Evidence of common descent
| Part of a series on |
| Evolutionary biology |
|---|
 |
Evidence of common descent of living organisms has been discovered by scientists researching in a variety of disciplines over many decades, demonstrating that all life on Earth comes from a single ancestor. This forms an important part of the evidence on which evolutionary theory rests, demonstrates that evolution does occur, and illustrates the processes that created Earth's biodiversity. It supports the modern evolutionary synthesis—the current scientific theory that explains how and why life changes over time. Evolutionary biologists document evidence of common descent, all the way back to the last universal common ancestor, by developing testable predictions, testing hypotheses, and constructing theories that illustrate and describe its causes.
Comparison of the DNA genetic sequences of organisms has revealed that organisms that are phylogenetically close have a higher degree of DNA sequence similarity than organisms that are phylogenetically distant. Genetic fragments such as pseudogenes, regions of DNA that are orthologous to a gene in a related organism, but are no longer active and appear to be undergoing a steady process of degeneration from cumulative mutations support common descent alongside the universal biochemical organization and molecular variance patterns found in all organisms. Additional genetic information conclusively supports the relatedness of life and has allowed scientists (since the discovery of DNA) to develop phylogenetic trees: a construction of organisms' evolutionary relatedness. It has also led to the development of molecular clock techniques to date taxon divergence times and to calibrate these with the fossil record.
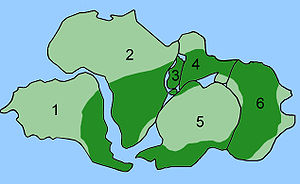
Evidence from animal coloration was gathered by some of Darwin's contemporaries; camouflage, mimicry, and warning coloration are all readily explained by natural selection. Special cases like the seasonal changes in the plumage of the ptarmigan, camouflaging it against snow in winter and against brown moorland in summer provide compelling evidence that selection is at work. Further evidence comes from the field of biogeography because evolution with common descent provides the best and most thorough explanation for a variety of facts concerning the geographical distribution of plants and animals across the world. This is especially obvious in the field of insular biogeography. Combined with the well-established geological theory of plate tectonics, common descent provides a way to combine facts about the current distribution of species with evidence from the fossil record to provide a logically consistent explanation of how the distribution of living organisms has changed over time.
The development and spread of
Evidence from comparative physiology and biochemistry
Genetics
One of the strongest evidences for common descent comes from gene sequences. Comparative sequence analysis examines the relationship between the DNA sequences of different species,[1] producing several lines of evidence that confirm Darwin's original hypothesis of common descent. If the hypothesis of common descent is true, then species that share a common ancestor inherited that ancestor's DNA sequence, as well as mutations unique to that ancestor. More closely related species have a greater fraction of identical sequence and shared substitutions compared to more distantly related species.

The simplest and most powerful evidence is provided by phylogenetic reconstruction. Such reconstructions, especially when done using slowly evolving protein sequences, are often quite robust and can be used to reconstruct a great deal of the evolutionary history of modern organisms (and even in some instances of the evolutionary history of extinct organisms, such as the recovered gene sequences of mammoths or Neanderthals). These reconstructed phylogenies recapitulate the relationships established through morphological and biochemical studies.[2] The most detailed reconstructions have been performed on the basis of the mitochondrial genomes shared by all eukaryotic organisms,[3] which are short and easy to sequence; the broadest reconstructions have been performed either using the sequences of a few very ancient proteins or by using ribosomal RNA sequence.[citation needed]
Phylogenetic relationships extend to a wide variety of nonfunctional sequence elements, including repeats,
Universal biochemical organisation and molecular variance patterns
All known
DNA sequencing
Comparison of DNA sequences allows organisms to be grouped by sequence similarity, and the resulting
Some DNA sequences are shared by very different organisms. It has been predicted by the theory of evolution that the differences in such DNA sequences between two organisms should roughly resemble both the biological difference between them according to their
Proteins
Pseudogenes
Pseudogenes, also known as
Other mechanisms
A large body of molecular evidence supports a variety of mechanisms for large evolutionary changes, including:
Since
Specific examples from comparative physiology and biochemistry
Chromosome 2 in humans
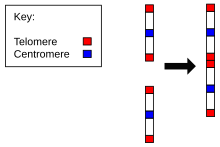
Evidence for the evolution of
The evidence for this includes:
- The correspondence of chromosome 2 to two ape chromosomes. The closest human relative, the chimpanzee, has near-identical DNA sequences to human chromosome 2, but they are found in two separate chromosomes. The same is true of the more distant gorilla and orangutan.[27][28]
- The presence of a vestigial centromere. Normally a chromosome has just one centromere, but in chromosome 2 there are remnants of a second centromere.[29]
- The presence of vestigial telomeres. These are normally found only at the ends of a chromosome, but in chromosome 2 there are additional telomere sequences in the middle.[30]
Chromosome 2 thus presents strong evidence in favour of the common descent of humans and other apes. According to J. W. Ijdo, "We conclude that the locus cloned in cosmids c8.1 and c29B is the relic of an ancient telomere-telomere fusion and marks the point at which two ancestral ape chromosomes fused to give rise to human chromosome 2."[30]
Cytochrome c and b
A classic example of biochemical evidence for evolution is the variance of the ubiquitous (i.e. all living organisms have it, because it performs very basic life functions) protein Cytochrome c in living cells. The variance of cytochrome c of different organisms is measured in the number of differing amino acids, each differing amino acid being a result of a base pair substitution, a mutation. If each differing amino acid is assumed the result of one base pair substitution, it can be calculated how long ago the two species diverged by multiplying the number of base pair substitutions by the estimated time it takes for a substituted base pair of the cytochrome c gene to be successfully passed on. For example, if the average time it takes for a base pair of the cytochrome c gene to mutate is N years, the number of amino acids making up the cytochrome c protein in monkeys differ by one from that of humans, this leads to the conclusion that the two species diverged N years ago.
The primary structure of cytochrome c consists of a chain of about 100 amino acids. Many higher order organisms possess a chain of 104 amino acids.[31]
The cytochrome c molecule has been extensively studied for the glimpse it gives into evolutionary biology. Both
What makes these homologous similarities particularly suggestive of common ancestry in the case of cytochrome c, in addition to the fact that the phylogenies derived from them match other phylogenies very well, is the high degree of functional redundancy of the cytochrome c molecule. The different existing configurations of amino acids do not significantly affect the functionality of the protein, which indicates that the base pair substitutions are not part of a directed design, but the result of random mutations that are not subject to selection.[33]
In addition, Cytochrome b is commonly used as a region of
Endogenous retroviruses
Recent African origin of modern humans
Mathematical models of evolution, pioneered by the likes of
Evidence from comparative anatomy
Atavisms
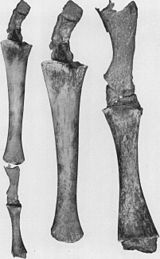
Once thought of as a refutation to evolutionary theory, atavisms are "now seen as potent evidence of how much genetic potential is retained...after a particular structure has disappeared from a species".[43] "Atavisms are the reappearance of a lost character typical of remote ancestors and not seen in the parents or recent ancestors..."[44] and are an "[indication] of the developmental plasticity that exists within embryos..."[44] Atavisms occur because genes for previously existing phenotypical features are often preserved in DNA, even though the genes are not expressed in some or most of the organisms possessing them.[45] Numerous examples have documented the occurrence of atavisms alongside experimental research triggering their formation. Due to the complexity and interrelatedness of the factors involved in the development of atavisms, both biologists and medical professionals find it "difficult, if not impossible, to distinguish [them] from malformations."[46]
Some examples of atavisms found in the scientific literature include:
- Hind limbs in whales.[44] (see figure 2a)
- Reappearance of limbs in limbless vertebrates.[43][44][47]
- Back pair of flippers on a bottlenose dolphin.[48]
- Extra toes of the modern horse.[44][49][50]
- Human tails (not pseudo-tails)[46][51] and extra nipples in humans.[44]
- Re-evolution of sexuality from parthenogenesis in orbitid mites.[52]
- Teeth in chickens.[53]
- Dewclaws in dogs.[44]
- Reappearance of wings on wingless stick insects[54] and earwigs.[44]
- Atavistic muscles in several birds[55][56] and mammals such as the beagle[57] and the jerboa.[55]
- Extra toes in
Evolutionary developmental biology and embryonic development
Evolutionary developmental biology is the biological field that compares the developmental process of different organisms to determine ancestral relationships between species. A large variety of organism's genomes contain a
Some examples include:
- Hair growth and loss (lanugo) during human development.[59]
- Development and degeneration of a yolk sac.
- Terrestrial frogs and salamanders passing through the larval stage within the egg—with features of typically aquatic larvae—but hatch ready for life on land;[60]
- The appearance of gill-like structures (pharyngeal arch) in vertebrate embryo development. Note that in fish, the arches continue to develop as branchial arches while in humans, for example, they give rise to a variety of structures within the head and neck.
Homologous structures and divergent (adaptive) evolution
If widely separated groups of organisms are originated from a common ancestry, they are expected to have certain basic features in common. The degree of resemblance between two organisms should indicate how closely related they are in evolution:
- Groups with little in common are assumed to have diverged from a common ancestormuch earlier in geological history than groups with a lot in common;
- In deciding how closely related two animals are, a comparative anatomist looks for structures that are fundamentally similar, even though they may serve different functions in the adult. Such structures are described as homologous and suggest a common origin.
