File:Subhash nucleoid 10.png
Page contents not supported in other languages.

Size of this preview: 523 × 600 pixels. Other resolutions: 209 × 240 pixels | 418 × 480 pixels | 669 × 768 pixels | 893 × 1,024 pixels | 1,785 × 2,048 pixels | 2,693 × 3,089 pixels.
Original file (2,693 × 3,089 pixels, file size: 424 KB, MIME type: image/png)
| This is a file from the Wikimedia Commons. Information from its description page there is shown below. Commons is a freely licensed media file repository. You can help. |
Summary
| DescriptionSubhash nucleoid 10.png |
English: Models for DNA organization by MatP and MukBEF A. A matS-bridging model for DNA organization in the Ter macrodomain by MatP. MatP recognizes a 13-bp signature DNA sequence called matS that is present exclusively in the Ter macrodomain. There are 23 matS sites separated by one another by an average of 35-kb. MatP binds to a matS site as a dimer, and the tetramerization of the DNA-bound dimers bridges matS sites forming large DNA loops. B. The architecture of the E. coli MukBEF complex. The complex is formed by protein-protein interactions between MukB (blue), MukF (dark orange) and MukE (light orange). MukB, which belongs to the family of structural maintenance of chromosomes (SMCs) proteins, forms a dimer (monomers are shown by dark and light blue colors) consisting of an ATPase head domain and a 100 nm long intramolecular coiled-coil with a hinge region in the middle. Because of the flexibility of the hinge region, MukB adopts a characteristic V-shape of the SMC family. MukF also tends to exist as a dimer because of the strong dimerization affinity between monomers.[135][136] The C-terminal domain of MukF can interact with the head domain of MukB while its central domain can interact with MukE. Two molecules of MukE and one molecule of MukF associate with each other independent of MukB to form a trimeric complex (MukE2F). Since MukF tends to exist in a dimeric form, the dimerization of MukF results in an elongated hexameric complex (MukE2F)2.[137] In the absence of ATP, the (MukE2F)2 complex binds to the MukB head domains through the C-terminal domain of MukF to form a symmetric MukBEF complex (shown on the left). The stoichiometry of the symmetric complex is B2(E2F)2. The ATP binding between the MukB head domains forces the detachment of one MukF molecule and two MukE molecules.[138][137] As a result, an asymmetric MukBEF complex of the stoichiometry B2(E2F)1 is formed. Since MukF readily dimerizes, the MukF dimerization can potentially join two ATP-bound asymmetric molecules resulting in the formation of a dimer of dimers with the stoichiometry of B4(E2F)2 (shown on the right). The stoichiometry of the MukBEF complex in vivo is estimated to be B4(E2F)2 suggesting that a dimer of dimers is the functional unit in vivo.[139] C. A model for loop extrusion by a MukBEF dimer of dimers. A dimer of dimer loads onto DNA (depicted as a grey line) through DNA binding domains of MukB. MukB has been shown to bind DNA via its hinge region and the top region of its head domain.[45][140] The translocation of the complex away from its loading site then extrudes DNA loops. The loops are extruded in a rock-climbing manner by the coordinated opening and closing of the MukBEF ring through the MukB head disengagement that occurs due to coordinated ATP hydrolysis in the two dimers.[139] Dark and light blue circles represent ATP binding and hydrolysis events respectively. MukE is not shown in the complex for simplicity. |
| Date | |
| Source | doi:10.1371/journal.pgen.1008456 Verma SC, Qian Z, Adhya SL (2019) Architecture of the Escherichia coli nucleoid. PLoS Genet 15(12): e1008456. |
| Author | Subhash C. Verma, Zhong Qian, Sankar L. Adhya |
Licensing
This file is licensed under the Creative Commons Attribution 4.0 International license.
- You are free:
- to share – to copy, distribute and transmit the work
- to remix – to adapt the work
- Under the following conditions:
- attribution – You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
Captions
Add a one-line explanation of what this file represents
Items portrayed in this file
depicts
12 December 2019
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 00:15, 18 December 2019 | 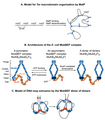 | 2,693 × 3,089 (424 KB) | Evolution and evolvability | User created page with UploadWizard |
File usage
The following pages on the English Wikipedia use this file (pages on other projects are not listed):
Global file usage
The following other wikis use this file:
- Usage on en.wikiversity.org
Metadata
This file contains additional information, probably added from the digital camera or scanner used to create or digitize it.
If the file has been modified from its original state, some details may not fully reflect the modified file.
| Short title |
|
|---|---|
| Width | 3,400 px |
| Height | 4,400 px |
| Pixel composition | RGB |
| Number of components | 3 |
| Horizontal resolution | 400 dpi |
| Vertical resolution | 400 dpi |
| Color space | sRGB |
| Image width | 3,400 px |
| Image height | 4,400 px |
| Bits per component |
|
| Software used | Adobe Illustrator CC 23.0 (Macintosh) |
| Date and time of digitizing | 11:53, 5 April 2019 |
| File change date and time | 11:56, 5 April 2019 |
| Date metadata was last modified | 11:56, 5 April 2019 |
| Unique ID of original document | uuid:9E3E5C9A8C81DB118734DB58FDDE4BA7 |
Retrieved from "https://en.wikipedia.org/wiki/File:Subhash_nucleoid_10.png"
