Fibroblast growth factor receptor 1
Fibroblast growth factor receptor 1 (FGFR1), also known as basic fibroblast growth factor receptor 1, fms-related tyrosine kinase-2 / Pfeiffer syndrome, and
Gene
The FGFR1 gene is located on human chromosome 8 at position p11.23 (i.e. 8p11.23), has 24 exons, and codes for a
Protein
Receptor
FGFR1 is a member of the
Cell activation
FGFR1, when bound to a proper FGF, elicits cellular responses by activating signaling pathways that include the: a)

Activation of the gamma isoforms of
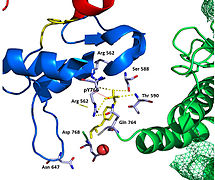
Recent publication on the 2.5 Å crystal structure of PLCγ in complex with FGFR1 kinase (PDB: 3GQI) provides new insights in understanding the molecular mechanism of FGFR1's recruitment of PLCγ by its SH2 domains. Figure 1 on the extreme right shows the PLCγ-FGFR1 kinase complex with the c-SH2 domain colored in red, n-SH2 domain colored in blue, and the interdomain linker colored in yellow. The structure contains typical SH2 domain, with two α-helices and three antiparallel β-strands in each SH2 domain. In this complex, the phosphorylated tyrosine (pY766) on the C-terminal tail of FGFR1 kinase binds preferentially to the nSH2 domain of PLCγ. The phosphorylation of tyrosine residue 766 on FGFR1 kinase forms hydrogen bonds with the n-SH2 to stabilize the complex. Hydrogen bonds in the binding pocket help to stabilize the PLCγ-FGFR1 kinase complex. The water molecule as shown mediates the interaction of asparagine 647 (N647) and aspartate 768 (D768) to further increase the binding affinity of the n-SH2 and FGFR1 kinase complex. (Figure 2). The phosphorylation of tyrosine 653 and tyrosine 654 in the active kinase conformation causes a large conformation change in the activation segment of FGFR1 kinase. Threonine 658 is moved by 24Å from the inactive form (Figure 3.) to the activated form of FGFR1 kinase (Figure 4.). The movement causes the closed conformation in the inactive form to open to enable substrate binding. It also allows the open conformation to coordinate Mg2+ with AMP-PCP (analog of ATP). In addition, pY653 and pY654 in the active form helps to maintain the open conformation of the SH2 and FGFR1 kinase complex. However, the mechanism by which the phosphorylation at Y653 and Y654 helps to recruit the SH2 domain to its C-terminal tail upon phosphorylation of Y766 remains elusive. Figure 5 shows the overlay structure of active and inactive forms of FGFR1 kinase. Figure 6 shows the dots and contacts on phosphorylated tyrosine residues 653 and 654. Green dots show highly favorable contacts between pY653 and pY654 with surrounding residues. Red spikes show unfavorable contacts in the activation segment. The figure is generated through Molprobity extension on Pymol.

The tyrosine kinase region of FGFR1 binds to the N-SH2 domain of PLCγ primarily through charged amino acids. Arginine residue (R609) on the N-SH2 domain forms a salt bridge to aspartate 755 (D755) on the FGFR1 domain. The acid base pairs located in the middle of the interface are nearly parallel to each other, indicating a highly favorable interaction. The N-SH2 domain makes an additional polar contact through water-mediated interaction that takes place between the N-SH2 domain and the FGFR1 kinase region. The arginine residue 609 (R609) on the FGFR1 kinase also forms a salt bridge to the aspartate residue (D594) on the N-SH2 domain. The acid-base pair interacts with each other carry out a reduction–oxidation reaction that stabilizes the complex (Figure 7). Previous studies have done to elucidate the binding affinity of the n-SH2 domain with the FGFR1 kinase complex by mutating these phenylalanine or valine amino acids. The results from isothermal titration calorimetry indicated that the binding affinity of the complex decreased by 3 to 6-fold, without affecting the phosphorylation of the tyrosine residues.[12]
Cell inhibition
FGF-induced activation of FGFR1 also stimulates the activation of
Function
Mice genetically engineered to lack a functional Fgfr1 gene (
Clinical significance
Congenital diseases
Hereditary mutations in the FGFR1 gene are associated with various congenital malformations of the
Cancers
Somatic mutations and epigenetic changes in the expression of the FGFR1 gene occur in and are thought to contribute to various types of lung, breast, hematological, and other types of cancers.