- In cases where the similar structures serve different functions in adults, it may be necessary to trace their origin and embryonic development. A similar developmental origin suggests they are the same structure, and thus likely derived from a common ancestor.
When a group of organisms share a homologous structure that is specialized to perform a variety of functions to adapt different environmental conditions and modes of life, it is called adaptive radiation. The gradual spreading of organisms with adaptive radiation is known as divergent evolution.
Nested hierarchies and classification
All the foregoing rules and aids and difficulties in classification are explained, if I do not greatly deceive myself, on the view that the natural system is founded on descent with modification; that the characters which naturalists consider as showing true affinity between any two or more species, are those which have been inherited from a common parent, and, in so far, all true classification is genealogical; that community of descent is the hidden bond which naturalists have been unconsciously seeking, ...
— Charles Darwin, On the Origin of Species, page 577
Evolutionary trees
An
Vestigial structures
Evidence for common descent comes from the existence of vestigial structures.[72] These rudimentary structures are often homologous to structures that correspond in related or ancestral species. A wide range of structures exist such as mutated and non-functioning genes, parts of a flower, muscles, organs, and even behaviors. This variety can be found across many different groups of species. In many cases they are degenerated or underdeveloped. The existence of vestigial organs can be explained in terms of changes in the environment or modes of life of the species. Those organs are typically functional in the ancestral species but are now either semi-functional, nonfunctional, or re-purposed.
Scientific literature concerning vestigial structures abounds. One study compiled 64 examples of vestigial structures found in the literature across a wide range of disciplines within the 21st century.[73] The following non-exhaustive list summarizes Senter et al. alongside various other examples:
- The presence of remnant Giardia intestinalis.[74]
- Remnant Epifagus americana.[75]
- Missing Gethyum flowers.[76]
- Non-functioning
- Remnant stamens on female flowers of Fragaria virginiana;[78] all species in the genus Schiedea;[79] and on Penstemon centranthifolius, P. rostriflorus, P. ellipticus, and P. palmeri.[80]
- Vestigial anthers on Nemophila menziesii.[81]
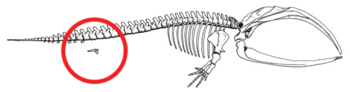
- Reduced hindlimbs and display an example of an atavism predicted from their common ancestry.
- Nonfunctional hind wings in Carabus solieri[87] and other beetles.[83]
- Remnant eyes (and eye structures) in animals that have lost sight such as blind cavefish (e.g.
- Vestigial eye in the extant
- Functionless wings in flightless birds such as ostriches, kiwis, cassowaries, and emus.[93][94]
- The presence of the plica semilunaris in the human eye—a vestigial remnant of the nictitating membrane.[95]
- Harderian gland in primates.[96]
- Reduced hind limbs and pelvic girdle structures in
- Reduced and missing olfactory apparatus in whales that still possess vestigial olfactory receptor subgenomes.[100]
- Vestigial teeth in narwhal.[101]
- Rudimentary digits of Perodicticus potto.[102]
- Vestigial dental primordia in the embryonic tooth pattern in mice.[103]
- Reduced or absent vomeronasal organ in humans and Old World monkeys.[104][105]
- Presence of non-functional sinus hair muscles in humans used in whisker movement.[106]
- Degenerating palmaris longus muscle in humans.[107]
- ascorbic acid), yet still possess the genes involved. This inability is due to mutations of the L-gulono-γ-lactone oxidase (GLO) gene— and in primates, teleost fish, and guinea pigs it is irreversible.[108]
- Remnant abdominal segments in cirripedes (barnacles).[109]
- Non-mammalian vertebrate embryos depend on nutrients from the yolk sac. Humans and other mammal genomes contain broken, non-functioning genes that code for the production of yolk. alongside the presence of an empty yolk sac with the embryo.[110][111][112]
- Dolphin embryonic limb buds.[113]
- Leaf formation in some cacti species.[114]
- Presence of a vestigial endosymbiont Lepidodinium viride within the dinoflagellate Gymnodinium chlorophorum.[115]
- The species Dolabrifera dolabrifera has an ink gland but is "incapable of producing ink or its associated anti-predator proteins".[116]
Specific examples from comparative anatomy
This section needs additional citations for verification. (April 2021) |
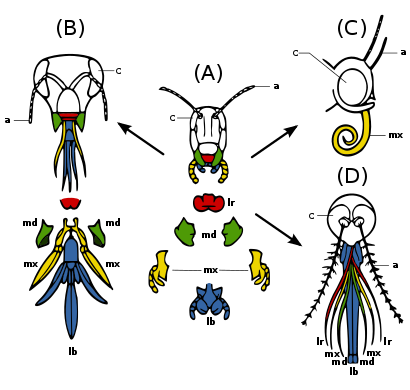
(A) Primitive state — biting and chewing: e.g.
Insect mouthparts and appendages
Many different species of insects have mouthparts derived from the same embryonic structures, indicating that the mouthparts are modifications of a common ancestor's original features. These include a
Insect mouthparts and antennae are considered homologues of insect legs. Parallel developments are seen in some
Pelvic structure of dinosaurs
Similar to the pentadactyl limb in mammals, the earliest
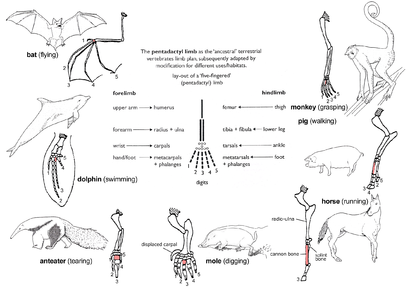
Pentadactyl limb
The pattern of limb bones called
(digits). Throughout the tetrapods, the fundamental structures of pentadactyl limbs are the same, indicating that they originated from a common ancestor. But in the course of evolution, these fundamental structures have been modified. They have become superficially different and unrelated structures to serve different functions in adaptation to different environments and modes of life. This phenomenon is shown in the forelimbs of mammals. For example:- In monkeys, the forelimbs are much elongated, forming a grasping hand used for climbing and swinging among trees.
- Pigs have lost their first digit, while the second and fifth digits are reduced. The remaining two digits are longer and stouter than the rest and bear a hoof for supporting the body.
- In horses, the forelimbs are highly adapted for strength and support. Fast and long-distance running is possible due to the extensive elongation of the third digit that bears a hoof.
- The burrowing.
- Anteaters use their enlarged third digit for tearing into ant and termite nests.
- In cetaceans, the forelimbs become flippers for steering and maintaining equilibrium during swimming.
- In bats, the forelimbs have become highly modified and evolved into functioning wings. Four digits have become elongated, while the hook-like first digit remains free and is used to grip.
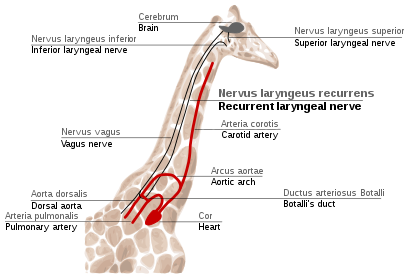
Recurrent laryngeal nerve in giraffes
The
This path is suboptimal even for humans, but for
The indirect route of this nerve is the result of evolution of mammals from fish, which had no neck and had a relatively short nerve that innervated one gill slit and passed near the gill arch. Since then, the gill it innervated has become the larynx and the gill arch has become the dorsal aorta in mammals.[117][118]
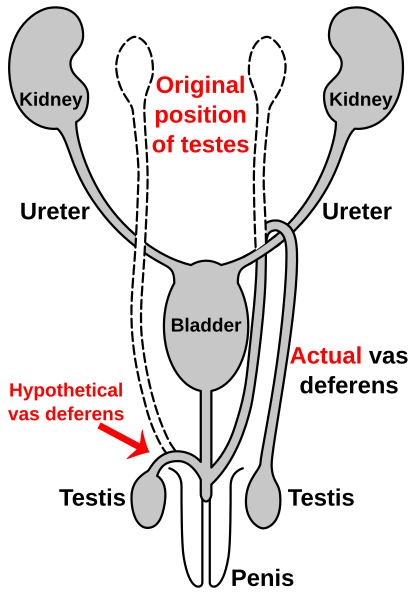
Route of the vas deferens
Similar to the laryngeal nerve in giraffes, the
Evidence from paleontology
When organisms die, they often
For fossilization to take place, the traces and remains of organisms must be quickly buried so that weathering and decomposition do not occur. Skeletal structures or other hard parts of the organisms are the most commonly occurring form of fossilized remains. There are also some trace "fossils" showing moulds, cast or imprints of some previous organisms.
As an animal dies, the organic materials gradually decay, such that the
. Fossilization can sometimes be a trace, an impression of a form. Examples include leaves and footprints, the fossils of which are made in layers that then harden.Fossil record
It is possible to decipher how a particular group of organisms evolved by arranging its fossil record in a chronological sequence. Such a sequence can be determined because fossils are mainly found in
A succession of animals and plants can also be seen from fossil discoveries. By studying the number and complexity of different fossils at different stratigraphic levels, it has been shown that older fossil-bearing rocks contain fewer types of fossilized organisms, and they all have a simpler structure, whereas younger rocks contain a greater variety of fossils, often with increasingly complex structures.[121]
For many years, geologists could only roughly estimate the ages of various strata and the fossils found. They did so, for instance, by estimating the time for the formation of sedimentary rock layer by layer. Today, by measuring the proportions of radioactive and stable elements in a given rock, the ages of fossils can be more precisely dated by scientists. This technique is known as radiometric dating.
