target, the transpeptidase from the bacteria Streptomyces R61 (the protein is shown as a ribbon diagram).
Antibiotic drug design is facilitated when an enzyme that is essential to the pathogen's survival is absent or very different in humans.[104] Humans do not make peptidoglycan, therefore antibiotics that inhibit this process are selectively toxic to bacteria.[105] Selective toxicity is also produced in antibiotics by exploiting differences in the structure of the ribosomes in bacteria,[106] or how they make fatty acids.[107]
Antivirals
Drugs that inhibit enzymes needed for the
Pesticides
Many
Herbicides
The herbicide
Discovery and design

New drugs are the products of a long drug development process, the first step of which is often the discovery of a new enzyme inhibitor.[123] There are two principle approaches of discovering these inhibitors.[124]
The first general method is
The second way of discovering new enzyme inhibitors is
Hits from any of the above approaches can be
See also
- Activity-based proteomics – a branch of proteomics that uses covalent enzyme inhibitors as reporters to monitor enzyme activity.
- Antimetabolite – an enzyme inhibitor that is used to interfere with cell growth and division
- Transition state analogue– a type of enzyme inhibitor that mimics the transition state of the chemical reaction catalysed by the enzyme
References
- ^ ISBN 978-1-118-48813-3.
- ^ PMID 28202588.
- ^ ISBN 978-0-471-67557-0. Archivedfrom the original on 28 March 2023. Retrieved 14 April 2022.
- ^ PMID 12821301.
- ISBN 978-3-527-65970-8.
- PMID 30809526.
- PMID 1998683.
- S2CID 215772580.
- ^ ISBN 978-0-08-091488-6. Archivedfrom the original on 28 March 2023. Retrieved 20 July 2022.
- PMID 30917186.
- ISBN 978-0-19-969739-7. Archivedfrom the original on 20 July 2022. Retrieved 20 July 2022.
- ISBN 978-1-135-08892-7. Archivedfrom the original on 7 June 2022. Retrieved 7 June 2022.
- ^ PMID 29689165.
- ^ PMID 14021668.
- ISBN 978-0-7167-4955-4. Archivedfrom the original on 26 September 2009. Retrieved 31 August 2017.
- ^ a b c "Types of Inhibition". NIH Center for Translational Therapeutics. Archived from the original on 8 September 2011. Retrieved 2 April 2012.
- ^ ISBN 978-3-527-66549-5.
- ^ ISBN 978-0-471-30309-1.
- ISBN 978-1-904275-27-5.
- from the original on 28 November 2021. Retrieved 3 April 2022.
- ^ ISBN 978-1-118-91840-1.
- S2CID 73480979.
In some cases, the inhibitor may bind to a distinct site on the enzyme that is in allosteric communication with the substrate binding pocket. In many cases, allosteric, substrate competitive compounds result in conformational changes to the enzyme that change the ability of the enzyme to bind substrate.
- ISBN 978-0-8053-5680-9.
- ^ ISBN 978-3-527-80647-8.
- ISBN 978-0-471-15985-8.
- ISBN 978-0-471-30309-1.
- ^ PMID 22038120.
- ^ PMID 17307293.
- ^ Walsh R (May 2012). "Alternative perspectives of enzyme kinetic modeling". Medicinal Chemistry and Drug Design. (16). InTech: 357–372.
- from the original on 15 June 2022. Retrieved 9 April 2022.
- PMID 11464510.
- PMID 2077683.
- ^ PMID 2246896.
- ISBN 978-0-470-20745-1.
- PMID 7791615.
- PMID 319722.
- PMID 2709379.
- .
- PMID 9417034.
- PMID 20194791.
- S2CID 21808708.
- ^ ISBN 978-0-444-62667-7.
- PMID 16611117.
- S2CID 25269012.
- PMID 21864279.
- ^ PMID 30335982.
- PMID 10702632.
- PMID 14529528.
- PMID 25494294.
- PMID 16182621.
- ^ ISBN 978-0-19-874969-1. Archivedfrom the original on 28 March 2023. Retrieved 3 June 2022.
- S2CID 56480231.
- ISBN 978-0-8493-1983-9.
- ISBN 978-0-444-82970-2.
Enzyme inactivation is generally explained as a chemical process involving several phenomena like aggregation, dissociation into subunits, or denaturation (conformational changes), which occur simultaneously during the inactivation of a specific enzyme.
- ISBN 978-0-12-564710-6.
- PMID 11182321.
- PMID 11741224.
- S2CID 11766917.
- ISBN 978-0-12-380925-4. Archivedfrom the original on 20 July 2022. Retrieved 20 July 2022.
- PMID 22902804.
- ^ PMID 1730582.
- PMID 7791610.
- PMID 3511964.
- PMID 4322209.
- PMID 2553730.
- PMID 15102853.
- ISBN 978-0-12-181860-9. Archived from the originalon 28 February 2018.
- ISBN 978-0-7216-7757-6.
- PMID 6433782.
- PMID 15102853.
- ISBN 978-3-527-34205-1. Archived(PDF) from the original on 5 December 2022. Retrieved 3 April 2022.
- PMID 16041744.
- ISBN 978-0-7020-7819-4. Archivedfrom the original on 25 April 2013. Retrieved 3 June 2022.
- ISBN 978-0-203-64683-0. Archivedfrom the original on 28 March 2023. Retrieved 14 April 2022.
- from the original on 17 January 2023. Retrieved 14 April 2022.
- ISBN 978-3-642-13865-2.
- S2CID 24586866.
- ISBN 978-0-19-850229-6.
- from the original on 28 March 2023. Retrieved 14 April 2022.
- PMID 2696173.
- ISBN 978-981-19-0932-0.
- ISBN 978-3-540-00118-8.