Lung cancers
Breast cancers
Amplification of FGFR1 also occurs in ~10% of estrogen receptor positive breast cancers, particularly of the luminal subtype B form of breast cancer. The presence of FGFR1 amplification has been correlated with resistance to hormone blocking therapy and found to be a poor prognostic factor in the disease.[9][10]
Hematological cancers
In certain rare hematological cancers, the
| Gene | locus | notation | gene | locus | notation | Gene | locus | notation | gene | locus | notation | gene | locus | notation | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MYO18A | 17q11 | t(8;17)(p11;q11) | CPSF6 | 12q15 | t(8;12)(p11;q15) | TPR | 1q25 | t(1;8)(q25p11;; | HERV-K |
10q13 | t(8;13)(p11-q13) | FGFR1OP2 | 12p11 | t(8;12)(p11;q12) | ||||
| ZMYM2 | 13q12 | t(8;13)(p11;q12) | CUTL1 | 7q22 | t(7;8)(q22;p11) | SQSTM1 |
5q35 | t(5;8)(q35;p11 | RANBP2 | 2q13 | t(2;8)(q13;p11) | LRRFIP1 | 2q37 | t(8;2)(p11;q37) | ||||
| CNTRL | 9q33 | t(8;9)(p11;q33) | FGFR1OP | 6q27 | t(6;8)(q27;p11) | BCR | 22q11 | t(8;22)(p11;q11 | NUP98 | 11p15 | t(8;11)(p11-p15) | MYST3 |
8p11.21 | multiple[21] | ||||
CEP110 |
16p12 | t(8;16)(p11;p12) |
These cancers are sometimes termed 8p11
Unlike many other myeloid neoplasms with eosinophil such as those caused by
Phosphaturic mesenchymal tumor
Rhabdomyosarcoma
Elevated expression of FGFR1 protein was detected in 10 of 10 human
Other types of cancers
Acquired abnormalities if the FGFR1 gene are found in: ~14% of urinary bladder
FGFR inhibitors
FGFR-targeted drugs exert direct as well as indirect anticancer effects, due to the fact that FGFRs on cancer cells and endothelial cells are involved in tumorigenesis and vasculogenesis, respectively.
| PD173074 | Dovitinib |
Ki23057 | Lenvatinib | Brivanib |
Nintedanib | Ponatinib | MK-2461 | Lucitanib | AZD4547 |
|---|---|---|---|---|---|---|---|---|---|
| 26 | 8 | NA | 46 | 148 | 69 | 2.2 | 65 | 18 | 0.2 |
FGFR1 mutation in breast and lung cancer as a result of
FGFR inhibitors have been predicted to be effective on relapsed tumors because of the clonal evolution of an FGFR-activated minor subpopulation after therapy targeted to EGFRs or VEGFRs. Because there are multiple mechanisms of action for FGFR inhibitors to overcome drug resistance in human cancer, FGFR-targeted therapy might be a promising strategy for the treatment of refractory cancer.[31]
AZD4547 has undergone a phase II clinical trial in gastric cancer and reported some results.[32]
Interactions
Fibroblast growth factor receptor 1 has been shown to
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000077782 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031565 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 2162671.
- ^ S2CID 42668440.
- ^ "FGFR1 fibroblast growth factor receptor 1 [Homo sapiens (human)] - Gene - NCBI".
- PMID 26277103.
- ^ S2CID 27412585.
- ^ PMID 23880303.
- ^ PMID 26224133.
- PMID 19665973.
- PMID 9169049.
- PMID 21479780.
- ^ "OMIM Entry - * 136350 - FIBROBLAST GROWTH FACTOR RECEPTOR 1; FGFR1".
- ^ "FGFR1 Alterations". Archived from the original on 2015-04-27. Retrieved 2015-04-21.[full citation needed]
- PMID 23182986.
- ^ S2CID 10435391.
- ^ PMID 28028030.
- PMID 28010895.
- ^ S2CID 5256456.
- S2CID 9887919.
- PMID 27443518.
- S2CID 8865648.
- PMID 21666686.
- PMID 22837387.
- PMID 26661925.
- ^ PMID 23658459.
- S2CID 238231910.
- PMID 34830836.
- PMID 24587832.
- ^ "A randomized, open-label phase II study of AZD4547 (AZD) versus Paclitaxel (P) in previously treated patients with advanced gastric cancer (AGC) with Fibroblast Growth Factor Receptor 2 (FGFR2) polysomy or gene amplification (amp): SHINE study". Archived from the original on 2016-03-24. Retrieved 2016-07-07.
- PMID 25193991.
- PMID 11030354.
- PMID 8576175.
- PMID 11877385.
- PMID 10629055.
- PMID 9660748.
- PMID 11090629.
- S2CID 4387190.
- PMID 10713090.
- PMID 7537362.
Further reading
- Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, et al. (December 2010). "Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer". Science Translational Medicine. 2 (62): 62ra93. PMID 21160078.
- Johnson DE, Williams LT (1992). Structural and Functional Diversity in the FGF Receptor Multigene Family. Advances in Cancer Research. Vol. 60. pp. 1–41. PMID 8417497.
- Macdonald D, Reiter A, Cross NC (2002). "The 8p11 myeloproliferative syndrome: a distinct clinical entity caused by constitutive activation of FGFR1". Acta Haematologica. 107 (2): 101–107. S2CID 9582122.
- Groth C, Lardelli M (2002). "The structure and function of vertebrate fibroblast growth factor receptor 1". The International Journal of Developmental Biology. 46 (4): 393–400. PMID 12141425.
- Wilkie AO (April 2005). "Bad bones, absent smell, selfish testes: the pleiotropic consequences of human FGF receptor mutations". Cytokine & Growth Factor Reviews. 16 (2): 187–203. PMID 15863034.
External links
- GeneReviews/NIH/NCBI/UW entry on FGFR-Related Craniosynostosis Syndromes
- GeneReviews/NCBI/NIH/UW entry on Kallmann syndrome
- FGFR1+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Fibroblast growth factor receptor 1 on the Atlas of Genetics and Oncology
- FGFR1 human gene location in the UCSC Genome Browser.
- FGFR1 human gene details in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: P11362 (Human Fibroblast growth factor receptor 1) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: P16092 (Mouse Fibroblast growth factor receptor 1) at the PDBe-KB.
This article incorporates text from the United States National Library of Medicine, which is in the public domain.