Throughout the fossil record, many species that appear at an early stratigraphic level disappear at a later level. This is interpreted in evolutionary terms as indicating the times when species originated and became extinct. Geographical regions and climatic conditions have varied throughout Earth's history. Since organisms are adapted to particular environments, the constantly changing conditions favoured species that adapted to new environments through the mechanism of natural selection.
Extent of the fossil record
Despite the relative rarity of suitable conditions for fossilization, an estimated 250,000 fossil species have been named.
Limitations
The fossil record is an important source for scientists when tracing the evolutionary history of organisms. However, because of limitations inherent in the record, there are not fine scales of intermediate forms between related groups of species. This lack of continuous fossils in the record is a major limitation in tracing the descent of biological groups. When transitional fossils are found that show intermediate forms in what had previously been a gap in knowledge, they are often popularly referred to as "missing links".
There is a gap of about 100 million years between the beginning of the
Some of the reasons for the incompleteness of fossil records are:
- In general, the probability that an organism becomes fossilized is very low;
- Some species or groups are less likely to become fossils because they are soft-bodied;
- Some species or groups are less likely to become fossils because they live (and die) in conditions that are not favourable for fossilization;
- Many fossils have been destroyed through erosion and tectonic movements;
- Most fossils are fragmentary;
- Some evolutionary change occurs in populations at the limits of a species' ecological range, and as these populations are likely small, the probability of fossilization is lower (see punctuated equilibrium);
- Similarly, when environmental conditions change, the population of a species is likely to be greatly reduced, such that any evolutionary change induced by these new conditions is less likely to be fossilized;
- Most fossils convey information about external form, but little about how the organism functioned;
- Using present-day biodiversity as a guide, this suggests that the fossils unearthed represent only a small fraction of the large number of species of organisms that lived in the past.
Specific examples from paleontology
Evolution of the horse
Due to an almost-complete fossil record found in North American sedimentary deposits from the early
This evolutionary sequence starts with a small animal called
The probable course of development of horses from Hyracotherium to Equus (the modern horse) involved at least 12 genera and several hundred species. The major trends seen in the development of the horse to changing environmental conditions may be summarized as follows:
- Increase in size (from 0.4 m to 1.5 m — from 15 in to 60 in);
- Lengthening of limbs and feet;
- Reduction of lateral digits;
- Increase in length and thickness of the third digit;
- Increase in width of incisors;
- Replacement of premolars by molars; and
- Increases in tooth length, crown height of molars.
Fossilized plants found in different strata show that the marshy, wooded country in which Hyracotherium lived became gradually drier. Survival now depended on the head being in an elevated position for gaining a good view of the surrounding countryside, and on a high turn of speed for escape from
Transition from fish to amphibians
Prior to 2004, paleontologists had found fossils of amphibians with necks, ears, and four legs, in rock no older than 365 million years old. In rocks more than 385 million years old they could only find fish, without these amphibian characteristics. Evolutionary theory predicted that since amphibians evolved from fish, an intermediate form should be found in rock dated between 365 and 385 million years ago. Such an intermediate form should have many fish-like characteristics, conserved from 385 million years ago or more, but also have many amphibian characteristics as well. In 2004, an expedition to islands in the Canadian arctic searching specifically for this fossil form in rocks that were 375 million years old discovered fossils of Tiktaalik.[132] Some years later, however, scientists in Poland found evidence of fossilised tetrapod tracks predating Tiktaalik.[133]
Evidence from biogeography
Data about the presence or absence of species on various continents and islands (biogeography) can provide evidence of common descent and shed light on patterns of speciation.
Continental distribution
All organisms are adapted to their environment to a greater or lesser extent. If the abiotic and biotic factors within a
- Africa has Old World monkeys, apes, elephants, leopards, giraffes, and hornbills.
- South America has New World monkeys, cougars, jaguars, sloths, llamas, and toucans.
- Deserts in North and South America have native succulent (apart from Rhipsalis baccifera) which are native euphorbs that resemble cacti but are very different.[134]
Even greater differences can be found if
Other animal distribution examples include
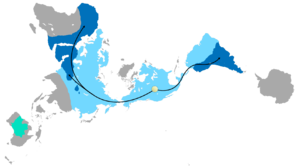
The same kinds of fossils are found from areas known to be adjacent to one another in the past but that, through the process of continental drift, are now in widely divergent geographic locations. For example, fossils of the same types of ancient amphibians, arthropods and ferns are found in South America, Africa, India, Australia and Antarctica, which can be dated to the Paleozoic Era, when these regions were united as a single landmass called Gondwana.[139]
Island biogeography
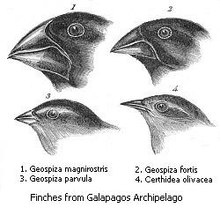

Types of species found on islands
Evidence from
Endemism
Many of the species found on remote islands are
Other types of endemism do not have to include, in the strict sense, islands. Islands can mean isolated lakes or remote and isolated areas. Examples of these would include the highlands of
Adaptive radiations
Oceanic islands are frequently inhabited by clusters of closely related species that fill a variety of
Ring species
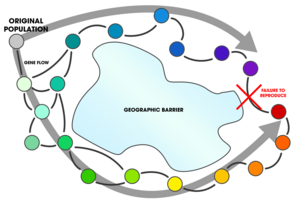
A ring species is a connected series of populations, each of which can interbreed with its neighbors, with at least two "end" populations which are too distantly related to interbreed, though with the potential for
Specific examples from biogeography
Distribution of Glossopteris
The combination of continental drift and evolution can sometimes be used to predict what will be found in the fossil record.
Metatherian distribution
The history of
Migration, isolation, and distribution of the camel
The history of the
Evidence from selection
Scientists have observed and documented a multitude of events where natural selection is in action. The most well known examples are antibiotic resistance in the medical field along with better-known laboratory experiments documenting evolution's occurrence. Natural selection is tantamount to common descent in that long-term occurrence and selection pressures can lead to the diversity of life on earth as found today. All adaptations—documented and undocumented changes concerned—are caused by natural selection (and a few other minor processes). It is well established that, "...natural selection is a ubiquitous part of speciation...",[166] and is the primary driver of speciation.[167]
Artificial selection and experimental evolution
Experimental evolution uses controlled experiments to test hypotheses and theories of evolution. In one early example, William Dallinger set up an experiment shortly before 1880, subjecting microbes to heat with the aim of forcing adaptive changes. His experiment ran for around seven years, and his published results were acclaimed, but he did not resume the experiment after the apparatus failed.[169]
A large-scale example of experimental evolution is
Invertebrates
Historical lead tolerance in Daphnia
A study of species of Daphnia and lead pollution in the 20th century predicted that an increase in lead pollution would lead to strong selection of lead tolerance. Researchers were able to use "resurrection ecology", hatching decades-old Daphnia eggs from the time when lakes were heavily polluted with lead. The hatchlings in the study were compared to current-day Daphnia, and demonstrated "dramatic fitness differences between old and modern phenotypes when confronted with a widespread historical environmental stressor". Essentially, the modern-day Daphnia were unable to resist or tolerate high levels of lead (this is due to the huge reduction of lead pollution in 21st century lakes). The old hatchlings, however, were able to tolerate high lead pollution. The authors concluded that "by employing the techniques of resurrection ecology, we were able to show clear phenotypic change over decades...".[173]
Peppered moths
A classic example was the phenotypic change, light-to-dark color adaptation, in the peppered moth, due to pollution from the Industrial Revolution in England.[174][175]
Microbes
Antimicrobial resistance
The development and spread of
All classes of microbes develop resistance: including fungi (
resistance). This is to be expected when considering that all life exhibits universal genetic code and is therefore subject to the process of evolution through its various mechanisms.Nylon-eating bacteria
Another example of organisms adapting to human-caused conditions are Nylon-eating bacteria: a strain of Flavobacterium that are capable of digesting certain byproducts of nylon 6 manufacturing. There is scientific consensus that the capacity to synthesize nylonase most probably developed as a single-step mutation that survived because it improved the fitness of the bacteria possessing the mutation. This is seen as a good example of evolution through mutation and natural selection that has been observed as it occurs and could not have come about until the production of nylon by humans.[177][178][179][180]
Plants and fungi
Monkeyflower radiation