- PMID 16515527.
- PMID 12769688.
- .
- S2CID 13701908.
- S2CID 36465515.
- PMID 9604278.
- PMID 12369866.
- PMID 11577938.
- ISBN 978-0-12-227055-0.
- PMID 31174319.
- PMID 31216687.
- PMID 15450171.
- ^ ISBN 978-0-7817-7155-9. Archivedfrom the original on 28 March 2023. Retrieved 14 April 2022.
- ^ PMID 27910877.
Figure 1C: Clinical success of privileged protein family classes (% approved drugs targeting each target class): Reductase 7.62, Kinase 5.94, Protease 3.35, Hydrolase 2.76, NPTase 2.09, Transferase 1.92, Lyase 1.59, Isomerase 1.51, Phosphodiesterase 1.50, Cytochrome p450 0.84, Epigenetic eraser 0.33, Total enzyme targets of approved drugs = 29.45%
- PMID 30259761.
- S2CID 237311419.
- PMID 14529544.
- S2CID 52945888.
- PMID 10913932.
- from the original on 28 March 2023. Retrieved 20 July 2022.
- PMID 17894443.
- PMID 33367978.
- from the original on 9 April 2022. Retrieved 9 April 2022.
- S2CID 231761270.
- PMID 27668701.
- OCLC 1240584737.
- S2CID 195356119.
- S2CID 4489039.
- PMID 35847492.
- S2CID 212651676.
- S2CID 53763831.
- ISBN 978-0-87819-052-2.
- ISBN 978-1-00-009898-3.
- ISBN 978-0-08-054310-9. Archivedfrom the original on 21 July 2022. Retrieved 21 July 2022.
- PMID 29655452.
- ^ Butterworth JF, IV, Mackey DC, Wasnick JD, eds. (2013). "Chapter 12. Cholinesterase Inhibitors & Other Pharmacologic Antagonists to Neuromuscular Blocking Agents.". Morgan & Mikhail's Clinical Anesthesiology (5th ed.). McGraw Hill. Archived from the original on 30 June 2022. Retrieved 9 April 2022.
- PMID 28117022.
- S2CID 2358278.
- ISBN 978-3-642-59416-8. Archivedfrom the original on 27 June 2022. Retrieved 27 June 2022.
Chapter 10.2.1: Sulfonylurea acetolactate synthase inhibitors
- PMID 1980104.
- ISBN 978-1-00-043721-8.
- S2CID 231633284.
- ^ Lindquist RN (October 2013). "The design of enzyme inhibitors: Transition state analogues.". In Ariëns EJ (ed.). Drug Design: Medicinal Chemistry: A Series of Monographs. Vol. 5. Elsevier. pp. 24–80.
- PMID 16796557.
- PMID 12203463.
- PMID 16713909.
- PMID 17959370.
- S2CID 232059044.
- ISBN 978-1-315-11347-0.
- S2CID 226271153.
- PMID 33810118.
External links
- "BRENDA". Archived from the original on 1 April 2022., Database of enzymes giving lists of known inhibitors for each entry
- "PubChem". National Center for Biotechnology Information. National Library of Medicine. Database of drugs and enzyme inhibitors
- "Symbolism and Terminology in Enzyme Kinetics". Archived from the original on 20 June 2006. Recommendations of the Nomenclature Committee of the International Union of Biochemistry (NC-IUB) on enzyme inhibition terminology

![{\displaystyle {\begin{aligned}{\cfrac {V_{\max }}{1+{\cfrac {\ce {[I]}}{K_{i}}}}}&={V_{\max }}\left({\cfrac {K_{i}}{K_{i}+[{\ce {I}}]}}\right)&&{\text{multiply by }}{\cfrac {K_{i}}{K_{i}}}=1\\&={V_{\max }}\left({\cfrac {K_{i}+[{\ce {I}}]-[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{add }}[{\ce {I}}]-[{\ce {I}}]=0{\text{ to numerator}}\\&={V_{\max }}\left(1-{\cfrac {[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{simplify }}{\cfrac {K_{i}+[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}=1\\&=V_{\max }-V_{\max }{\cfrac {\ce {[I]}}{K_{i}+[{\ce {I}}]}}&&{\text{multiply out by }}V_{\max }\end{aligned}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/37eda4dec307f8acfca89b2d8f4811474ea764ec)
![{\displaystyle {\cfrac {\ce {[S]}}{[{\ce {S}}]+K_{m}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/eb08dd139085a394e6e7370f47ebfa255f1ad685)
![{\displaystyle {\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/9ed50a1f7a5f2c52f406b52263916ab48b268e07)
![{\displaystyle V_{\max }-\Delta V_{\max }{\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/7dff424ec79284c3a1cea14f0f82b0eaace53c69)
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/f3874623edd9524ba2741fe448927bf5cf0ab257)
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/306d44733a89308883053e3b8372a8cf9ce0239b)
![{\displaystyle K_{m1}-(K_{m1}-K_{m2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/cb4e0de216e1e625bb803ee725bf85c9989a15f5)
![{\displaystyle V={\frac {V_{max}[S]}{\alpha K_{m}+\alpha ^{\prime }[S]}}={\frac {(1/\alpha ^{\prime })V_{max}[S]}{(\alpha /\alpha ^{\prime })K_{m}+[S]}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4a8f0a9dda1d308de7f090f99c2833f944f11a09)
![{\displaystyle \alpha =1+{\frac {[I]}{K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/57fcf54938a9784f9313437681b220079ff43ee5)
![{\displaystyle \alpha ^{\prime }=1+{\frac {[I]}{K_{i}^{\prime }}}.}](https://wikimedia.org/api/rest_v1/media/math/render/svg/65bf16742482cae7b0743781f47c327ddcf537e3)