Fluorescence in situ hybridization

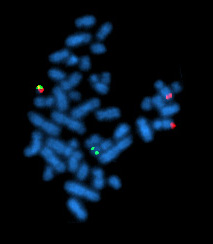
Fluorescence in situ hybridization (FISH) is a
Probes – RNA and DNA

In biology, a probe is a single strand of DNA or RNA that is complementary to a nucleotide sequence of interest.
RNA probes can be designed for any gene or any sequence within a gene for visualization of

Probes are often derived from fragments of DNA that were isolated, purified, and amplified for use in the
Preparation and hybridization process – RNA
The purpose of using RNA FISH is to detect target mRNA transcripts in cells, tissue sections, or even whole-mounts.[10] The process is done in 3 main procedures: tissue preparation (pre-hybridization), hybridization, and washing (post-hybridization).
The tissue preparation starts by collecting the appropriate tissue sections to perform RNA FISH. First, cells, circulating tumor cells (CTCs), formalin-fixed paraffin-embedded (FFPE), or frozen tissue sections are fixed. Some commonly used fixatives are 4% formaldehyde or paraformaldehyde (PFA) in phosphate buffered saline (PBS).[10] FISH has also been successfully done on unfixed cells.[11] After fixation, samples are permeabilized to allow the penetration of hybridization reagents. The use of detergents at a 0.1% concentration is commonly used to enhance the tissue permeability such as Tween-20 or Triton X-100.[12]
It is critical for the hybridization process to have all optimal conditions to have a successful in situ result, including temperature, pH, salt concentration, and time of the hybridization reaction. After checking all the necessary conditions, hybridization steps can be started by first adding a target-specific probe, composed of 20 oligonucleotide pairs, hybridizes to the target RNA(s). Separate but compatible signal amplification systems enable the multiplex assay (up to two targets per assay). Signal amplification is achieved via series of sequential hybridization steps.[13]
After the hybridization steps, washing steps are performed. These steps aim to remove nonspecific hybrids and get rid of unbound probe molecules from the samples to reduce any background signaling. The use of ethanol washes are typically used at this stage to reduce autofluorescence in tissues or cells.[14] At the end of the assay the tissue samples are visualized under a fluorescence microscope such as the confocal fluorescence microscope and the Keyence microscope.[12]
Preparation and hybridization process – DNA

First, a probe is constructed. The probe must be large enough to hybridize specifically with its target but not so large as to impede the hybridization process. The probe is
Then, an
If the fluorescent signal is weak, amplification of the signal may be necessary in order to exceed the detection threshold of the
Variations on probes and analysis
FISH is a very general technique. The differences between the various FISH techniques are usually due to variations in the sequence and labeling of the probes; and how they are used in combination. Probes are divided into two generic categories: cellular and acellular. In fluorescent "in situ" hybridization refers to the cellular placement of the probe
Probe size is important because shorter probes hybridize less specifically than longer probes, so that long enough strands of DNA or RNA (often 10–25 nucleotides) which are complementary to a given target sequence are often used to locate a target. The overlap defines the resolution of detectable features. For example, if the goal of an experiment is to detect the breakpoint of a translocation, then the overlap of the probes — the degree to which one DNA sequence is contained in the adjacent probes — defines the minimum window in which the breakpoint may be detected.
The mixture of probe sequences determines the type of feature the probe can detect. Probes that hybridize along an entire chromosome are used to count the number of a certain chromosome, show translocations, or identify extra-chromosomal fragments of
A variety of other techniques uses mixtures of differently colored probes. A range of colors in mixtures of fluorescent dyes can be detected, so each human chromosome can be identified by a characteristic color using whole-chromosome probe mixtures and a variety of ratios of colors. Although there are more chromosomes than easily distinguishable fluorescent dye colors, ratios of probe mixtures can be used to create secondary colors. Similar to comparative genomic hybridization, the probe mixture for the secondary colors is created by mixing the correct ratio of two sets of differently colored probes for the same chromosome. This technique is sometimes called M-FISH.
The same physics that make a variety of colors possible for M-FISH can be used for the detection of translocations. That is, colors that are adjacent appear to overlap; a secondary color is observed. Some assays are designed so that the secondary color will be present or absent in cases of interest. An example is the detection of BCR/ABL translocations, where the secondary color indicates disease. This variation is often called double-fusion FISH or D-FISH. In the opposite situation—where the absence of the secondary color is pathological—is illustrated by an assay used to investigate translocations where only one of the breakpoints is known or constant. Locus-specific probes are made for one side of the breakpoint and the other intact chromosome. In normal cells, the secondary color is observed, but only the primary colors are observed when the translocation occurs. This technique is sometimes called "break-apart FISH".
Single-molecule RNA FISH
Single-molecule RNA FISH, also known as Stellaris® RNA FISH
Single-molecule RNA FISH assays can be performed in simplex or
.Fiber FISH
In an alternative technique to
Q-FISH
Q-FISH combines FISH with PNAs and computer software to quantify fluorescence intensity. This technique is used routinely in telomere length research.
Flow-FISH
Flow-FISH uses flow cytometry to perform FISH automatically using per-cell fluorescence measurements.
MA-FISH
Microfluidics-assisted FISH (MA-FISH) uses a microfluidic flow to increase DNA hybridization efficiency, decreasing expensive FISH probe consumption and reduce the hybridization time. MA-FISH is applied for detecting the
MAR-FISH
Microautoradiography FISH is a technique to combine radio-labeled substrates with conventional FISH to detect phylogenetic groups and metabolic activities simultaneously.[23]
Hybrid Fusion-FISH
Hybrid Fusion FISH (HF-FISH) uses primary additive excitation/emission combination of fluorophores to generate additional spectra through a labeling process known as dynamic optical transmission (DOT). Three primary fluorophores are able to generate a total of 7 readily detectable emission spectra as a result of combinatorial labeling using DOT. Hybrid Fusion FISH enables highly multiplexed FISH applications that are targeted within clinical oncology panels. The technology offers faster scoring with efficient probesets that can be readily detected with traditional fluorescent microscopes.
MERFISH
Multiplexed error-robust fluorescence in situ hybridization[24] is a highly multiplexed version of smFISH. It uses combinatorial labeling, followed by imaging, and then error-resistant encoding[25] to capture a high number of RNA molecules and spatial localization within the cell. The capture of a large number of RNA molecules enables elucidation of gene regulatory networks, prediction of function of unannotated genes, and identification of RNA molecule distribution patterns, which correlate with their associated proteins.
STARFISH
Starfish is a set of software tools developed in 2019 by a consortium of scientists to analyze data from nine different variations of FISH, since all variations produce the same set of data—gene expression values mapped to x and y coordinates in a cell. The software, created for all scientists, not just bioinformaticians, reads a set of images, removes noise, and identifies RNA molecules. This approach has set out to define a standard analysis scheme of FISH datasets in a similar way to single-cell transcriptomics analysis.[26]
Medical applications
Often parents of children with a
In medicine, FISH can be used to form a