Both subspecies Mimulus aurantiacus puniceus (red-flowered) and Mimulus aurantiacus australis (yellow-flowered) of monkeyflowers are isolated due to the preferences of their hummingbird and hawkmoth pollinators. The radiation of M. aurantiacus subspecies are mostly yellow colored; however, both M. a. ssp. puniceus and M. a. ssp. flemingii are red. Phylogenetic analysis suggests two independent origins of red-colored flowers that arose due to cis-regulatory mutations in the gene MaMyb2 that is present in all M. aurantiacus subspecies. Further research suggested that two independent mutations did not take place, but one MaMyb2 allele was transferred via introgressive hybridization.[181]
Radiotrophic fungi
Vertebrates
Guppies
While studying guppies (Poecilia reticulata) in Trinidad, biologist John Endler detected selection at work on the fish populations. To rule out alternative possibilities, Endler set up a highly controlled experiment to mimic the natural habitat by constructing ten ponds within a laboratory greenhouse at Princeton University. Each pond contained gravel to exactly match that of the natural ponds. After capturing a random sample of guppies from ponds in Trinidad, he raised and mixed them to create similar genetically diverse populations and measured each fish (spot length, spot height, spot area, relative spot length, relative spot height, total patch area, and standard body lengths). For the experiment he added Crenicichla alta (P. reticulata's main predator) in four of the ponds, Rivulus hartii (a non-predator fish) in four of the ponds, and left the remaining two ponds empty with only the guppies. After 10 generations, comparisons were made between each pond's guppy populations and measurements were taken again. Endler found that the populations had evolved dramatically different color patterns in the control and non-predator pools and drab color patterns in the predator pool. Predation pressure had caused a selection against standing out from background gravel.[184]
In parallel, during this experiment, Endler conducted a field experiment in Trinidad where he caught guppies from ponds where they had predators and relocated them to ponds upstream where the predators did not live. After 15 generations, Endler found that the relocated guppies had evolved dramatic and colorful patterns. Essentially, both experiments showed convergence due to similar selection pressures (i.e. predator selection against contrasting color patterns and sexual selection for contrasting color patterns).[184]
In a later study by David Reznick, the field population was examined 11 years later after Endler relocated the guppies to high streams. The study found that the populations has evolved in a number of different ways: bright color patterns, late maturation, larger sizes, smaller litter sizes, and larger offspring within litters.[185] Further studies of P. reticulata and their predators in the streams of Trinidad have indicated that varying modes of selection through predation have not only changed the guppies color patterns, sizes, and behaviors, but their life histories and life history patterns.[186]
Humans
Natural selection is observed in contemporary human populations, with recent findings demonstrating that the population at risk of the severe debilitating disease
A well known example of selection occurring in human populations is lactose tolerance. Lactose intolerance is the inability to metabolize lactose, because of a lack of the required enzyme lactase in the digestive system. The normal mammalian condition is for the young of a species to experience reduced lactase production at the end of the weaning period (a species-specific length of time). In humans, in non-dairy consuming societies, lactase production usually drops about 90% during the first four years of life, although the exact drop over time varies widely.[190] Lactase activity persistence in adults is associated with two polymorphisms: C/T 13910 and G/A 22018 located in the MCM6 gene.[191] This gene difference eliminates the shutdown in lactase production, making it possible for members of these populations to continue consumption of raw milk and other fresh and fermented dairy products throughout their lives without difficulty. This appears to be an evolutionarily recent (around 10,000 years ago [and 7,500 years ago in Europe][192]) adaptation to dairy consumption,[193] and has occurred independently in both northern Europe and east Africa in populations with a historically pastoral lifestyle.[194][195]
Italian wall lizards
In 1971, ten adult specimens of
In the 1990s, scientists returned to Pod Mrčaru and found that the lizards there differed greatly from those on Kopište. While mitochondrial DNA analyses have verified that P. sicula currently on Mrčaru are genetically very similar to the Kopište source population,[196] the new Mrčaru population of P. sicula had a larger average size, shorter hind limbs, lower maximal sprint speed and altered response to simulated predatory attacks compared to the original Kopište population.[197] These changes were attributed to "relaxed predation intensity" and greater protection from vegetation on Mrčaru.[197]
In 2008, further analysis revealed that the Mrčaru population of P. sicula have significantly different head morphology (longer, wider, and taller heads) and increased bite force compared to the original Kopište population.
Another difference found between the two populations was the discovery, in the Mrčaru lizards, of
PAH resistance in killifish
A similar study was also done regarding the polycyclic aromatic hydrocarbons (PAHs) that pollute the waters of the Elizabeth River in Portsmouth, Virginia. This chemical is a product of creosote, a type of tar. The Atlantic killifish (Fundulus heteroclitus) has evolved a resistance to PAHs involving the AHR gene (the same gene involved in the tomcods). This particular study focused on the resistance to "acute toxicity and cardiac teratogenesis" caused by PAHs. that mutated within the tomcods in the Hudson River.[200]
PCB resistance in codfish
An example involving the direct observation of gene modification due to selection pressures is the resistance to PCBs in codfish. After
Urban wildlife
White Sands lizards

Animals that exhibit
New Mexico's White Sands are a recent geologic formation (approximately 6000 years old[210] to possibly 2000 years old[207]). This recent origin of these gypsum sand dunes suggests that species exhibiting lighter-colored variations have evolved in a relatively short time frame. The three lizard species previously mentioned have been found to display variable social signal coloration in coexistence with their ecotonal variants.[210] Not only have the three species convergently evolved their lighter variants due to the selection pressures from the environment, they have also evolved ecomorphological differences: morphology, behavior (in is case, escape behavior), and performance (in this case, sprint speed) collectively.[209] Roches' work found surprising results in the escape behavior of H. maculata and S. undulatus. When dark morphs were placed on white sands, their startle response was significantly diminished. This result could be due to varying factors relating to sand temperature or visual acuity; however, regardless of the cause, "…failure of mismatched lizards to sprint could be maladaptive when faced with a predator".[209]
Evidence from speciation
Speciation is the evolutionary process by which new biological species arise. Biologists research species using different theoretical frameworks for what constitutes a species (see
Research documenting speciation is abundant. Biologists have documented numerous examples of speciation in nature—with evolution having produced far more species than any observer would consider necessary. For example, there are well over 350,000 described species of beetles.[216] Examples of speciation come from the observations of island biogeography and the process of adaptive radiation, both explained previously. Evidence of common descent can also be found through paleontological studies of speciation within geologic strata. The examples described below represent different modes of speciation and provide strong evidence for common descent. It is important to acknowledge that not all speciation research directly observes divergence from "start-to-finish". This is by virtue of research delimitation and definition ambiguity, and occasionally leads research towards historical reconstructions. In light of this, examples abound, and the following are by no means exhaustive—comprising only a small fraction of the instances observed. Once again, take note of the established fact that, "...natural selection is a ubiquitous part of speciation...",[166] and is the primary driver of speciation,[167] so; hereinafter, examples of speciation will often interdepend and correspond with selection.
Fossils
Limitations exist within the fossil record when considering the concept of what constitutes a species. Paleontologists largely rely on a different framework: the morphological species concept.
Globorotalia
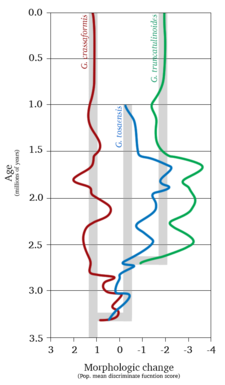
Extensive research on the planktonic foraminifer Globorotalia truncatulinoides has provided insight into paleobiogeographical and paleoenvironmental studies alongside the relationship between the environment and evolution. In an extensive study of the paleobiogeography of G. truncatulinoides, researchers found evidence that suggested the formation of a new species (via the sympatric speciation framework). Cores taken of the sediment containing the three species G. crassaformis, G. tosaensis, and G. truncatulinoides found that before 2.7 Ma, only G. crassaformis and G. tosaensis existed. A speciation event occurred at that time, whereby intermediate forms existed for quite some time. Eventually G. tosaensis disappears from the record (suggesting extinction) but exists as an intermediate between the extant G. crassaformis and G. truncatulinoides. This record of the fossils also matched the already existing phylogeny constructed by morphological characters of the three species.[219] See figure 6a.
