Exon
An exon is any part of a gene that will form a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term exon refers to both the DNA sequence within a gene and to the corresponding sequence in RNA transcripts. In RNA splicing, introns are removed and exons are covalently joined to one another as part of generating the mature RNA. Just as the entire set of genes for a species constitutes the genome, the entire set of exons constitutes the exome.
History
The term exon derives from the expressed region and was coined by American biochemist Walter Gilbert in 1978: "The notion of the cistron... must be replaced by that of a transcription unit containing regions which will be lost from the mature messenger – which I suggest we call introns (for intragenic regions) – alternating with regions which will be expressed – exons."[1]
This definition was originally made for protein-coding transcripts that are spliced before being translated. The term later came to include sequences removed from
Contribution to genomes and size distribution
Although unicellular
Across all eukaryotic genes in GenBank, there were (in 2002), on average, 5.48 exons per protein coding gene. The average exon encoded 30-36
Structure and function
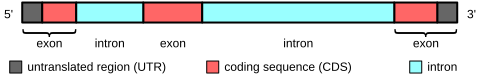
In protein-coding genes, the exons include both the protein-coding sequence and the 5′- and 3′-untranslated regions (UTR). Often the first exon includes both the 5′-UTR and the first part of the coding sequence, but exons containing only regions of 5′-UTR or (more rarely) 3′-UTR occur in some genes, i.e. the UTRs may contain introns.[11] Some non-coding RNA transcripts also have exons and introns.
Mature mRNAs originating from the same gene need not include the same exons, since different introns in the pre-mRNA can be removed by the process of alternative splicing.
Exonization is the creation of a new exon, as a result of mutations in
Experimental approaches using exons
Exon trapping or 'gene trapping' is a molecular biology technique that exploits the existence of the intron-exon splicing to find new genes.[13] The first exon of a 'trapped' gene splices into the exon that is contained in the insertional DNA. This new exon contains the ORF for a reporter gene that can now be expressed using the enhancers that control the target gene. A scientist knows that a new gene has been trapped when the reporter gene is expressed.
Splicing can be experimentally modified so that targeted exons are excluded from mature mRNA transcripts by blocking the access of splice-directing small nuclear ribonucleoprotein particles (snRNPs) to pre-mRNA using Morpholino antisense oligos.[14] This has become a standard technique in developmental biology. Morpholino oligos can also be targeted to prevent molecules that regulate splicing (e.g. splice enhancers, splice suppressors) from binding to pre-mRNA, altering patterns of splicing.
Common misuse of the term
Common incorrect uses of the term exon are that 'exons code for protein', or 'exons code for amino-acids' or 'exons are translated'. However, these sorts of definitions only cover
See also
- DBASS3/5
- Exitron
- Exon-intron database
- Exon shuffling
- Interrupted gene
- Outron
- Twintron
- Untranslated region (UTR)
References
- PMID 622185.
- PMID 3645543.
- PMID 343104.
- S2CID 233382870.
- PMID 6306255.
- PMID 11181995.
- PMID 11752290.
- PMID 15217358.
- PMID 26657562.
- PMID 22955988.
- S2CID 5808466.
- PMID 17709368.
- PMID 2247475.
- PMID 17493584.
- S2CID 233382870.
- PMID 27903072.
- PMID 16712733.
- ^ "Exon". Genome.gov. Archived from the original on 2023-03-16. Retrieved 2023-03-23.
- ^ "Exon". www.nature.com. Scitable. Archived from the original on 2023-03-23. Retrieved 2023-03-23.
Bibliography
- Zhang MQ (May 1998). "Statistical features of human exons and their flanking regions". Human Molecular Genetics. 7 (5): 919–32. PMID 9536098.
- Thanaraj TA, Robinson AJ (November 2000). "Prediction of exact boundaries of exons". Brief. Bioinform. 1 (4): 343–56. PMID 11465052.