Species identification
FISH has been extensively studied as a diagnostic technique for the identification of pathogens in the field of medical microbiology.[30] Although it has been proven to be a useful and applicable technique, it is still not widely applied in diagnostic laboratories. The short time to diagnosis (less than 2 hours) has been a major advantage compared with biochemical differentiation, but this advantage is challenged by MALDI-TOF-MS which allows the identification of a wider range of pathogens compared with biochemical differentiation techniques. Using FISH for diagnostic purposes has found its purpose when immediate species identification is needed, specifically for the investigation of blood cultures for which FISH is a cheap and easy technique for preliminary rapid diagnosis.[30]
FISH can also be used to compare the genomes of two biological
FISH is widely used in the field of
Comparative genomic hybridization
Comparative genomic hybridization can be described as a method that uses FISH in a parallel manner with the comparison of the hybridization strength to recall any major disruptions in the duplication process of the DNA sequences in the genome of the nucleus.[31]
Virtual karyotype
Spectral karyotype
Chromosome evolution

FISH can be used to study the evolution of chromosomes. Species that are related have similar chromosomes. This homology can be detected by gene or genome sequencing but also by FISH. For instance, human and chimpanzee chromosomes are very similar and FISH can demonstrate that two chimpanzee chromosomes fused to result in one human chromosome. Similarly, species that are more distantly related, have similar chromosomes but with increasing distance chromosomes tend to break and fuse and thus result in mosaic chromosomes. This can be impressively demonstrated by FISH (see figure).[32]
See also
- Chromogenic in situ hybridization (CISH)
- Eukaryotic chromosome fine structure
- G banding
- Gene mapping
- Genome evolution
- Happy mapping
- In situ hybridization, the technique used for labelling
- Molecular cytogenetics
- Virtual karyotype
Gallery
-
Another schematic of FISH process.
-
Microfluidic chip that lowered the cost-per-test of FISH by 90%.
-
Dual label FISH image; Bifidobacteria Cy3, Total bacteria FITC.
-
Paraspeckles visualized by single-molecule FISH against NEAT1 (Quasar 570) in U-2 OS cells (DAPI).
References
- PMID 6812046.
- S2CID 22498325.
- PMID 22851656.
- PMID 22446628.
- PMID 22253826.
- PMID 21233348.
- PMID 22840402.
- PMID 21518807.
- ^ PMID 18234275.
- ^ PMID 32219032.
- PMID 24060113.
- ^ PMID 27656642.
- PMID 29785002.
- PMID 20552552.
- PMID 26721487.
- PMID 29889208.
- PMID 18806792.
- ^ Biosearch Technologies Signs Exclusive License for Single Molecule FISH Technologies from UMDNJ. biosearchtech.com
- PMID 10610624.
- S2CID 26313219.
- PMID 8740834.
- PMID 27892928.
- .
- PMID 25858977.
- PMID 25858977.
- S2CID 201064966.
- ^ PMID 19254793.
- S2CID 4955677.
- ISBN 978-0387727165.
- ^ S2CID 25252460.
- ^ "Comparative Genomic Hybridization". McGraw-Hill Dictionary of Scientific and Technical Terms. Retrieved September 19, 2013.
- ^ PMID 36207754.
Further reading
- Pernthaler A, Pernthaler J, Amann R (June 2002). "Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria". Applied and Environmental Microbiology. 68 (6): 3094–3101. PMID 12039771.
- Wagner M, Horn M, Daims H (June 2003). "Fluorescence in situ hybridisation for the identification and characterisation of prokaryotes". Current Opinion in Microbiology. 6 (3): 302–309. PMID 12831908.
- Carthy JD (1965). Viewpoints In Biology. England: Butterworth & Co. p. 66.
External links
- Fluorescent+in+Situ+Hybridization at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Information on fiber FISH from the Olympus Corporation
- A guide to fiber FISH from Octavian Henegariu
- Fibre FISH protocol Archived 2006-10-23 at the Sanger Centre
- CARD-FISH, BioMineWiki Archived 2020-07-28 at the Wayback Machine
- Preparation of Complex DNA Probe Sets for 3D FISH with up to Six Different Fluorochromes
- FISH technical notes and protocols from GeneDetect.com
- Fluorescence in situ Hybridization Photos of bacteria Archived 2015-02-05 at the Wayback Machine
- Rational design of polynucleotide probe mixes to identify particular genes in defined taxa: www.dnaBaser.com/PolyPro