Radiolaria
In a large study of five species of
Rhizosolenia
Researchers conducted measurements on 5,000 Rhizosolenia (a planktonic diatom) specimens from eight sedimentary cores in the Pacific Ocean. The core samples spanned two million years and were chronologized using sedimentary magnetic field reversal measurements. All the core samples yielded a similar pattern of divergence: with a single lineage (R. bergonii) occurring before 3.1 Mya and two morphologically distinct lineages (daughter species: R. praebergonii) appearing after. The parameters used to measure the samples were consistent throughout each core.[222] An additional study of the daughter species R. praebergonii found that, after the divergence, it invaded the Indian Ocean.[217][223]
Turborotalia
A recent study was conducted involving the planktonic foraminifer Turborotalia. The authors extracted "51 stratigraphically ordered samples from a site within the oceanographically stable tropical North Pacific gyre". Two hundred individual species were examined using ten specific morphological traits (size, compression index, chamber aspect ratio, chamber inflation, aperture aspect ratio, test height, test expansion, umbilical angle, coiling direction, and the number of chambers in the final whorl). Utilizing multivariate statistical clustering methods, the study found that the species continued to evolve non-directionally within the Eocene from 45 Ma to about 36 Ma. However, from 36 Ma to approximately 34 Ma, the stratigraphic layers showed two distinct clusters with significantly defining characteristics distinguishing one another from a single species. The authors concluded that speciation must have occurred and that the two new species were descended from the prior species.[224]
Vertebrates
There exists evidence for vertebrate speciation despite limitations imposed by the fossil record. Studies have been conducted documenting similar patterns seen in marine invertebrates.[217] For example, extensive research documenting rates of morphological change, evolutionary trends, and speciation patterns in small mammals has significantly contributed to the scientific literature.[225]
A study of four mammalian genera: Hyopsodus, Pelycodus, Haplomylus (three from the Eocene), and Plesiadapis (from the Paleocene) found that—through a large number of stratigraphic layers and specimen sampling—each group exhibited, "gradual phyletic evolution, overall size increase, iterative evolution of small species, and character divergence following the origin of each new lineage".[226] The authors of this study concluded that speciation was discernible. In another study concerning morphological trends and rates of evolution found that the European arvicolid rodent radiated into 52 distinct lineages over a time frame of 5 million years while documenting examples of phyletic gradualism, punctuation, and stasis.[227]
Invertebrates
Drosophila melanogaster

William R. Rice and George W. Salt found experimental evidence of
Gall wasps
A study of the
In an additional study, the researchers studied two gall wasp species B. treatae and Disholcaspis quercusvirens and found strong morphological and behavioral variation among host-associated populations. This study further confounded prerequisites to speciation.[232]
Hawthorn fly
One example of evolution at work is the case of the hawthorn fly,
London Underground mosquito
The London Underground mosquito is a species of mosquito in the genus Culex found in the London Underground. It evolved from the overground species Culex pipiens. This mosquito, although first discovered in the London Underground system, has been found in underground systems around the world. It is suggested that it may have adapted to human-made underground systems since the last century from local above-ground Culex pipiens,[239] although more recent evidence suggests that it is a southern mosquito variety related to Culex pipiens that has adapted to the warm underground spaces of northern cities.[240]
The two species have very different behaviours,
The genetic data indicates that the molestus form in the London Underground mosquito appears to have a common ancestry, rather than the population at each station being related to the nearest aboveground population (i.e. the pipiens form). Byrne and Nichols' working hypothesis was that adaptation to the underground environment had occurred locally in London only once. These widely separated populations are distinguished by very minor genetic differences, which suggest that the molestus form developed: a single mtDNA difference shared among the underground populations of ten Russian cities;
Snapping shrimp and the isthmus of Panama
Debate exists determining when the isthmus of Panama closed. Much of the evidence supports a closure approximately 2.7 to 3.5 mya using "...multiple lines of evidence and independent surveys".[244] However, a recent study suggests an earlier, transient bridge existed 13 to 15 mya.[245] Regardless of the timing of the isthmus closer, biologists can study the species on the Pacific and Caribbean sides in, what has been called, "one of the greatest natural experiments in evolution."[244] Studies of snapping shrimp in the genus Alpheus have provided direct evidence of allopatric speciation events,[246] and contributed to the literature concerning rates of molecular evolution.[247] Phylogenetic reconstructions using "multilocus datasets and coalescent-based analytical methods" support the relationships of the species in the group[244] and molecular clock techniques support the separation of 15 pairs of Alpheus species between 3 and 15 million years ago.[247]
Plants
The botanist Verne Grant pioneered the field of plant speciation with his research and major publications on the topic.[248] As stated before, many biologists rely on the biological species concept, with some modern researchers utilizing the phylogenetic species concept. Debate exists in the field concerning which framework should be applied in the research.[248] Regardless, reproductive isolation is the primary role in the process of speciation and has been studied extensively by biologists in their respective disciplines.
Both hybridization and polyploidy have also been found to be major contributors to plant speciation.
Hybridization between two different species sometimes leads to a distinct phenotype. This phenotype can also be fitter than the parental lineage and as such, natural selection may then favor these individuals. Eventually, if reproductive isolation is achieved, it may lead to a separate species. However, reproductive isolation between hybrids and their parents is particularly difficult to achieve and thus hybrid speciation is considered a rare event. However, hybridization resulting in reproductive isolation is considered an important means of speciation in plants,[251] since polyploidy (having more than two copies of each chromosome) is tolerated in plants more readily than in animals.[252][253]
Polyploidy is a mechanism that has caused many rapid speciation events in sympatry because offspring of, for example, tetraploid x diploid matings often result in triploid sterile progeny.[257] Not all polyploids are reproductively isolated from their parental plants, and gene flow may still occur for example through triploid hybrid x diploid matings that produce tetraploids, or matings between meiotically unreduced gametes from diploids and gametes from tetraploids. It has been suggested that many of the existing plant and most animal species have undergone an event of polyploidization in their evolutionary history.[254][258] Reproduction of successful polyploid species is sometimes asexual, by parthenogenesis or apomixis, as for unknown reasons many asexual organisms are polyploid. Rare instances of polyploid mammals are known, but most often result in prenatal death.
Researchers consider reproductive isolation as key to speciation.[259] A major aspect of speciation research is to determine the nature of the barriers that inhibit reproduction. Botanists often consider the zoological classifications of prezygotic and postzygotic barriers as inadequate.[259] The examples provided below give insight into the process of speciation.
Mimulus peregrinus
The creation of a new allopolyploid species of monkeyflower (Mimulus peregrinus) was observed on the banks of the Shortcleuch Water—a river in Leadhills, South Lanarkshire, Scotland. Parented from the cross of the two species Mimulus guttatus (containing 14 pairs of chromosomes) and Mimulus luteus (containing 30-31 pairs from a chromosome duplication), M. peregrinus has six copies of its chromosomes (caused by the duplication of the sterile hybrid triploid). Due to the nature of these species, they have the ability to self-fertilize. Because of its number of chromosomes it is not able to pair with M. guttatus, M. luteus, or their sterile triploid offspring. M. peregrinus will either die, producing no offspring, or reproduce with itself effectively leading to a new species.[260][261]
Raphanobrassica
Senecio (groundsel)
The Welsh groundsel is an allopolyploid, a plant that contains sets of chromosomes originating from two different species. Its ancestor was Senecio × baxteri, an infertile hybrid that can arise spontaneously when the closely related groundsel (Senecio vulgaris) and Oxford ragwort (Senecio squalidus) grow alongside each other. Sometime in the early 20th century, an accidental doubling of the number of chromosomes in an S. × baxteri plant led to the formation of a new fertile species.[264][265]
The
Other hybrids descended from the same two parents are known. Some are infertile, such as S. x baxteri. Other fertile hybrids are also known, including S. vulgaris var. hibernicus, now common in Britain, and the allohexaploid S. cambrensis, which according to molecular evidence probably originated independently at least three times in different locations. Morphological and genetic evidence support the status of S. eboracensis as separate from other known hybrids.[266]
Thale cress

Kirsten Bomblies et al. from the Max Planck Institute for Developmental Biology discovered two genes in the thale cress plant, Arabidopsis thaliana. When both genes are inherited by an individual, it ignites a reaction in the hybrid plant that turns its own immune system against it. In the parents, the genes were not detrimental, but they evolved separately to react defectively when combined.[267] To test this, Bomblies crossed 280 genetically different strains of Arabidopsis in 861 distinct ways and found that 2 percent of the resulting hybrids were necrotic. Along with allocating the same indicators, the 20 plants also shared a comparable collection of genetic activity in a group of 1,080 genes. In almost all of the cases, Bomblies discovered that only two genes were required to cause the autoimmune response. Bomblies looked at one hybrid in detail and found that one of the two genes belonged to the NB-LRR class, a common group of disease resistance genes involved in recognizing new infections. When Bomblies removed the problematic gene, the hybrids developed normally.[267] Over successive generations, these incompatibilities could create divisions between different plant strains, reducing their chances of successful mating and turning distinct strains into separate species.[268]
Tragopogon (salsify)

T. dubius and T. pratensis mated in Europe but were never able to hybridize. A study published in March 2011 found that when these two plants were introduced to North America in the 1920s, they mated and doubled the number of chromosomes in there hybrid Tragopogon miscellus allowing for a "reset" of its genes, which in turn, allows for greater genetic variation. Professor Doug Soltis of the University of Florida said, "We caught evolution in the act…New and diverse patterns of gene expression may allow the new species to rapidly adapt in new environments".[270][271]
Vertebrates
Blackcap
The bird species,
Mollies
The shortfin molly (Poecilia mexicana) is a small fish that lives in the Sulfur Caves of Mexico. Years of study on the species have found that two distinct populations of mollies—the dark interior fish and the bright surface water fish—are becoming more genetically divergent.[274] The populations have no obvious barrier separating the two; however, it was found that the mollies are hunted by a large water bug (Belostoma spp). Tobler collected the bug and both types of mollies, placed them in large plastic bottles, and put them back in the cave. After a day, it was found that, in the light, the cave-adapted fish endured the most damage, with four out of every five stab-wounds from the water bugs sharp mouthparts. In the dark, the situation was the opposite. The mollies' senses can detect a predator's threat in their own habitats, but not in the other ones. Moving from one habitat to the other significantly increases the risk of dying. Tobler plans on further experiments, but believes that it is a good example of the rise of a new species.[275]
Polar bear
Natural selection, geographic isolation, and speciation in progress are illustrated by the relationship between the polar bear (Ursus maritimus) and the brown bear (Ursus arctos). Considered separate species throughout their ranges;[276] however, it has been documented that they possess the capability to interbreed and produce fertile offspring. This introgressive hybridization has occurred both in the wild and in captivity and has been documented[277] and verified with DNA testing.[278] The oldest known fossil evidence of polar bears dates around 130,000 to 110,000 years ago;[279] however, molecular data has revealed varying estimates of divergence time. Mitochondrial DNA analysis has given an estimate of 150,000 years ago[279] while nuclear genome analysis has shown an approximate divergence of 603,000 years ago.[280] Recent research using the complete genomes (rather than mtDNA or partial nuclear genomes) establishes the divergence of polar and brown bears between 479 and 343 thousand years ago.[281] Despite the differences in divergence rates, molecular research suggests the sister species have undergone a highly complex process of speciation and admixture between the two.[282]
The polar bear has acquired anatomical and physiological differences from the brown bear that allow it to comfortably survive in conditions that the brown bear likely could not. Notable examples include the ability to swim sixty miles or more at a time in freezing waters, fur that blends with the snow, and to stay warm in the arctic environment, an elongated neck that makes it easier to keep their heads above water while swimming, and oversized and heavy-matted webbed feet that act as paddles when swimming. It has also evolved small papillae and vacuole-like suction cups on the soles to make them less likely to slip on the ice, alongside smaller ears for a reduction of heat loss, eyelids that act like sunglasses, accommodations for their all-meat diet, a large stomach capacity to enable opportunistic feeding, and the ability to fast for up to nine months while recycling their urea.[283][284]
Evidence from coloration

Mimicry and aposematism
Bates and Müller described forms of mimicry that now carry their names, based on their observations of tropical butterflies. These highly specific patterns of coloration are readily explained by natural selection, since predators such as birds which hunt by sight will more often catch and kill insects that are less good mimics of distasteful models than those that are better mimics; but the patterns are otherwise hard to explain.
Camouflage
In 1889, Wallace noted that
Evidence from behavior
Some primitive reflexes are critical for the survival of neonates. There is evidence confirming that closely related species share more similar primitive reflexes, such as the type of fur-grasping in primates and their relationship to manual dexterity. The exact selection pressures for their development are not fully determined and some reflexes are understood to have evolved multiple times independently (convergent evolution).[295]
Evidence from mathematical modeling and simulation
Computer science allows the iteration of self-changing complex systems to be studied, allowing a mathematical understanding of the nature of the processes behind evolution; providing evidence for the hidden causes of known evolutionary events. The evolution of specific cellular mechanisms like spliceosomes that can turn the cell's genome into a vast workshop of billions of interchangeable parts that can create tools that create us can be studied for the first time in an exact way.
"It has taken more than five decades, but the electronic computer is now powerful enough to simulate evolution",[296] assisting bioinformatics in its attempt to solve biological problems.
Computational evolutionary biology has enabled researchers to trace the evolution of a large number of organisms by measuring changes in their DNA, rather than through physical taxonomy or physiological observations alone. It has compared entire genomes permitting the study of more complex evolutionary events, such as gene duplication, horizontal gene transfer, and the prediction of factors important in speciation. It has also helped build complex computational models of populations to predict the outcome of the system over time and track and share information on an increasingly large number of species and organisms.
Future endeavors are to reconstruct a now more complex tree of life.
Christoph Adami, a professor at the Keck Graduate Institute made this point in Evolution of biological complexity:
- To make a case for or against a trend in the evolution of complexity in biological evolution, complexity must be both rigorously defined and measurable. A recent information-theoretic (but intuitively evident) definition identifies genomic complexity with the amount of information a sequence stores about its environment. We investigate the evolution of genomic complexity in populations of digital organisms and monitor in detail the evolutionary transitions that increase complexity. We show that, because natural selection forces genomes to behave as a natural "Maxwell Demon", within a fixed environment, genomic complexity is forced to increase.[297]
David J. Earl and Michael W. Deem—professors at Rice University made this point in Evolvability is a selectable trait:
- Not only has life evolved, but life has evolved to evolve. That is, correlations within protein structure have evolved, and mechanisms to manipulate these correlations have evolved in tandem. The rates at which the various events within the hierarchy of evolutionary moves occur are not random or arbitrary but are selected by Darwinian evolution. Sensibly, rapid or extreme environmental change leads to selection for greater evolvability. This selection is not forbidden by causality and is strongest on the largest-scale moves within the mutational hierarchy. Many observations within evolutionary biology, heretofore considered evolutionary happenstance or accidents, are explained by selection for evolvability. For example, the vertebrate immune system shows that the variable environment of antigens has provided selective pressure for the use of adaptable codons and low-fidelity polymerases during influenza A.[298]
"Computer simulations of the evolution of linear sequences have demonstrated the importance of recombination of blocks of sequence rather than point mutagenesis alone. Repeated cycles of point mutagenesis, recombination, and selection should allow in vitro molecular evolution of complex sequences, such as proteins."[299] Evolutionary molecular engineering, also called directed evolution or in vitro molecular evolution involves the iterated cycle of mutation, multiplication with recombination, and selection of the fittest of individual molecules (proteins, DNA, and RNA). Natural evolution can be relived showing us possible paths from catalytic cycles based on proteins to based on RNA to based on DNA.[299][300][301][302]
The Avida artificial life software platform has been used to explore common descent and natural selection.[303][304] It has been used to demonstrate that natural selection can favor altruism, something that had been predicted but is difficult to test empirically. At the higher replication rates allowed by the platform it becomes observable.[305]
See also
References
- ISBN 978-0-87969-608-5.
- S2CID 4270111.
- ^ "Eukaryotes". www.tolweb.org. Retrieved 23 June 2018.
- ^ Max, Edward (5 May 2003). "Plagiarized Errors and Molecular Genetics". The Talk Origins Archive. Retrieved 4 May 2018.
- ISBN 978-0-87893-189-7.
- ISBN 978-0-19-854980-2.
- PMID 14602077.
- ISBN 0-19-850968-5.
- PMID 1973846.
- PMID 11170892.
- ^
Cooper, G.M.; Brudno, M.; Green, E.D.; Batzoglou, S.; Sidow, A. (2003). "Quantitative Estimates of Sequence Divergence for Comparative Analyses of Mammalian Genomes". Genome Res. 13 (5): 813–20. PMID 12727901.
- ^ The picture labeled "Human Chromosome 2 and its analogs in the apes" in the article Comparison of the Human and Great Ape Chromosomes as Evidence for Common Ancestry Archived 23 July 2011 at the Wayback Machine is literally a picture of a link in humans that links two separate chromosomes in the nonhuman apes creating a single chromosome in humans. Also, while the term originally referred to fossil evidence, this too is a trace from the past corresponding to some living beings that, when alive, physically embodied this link.
- ^ The New York Times report Still Evolving, Human Genes Tell New Story, based on A Map of Recent Positive Selection in the Human Genome, states the International HapMap Project is "providing the strongest evidence yet that humans are still evolving" and details some of that evidence.
- PMID 270744.
- PMID 2112744.
- ISBN 978-0-8153-3218-3.
- ^ "Converging Evidence for Evolution." Archived 1 December 2010 at the Wayback Machine Phylointelligence: Evolution for Everyone. 26 November 2010.
- PMID 10833048.
- ^ PMID 25391452.
- PMID 3909943.
- PMID 17568002. Retrieved 23 June 2018.
- ^ "What is Junk DNA?". News-Medical.net. 7 May 2010. Retrieved 23 June 2018.
- S2CID 22081618.
- PMID 16891155.
- ^ MacAndrew, Alec. Human Chromosome 2 is a fusion of two ancestral chromosomes. Accessed 18 May 2006.
- ^ Evidence of Common Ancestry: Human Chromosome 2 (video) 2007
- PMID 7063861.
- ^ Human and Ape Chromosomes Archived 23 July 2011 at the Wayback Machine; accessed 8 September 2007.
- S2CID 1441285.
- ^ PMID 1924367.
- ^ a b Amino acid sequences in cytochrome c proteins from different species Archived 28 December 2013 at the Wayback Machine, adapted from Strahler, Arthur; Science and Earth History, 1997. page 348.
- ISBN 978-1-4051-5089-7.
- ^ a b Theobald, Douglas (2004). "29+ Evidences for Macroevolution; Protein functional redundancy". The Talk Origins Archive.
- PMID 11264397.
- PMID 10482547.
- PMID 10655035.
- PMID 15044706.
- PMID 6181510.
- PMID 10468595.
- ISBN 978-1-85828-946-5.
- S2CID 4456458.
- ^ Andrews, Roy Chapman (3 June 1921). "A Remarkable Case of External Hind Limbs in a Humpback Whale" (PDF). American Museum Novitates. Archived from the original (PDF) on 13 June 2007. Retrieved 26 February 2013.
- ^ S2CID 27868367
- ^ S2CID 29258934
- ^ Lambert, Katie. (29 October 2007) HowStuffWorks "How Atavisms Work". Animals.howstuffworks.com. Retrieved on 2011-12-06.
- ^ S2CID 40851098
- ^ Raynaud, A. (1977), Somites and early morphogenesis in reptile limbs. In Vertebrate Limb and Somite Morphogenesis, Cambridge University Press, London, pp. 373–386
- ^ Tabuchi, Hiroko (2006), Dolphin May Have 'Remains' of Legs, Livescience.com
- PMID 15373256.
- ^ Simpson, G. G. (1951), Horses: The story of the horse family in the modern world and through sixty million years of evolution, Oxford University Press
- PMID 6373560
- PMID 17438282
- PMID 16488870
- S2CID 962571
- ^ JSTOR 1367290
- JSTOR 1366113
- S2CID 33397424
- ^ Castle, William E. (1906), The origin of a polydactylous race of guinea-pigs (49 ed.), Carnegie Institution of Washington
- S2CID 12112693.
- ^ ISBN 978-0-87893-189-7.
- ^ a b 29+ Evidences for Macroevolution: Part 1. Talkorigins.org. Retrieved on 6 December 2011.
- ISBN 978-0-670-02053-9.
- ^ Darwin, Charles (1859). On the Origin of Species. John Murray. p. 420.
- JSTOR 2992299.
- PMID 14519199.
- PMID 20525607. Archived from the original(PDF) on 27 September 2013.
- .
- ^ Kluge 1999
- ^ Laurin 2000
- ^ Fitzhugh 2006, p. 31
- ^ Kluge 1999, p. 432
- ISBN 978-1-933143-15-6.
- S2CID 85573136
- PMID 15985435
- PMID 12125757
- PMID 21665616
- S2CID 17169760
- S2CID 198153247
- PMID 21646167
- S2CID 19533785
- PMID 21642137
- S2CID 8448387
- ^ ISBN 978-0-670-02053-9.
- ISBN 978-0-19-512234-3.
- ^ "Example 1: Living whales and dolphins found with hindlimbs". Douglas Theobald. Retrieved 20 March 2011.
- PMID 18460088
- ISBN 978-0-670-02053-9.
- PMID 21113002
- S2CID 7504517
- ISBN 9780674330191
- ISBN 978-0-670-02053-9.
- S2CID 12494187
- ^ Michael G. Glasspool (1982), Atlas of Ophthalmology (1 ed.), MTP Press Unlimited, p. 9
- PMID 16856596
- S2CID 85768695
- S2CID 85991116
- PMID 18686195
- S2CID 22907605
- PMID 11842399
- S2CID 33141711
- PMID 12631698
- PMID 12826614
- S2CID 21055062
- PMID 19568443
- PMID 22294879
- S2CID 41665533
- PMID 18985616
- PMID 18351802
- Myers, P. Z. (2008), "Reproductive history writ in the genome", Pharyngula, ScienceBlogs
- PMID 9143876
- JSTOR 2441055
- ISBN 978-1-4051-0345-9.
- ^ ISBN 978-1-4165-9478-9.
- ISBN 978-0-19-506932-7.
- ^ "Confessions of a Darwinist". Niles Eldredge. Retrieved 22 June 2010.
- ISBN 978-0-670-02053-9.
- ^ Donald R. Prothero (2013), Bringing Fossils to Life: An Introduction to Paleobiology (3rd ed.), Columbia University Press, p. 21
- ^ "Frequently Asked Questions". The Natural History Museum of Los Angeles County Foundation. Archived from the original on 11 March 2011. Retrieved 21 February 2011.
- ISBN 978-0-521-55450-3.
- ^ Schadewald, Robert J. (1982). "Six "Flood" Arguments Creationists Can't Answer". Creation Evolution Journal. 3: 12–17. Archived from the original on 19 September 2015. Retrieved 21 February 2011.
- ^
- ISBN 0-618-00583-8
- S2CID 24895681.
- S2CID 4401274
- ISBN 978-3-923871-35-3.
- ^ Academy of Natural Sciences - Joseph Leidy - American Horses Archived 5 March 2012 at the Wayback Machine
- ISBN 978-0-375-42447-2.
- S2CID 4428903.
- ^ Cota-Sánchez, J. Hugo; Bomfim-Patrício, Márcia C. (2010). "Seed morphology, polyploidy and the evolutionary history of the epiphytic cactus Rhipsalis baccifera (Cactaceae)" (PDF). Polibotanica. 29: 107–129. Archived from the original (PDF) on 6 October 2018. Retrieved 28 February 2013.
- ISBN 978-0-19-550870-3.
- ISBN 9780643092044.
- ^ "Polar Bears/Habitat & Distribution". SeaWorld Parks & Entertainment. Archived from the original on 25 October 2013. Retrieved 21 February 2011.
- ^ "Sirenians of the World". Save the Manatee Club. Archived from the original on 26 October 2017. Retrieved 21 February 2011.
- ^ Continental Drift and Evolution Archived 11 February 2006 at the Wayback Machine. Biology.clc.uc.edu (25 March 2001). Retrieved on 2011-12-06.
- ^ ISBN 978-0-670-02053-9.
- ISBN 978-1-58834-073-3.
- ^ Burdick, Alan (25 March 2007). "The Wonder Land of Socotra, Yemen". ALAN BURDICK. Retrieved 8 July 2010.
- ^ "Tuatara". New Zealand Ecology: Living Fossils. TerraNature Trust. 2004. Retrieved 10 November 2006.
- ^ "Facts about tuatara". Conservation: Native Species. Threatened Species Unit, Department of Conservation, Government of New Zealand. Archived from the original on 30 September 2007. Retrieved 10 February 2007.
- ^ "New Caledonia's most wanted". Archived from the original on 29 September 2018. Retrieved 8 July 2010.
- ^ "Giant bushy-tailed cloud rat (Crateromys schadenbergi)". Archived from the original on 30 June 2010. Retrieved 8 July 2010.
- ^ Rabor, D.S. (1986). Guide to Philippine Flora and Fauna. Natural Resources Management Centre, Ministry of Natural Resources and University of the Philippines.
- ISBN 978-0816504367.
- ^ Schofield, James (27 July 2001). "Lake Baikal's Vanishing Nerpa Seal". The Moscow Times. Retrieved 27 September 2007.
- ^ Baldwin, B. G.; Robichaux, R. H. (1995). "Historical biogeography and ecology of the Hawaiian silversword alliance (Asteraceae). New molecular phylogenetic perspectives". In Wagner, W. L.; Funk, V. A. (eds.). Hawaiian biogeography: evolution on a hotspot archipelago. Washington: Smithsonian Institution Press. pp. 259–287.
- ^ "Adaptive Radiation and Hybridization in the Hawaiian Silversword Alliance". University of Hawaii Botany Department.
- ISBN 978-1-85828-946-5.
- S2CID 7108000. Archived from the original(PDF) on 29 May 2020. Retrieved 4 February 2018.
- ISBN 978-0-618-00583-3.
- ISBN 1-58834-156-9
- ISBN 978-0-670-02053-9.
- ^ Pioneer Productions (19 January 2010). "Episode Guide". How The Earth Was Made. Season 2. Episode 8. History channel.
- S2CID 18032860.
- PMID 20668664.
- S2CID 32304737.
- S2CID 2319361.
- .
- ISBN 978-1-57607-422-0
- ^
Goin, F.J.; Reguero, M.A.; Pascual, R.; von Koenigswald, W.; Woodburne, M.O.; Case, J.A.; Marenssi, S.A.; Vieytes, C.; Vizcaíno, S.F. (2006). "First gondwanatherian mammal from Antarctica". Geological Society, London, Special Publications. 258 (1): 135–144. S2CID 129493664.
- ISBN 978-0-8018-7135-1.
- ^ S2CID 10168162
- ^ ISBN 978-0-87893-089-0
- OCLC 183148564.
- ^ Haas, J. W. Jr. (June 2000). "The Rev. Dr. William H. Dallinger F.R.S.: Early Advocate of Theistic Evolution and Foe of Spontaneous Generation". Perspectives on Science and Christian Faith. 52: 107–117. Retrieved 15 June 2010.
- ^ a b Le Page, Michael (16 April 2008). "NS:bacteria make major evolutionary shift in the lab". New Scientist. Retrieved 9 July 2012.
- PMID 18524956.
- PMID 8041701.
- S2CID 205124469
- .
- S2CID 25407417.
- PMID 2501419.
- ^ Thwaites, W.M. (Summer 1985). "New Proteins Without God's Help". Creation Evolution Journal. 5 (2): 1–3.
- ^ Evolution and Information: The Nylon Bug. Nmsr.org. Retrieved on 6 December 2011.
- ^ Than, Ker (23 September 2005). "Why scientists dismiss 'intelligent design". NBC News.
- Only a Theory: Evolution and the Battle for America's Soul(2008) pp. 80–82
- PMID 26311673
- ^ Science News, Dark Power: Pigment seems to put radiation to good use, Week of 26 May 2007; Vol. 171, No. 21, p. 325 by Davide Castelvecchi
- PMID 17520016.
- ^ PMID 28563214.
- S2CID 18480502.
- PMID 28581096.
- ^ Medical Research Council (UK) (21 November 2009). "Brain Disease 'Resistance Gene' Evolves in Papua New Guinea Community; Could Offer Insights Into CJD". Science Daily (online). Science News. Retrieved 22 November 2009.
- PMID 19923577.
- PMID 19858476.
- ^ Soy and Lactose Intolerance Wayback: Soy Nutrition
- S2CID 21430931.
- PMID 23903732.
- PMID 14616060.
- ^ Harriet, Coles (20 January 2007). "The lactase gene in Africa: Do you take milk?". The Human Genome, Wellcome Trust. Archived from the original on 29 September 2008. Retrieved 18 July 2008.
- PMID 17159977.
- ^ PMID 18344323.
- ^ S2CID 55765174.
- ^ "Lizards Rapidly Evolve After Introduction to Island". National Geographic. 21 April 2008. Archived from the original on 23 April 2008.
- ^ Myers, PZ (23 April 2008). "Still just a lizard". ScienceBlogs.
- PMID 24003986.
- LiveScience. Retrieved 19 February 2011.
- ^ PMID 21330491.
- PMID 18316722.
- ^ "Evolution in the urban jungle". Archived from the original on 8 November 2012. Retrieved 8 July 2010.
- ^ "Human Activity Boosts Brain Size in Animals". Yale Scientific. 21 December 2013. Retrieved 16 March 2015.
- PMID 23966638.
- ^ S2CID 10765531
- S2CID 2322067
- ^
- ^
- S2CID 85083567
- S2CID 36627140
- ISBN 978-0-87893-089-0
- S2CID 85410953
- ISBN 978-0-87893-089-0
- ^ Liebherr, James K.; McHugh, Joseph V. in Resh, V. H.; Cardé, R. T. (Editors) 2003. Encyclopedia of Insects. Academic Press.
- ^ PMID 11403874
- ^ Erwin, D. H. & Anstey, R. L. (1995), New Approaches to Speciation in the Fossil Record, Columbia University Press, p. 22
- S2CID 88910977
- S2CID 88958553
- ISBN 978-0-8137-1126-3
- S2CID 5625312.
- S2CID 88897579
- S2CID 9145271.
- ^ "The Evolution of Reproductive Isolation as a Correlated Character Under Sympatric Conditions: Experimental Evidence" (PDF). William R. Rice, George W. Salt. Archived from the original (PDF) on 13 May 2012. Retrieved 23 May 2010.
- ^ "Observed Instances of Speciation, 5.3.5 Sympatric Speciation in Drosophila melanogaster". Joseph Boxhorn. Retrieved 23 May 2010.
- PMID 22337505
- PMID 23349952
- PMID 12663534.
- JSTOR 2413033.
- S2CID 9677456.
- PMID 28562891.
- S2CID 19076040.
- PMID 11038585.
- ^ a b c "London underground source of new insect forms". The Times. 26 August 1998.
- ^ S2CID 40158868. Archived from the original(PDF) on 23 July 2011.
- ^ a b Burdick, Alan (2001). "Insect From the Underground — London, England Underground home to different species of mosquitos". Natural History.
- PMID 10200079.
- ^ Vinogradova, E.B.; Shaikevich, E.V. (2007). "Morphometric, physiological and molecular characteristics of underground populations of the urban mosquito Culex pipiens Linnaeus f. molestus Forskål (Diptera: Culicidae) from several areas of Russia" (PDF). European Mosquito Bulletin. 22: 17–24.
- ^ S2CID 11820649
- PMID 25859042
- S2CID 31875676
- ^ PMC 1689526
- ^
- ^ PMID 26625345
- PMID 24452025
- S2CID 29320650
- S2CID 14856314.
- PMID 18006297.
- ^ S2CID 3329282.
- PMID 10860970.
- S2CID 2457445.
- S2CID 31637733.
- PMID 11092833.
- ^ S2CID 30147815
- PMID 23170069.
- PMID 25929999.
- ^ Karpechenko, G.D. (1927). "Polyploid hybrids of Raphanus sativus X Brassica oleracea L". Bull. Appl. Bot. 17: 305–408.
- .
- JSTOR 2446125.
- ISBN 978-0-670-02053-9.
- ^ Missouri Botanical Garden. "TROPICOS Web display Senecio vulgaris L". Nomenclatural and Specimen Data Base. Missouri State Library. Archived from the original on 2 August 2013. Retrieved 1 February 2008.
- ^ PMID 17803357.
- ^ "New plant species arise from conflicts between immune system genes". Ed Yong. Archived from the original on 3 February 2010. Retrieved 22 May 2010.
- ISBN 978-0-7167-9856-9.
- ^ Soltis, Pam (17 March 2011). "UF researcher: Flowering plant study 'catches evolution in the act'". EurekAlert, American Association for the Advancement of Science. Retrieved 28 March 2011.
- S2CID 18417406.
- ^ Yong, Ed (3 December 2009). "British birdfeeders split blackcaps into two genetically distinct groups". ScienceBlogs. Retrieved 21 May 2010.
- PMID 19443506.
- ^ "Giant insect splits cavefish into distinct populations". Ed Yong. Archived from the original on 1 February 2010. Retrieved 22 May 2010.
- ^ J. P. Doupé; J. H. England; M. Furze; D. Paetkau (2007), "Most Northerly Observation of a Grizzly Bear (Ursus arctos) in Canada: Photographic and DNA Evidence from Melville Island, Northwest Territories", Arctic, 60: 271–276
- ^ Gray, A. P. (1972), Mammalian hybrids. A check-list with bibliography (2nd ed.), Farnham Royal, Slough SL2 3BN, England: Commonwealth Agricultural Bureaux
- PMID 24903145.
- ^ PMID 20194737.
- S2CID 12671275.
- PMID 24813606
- PMID 22826254.
- ^ "Evolution". Polar Bears International. Archived from the original on 30 November 2016. Retrieved 30 January 2014.
- ^ Hansman, Jared (6 August 2008). "Adaptive Traits of the Polar Bear (Ursus maritimus)". Helium, Inc. Archived from the original on 25 January 2018. Retrieved 30 January 2014.
- ^ S2CID 153315450.
- ISBN 978-0-679-64288-6.
- ISBN 978-0-8018-4391-4.
- ^ a b Cott, Hugh B. (1940). Adaptive Coloration in Animals. Methuen. pp. 68–72.
- ^ PMID 22319093.
- ISSN 0014-3820.
- ISSN 0014-3820.
- ISSN 0014-3820.
- ISBN 978-1-4733-7510-9.
- ^ Poulton, Edward Bagnall (1890). The Colours of Animals. p. Foldout after page 339, and throughout.
- PMID 27883046.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - ^ Simulated Evolution Gets Complex. Trnmag.com (8 May 2003). Retrieved on 2011-12-06.
- PMID 10781045.
- PMID 15289608.
- ^ PMID 7938023.
- ^ Sauter, E. (27 March 2006). ""Accelerated Evolution" Converts RNA Enzyme to DNA Enzyme In Vitro". TSRI – News & Views. 6 (11).
- ^ Molecular evolution. kaist.ac.kr
- ^ In Vitro Molecular Evolution. Isgec.org (4 August 1975). Retrieved on 2011-12-06.
- ^ Oswald, Tom (18 July 2001). "Digital organisms used to confirm evolutionary process". EurekAlert! Science News. American Association for the Advancement of Science.
- ^ Dybas, Cheryl (7 May 2003). "Artificial life experiments show how complex functions can evolve". EurekAlert! Science News. American Association for the Advancement of Science.
- ^ Clune, J; Goldsby, H. J.; Ofria, C.; Pennock, R. T. (2011). "Selective pressures for accurate altruism targeting: Evidence from digital evolution for difficult-to-test aspects of inclusive fitness theory" (PDF). Proceedings of the Royal Society. 278. Archived from the original (PDF) on 7 December 2011.
Sources
- Biological science. Oxford. 2002.
- Clegg, C.J. (1998). Genetics & evolution. London: J. Murray. ISBN 978-0-7195-7552-5.
- ISBN 978-0-19-923084-6.
- ISBN 0-517-12320-7
- ISBN 978-1-4165-9478-9.
- Endler, John A. (1986). Natural selection in the wild. New Jersey: Princeton University Press. ISBN 978-0-691-08387-2.
- Fitzhugh, Kirk (2006). "-1-'Evidence' For Evolution Versus 'Evidence' For Intelligent Design: Parallel Confusions". Natural History Museum of Los Angeles County.
- Grant, Peter R.; Grant, B. Rosemary (2014). 40 Years of Evolution: Darwin's Finches on Daphne Major Island. Princeton University Press. p. 432. ISBN 978-0691160467.
- Hill, A.; Behrensmeyer, A. K. (1980). Fossils in the making: vertebrate taphonomy and paleoecology. Chicago: University of Chicago Press. ISBN 978-0-226-04169-8.
- Kluge, Arnold G. (1999). "The Science of Phylogenetic Systematics: Explanation, Prediction and Test". Cladistics. 15 (4): 429–436. S2CID 6034651.
- Laurin, Michel (2000). "Early tetrapod evolution" (PDF). Tree. 15 (3): 118–123. PMID 10675932. Archived from the original(PDF) on 22 July 2012. Retrieved 10 November 2017.
- ISBN 978-0-465-04426-9.
- Paul, C.R.C.; Donovan, S.K. (1998). The adequacy of the fossil record. New York: John Wiley. ISBN 978-0-471-96988-4.
- Prothero, Donald (2007). Evolution: What the Fossils Say and Why it Matters. Columbia University Press. ISBN 978-0231139625.
- Shubin, Neil (2008). Your Inner Fish:A Journey Into the 3.5 Billion-Year History of the Human Body. Random House. ISBN 978-0-375-42447-2.
- Sober, Elliott (2008). Evidence and Evolution: The logic behind the science. Cambridge University Press. ISBN 978-0-521-87188-4.
External links
- National Academies Evolution Resources
- TalkOrigins Archive – 29+ Evidences for Macroevolution: The Scientific Case for Common Descent
- TalkOrigins Archive – Transitional Vertebrate Fossils FAQ
- Understanding Evolution: Your one-stop source for information on evolution
- National Academy Press: Teaching About Evolution and the Nature of Science Archived 2 December 2008 at the Wayback Machine
- Evolution — Provided by PBS.
- Evolution News from Genome News Network (GNN)
- Evolution by Natural Selection — An introduction to the logic of the theory of evolution by natural selection
- Howstuffworks.com — How Evolution Works
- 15 Evolutionary Gems