Fluorescent tag

In molecular biology and biotechnology, a fluorescent tag, also known as a fluorescent label or fluorescent probe, is a molecule that is attached chemically to aid in the detection of a biomolecule such as a protein, antibody, or amino acid. Generally, fluorescent tagging, or labeling, uses a reactive derivative of a fluorescent molecule known as a fluorophore. The fluorophore selectively binds to a specific region or functional group on the target molecule and can be attached chemically or biologically.[1] Various labeling techniques such as enzymatic labeling, protein labeling, and genetic labeling are widely utilized. Ethidium bromide, fluorescein and green fluorescent protein are common tags. The most commonly labelled molecules are antibodies, proteins, amino acids and peptides which are then used as specific probes for detection of a particular target.[2]
History


The development of methods to detect and identify biomolecules has been motivated by the ability to improve the study of molecular structure and interactions. Before the advent of fluorescent labeling,
Ethidium bromide and variants were developed in the 1950s,[4] and in 1994, fluorescent proteins or FPs were introduced.[5] Green fluorescent protein or GFP was discovered by Osamu Shimomura in the 1960s and was developed as a tracer molecule by Douglas Prasher in 1987.[6] FPs led to a breakthrough of live cell imaging with the ability to selectively tag genetic protein regions and observe protein functions and mechanisms.[5] For this breakthrough, Shimomura was awarded the Nobel Prize in 2008.[7]
New methods for tracking biomolecules have been developed including the use of colorimetric biosensors, photochromic compounds,

Methods for tracking biomolecules
There are currently several labeling methods for tracking biomolecules. Some of the methods include the following.
Isotope markers
Common species that isotope markers are used for include proteins. In this case, amino acids with stable isotopes of either carbon, nitrogen, or hydrogen are incorporated into polypeptide sequences.[8] These polypeptides are then put through mass spectrometry. Because of the exact defined change that these isotopes incur on the peptides, it is possible to tell through the spectrometry graph which peptides contained the isotopes. By doing so, one can extract the protein of interest from several others in a group. Isotopic compounds play an important role as photochromes, described below.
Colorimetric biosensors
Biosensors are attached to a substance of interest. Normally, this substance would not be able to absorb light, but with the attached biosensor, light can be absorbed and emitted on a
Colorimetric assays are normally used to determine how much concentration of one species there is relative to another.[9]
Photochromic compounds
The most common organic molecule to be used as a photochrome is diarylethene.[12] Other examples of photoswitchable proteins include PADRON-C, rs-FastLIME-s and bs-DRONPA-s, which can be used in plant and mammalian cells alike to watch cells move into different environments.[11]
Biomaterials
Fluorescent biomaterials are a possible way of using external factors to observe a pathway more visibly. The method involves fluorescently labeling peptide molecules that would alter an organism's natural pathway. When this peptide is inserted into the organism's cell, it can induce a different reaction. This method can be used, for example to treat a patient and then visibly see the treatment's outcome.[13]
Electrochemical sensors
Electrochemical sensors can be used for label-free sensing of biomolecules. They detect changes and measure current between a probed metal electrode and an electrolyte containing the target analyte. A known potential to the electrode is then applied from a feedback current and the resulting current can be measured. For example, one technique using electrochemical sensing includes slowly raising the voltage causing chemical species at the electrode to be oxidized or reduced. Cell current vs voltage is plotted which can ultimately identify the quantity of chemical species consumed or produced at the electrode.[14] Fluorescent tags can be used in conjunction with electrochemical sensors for ease of detection in a biological system.
Fluorescent labels

Of the various methods of labeling biomolecules, fluorescent labels are advantageous in that they are highly sensitive even at low concentration and non-destructive to the target molecule folding and function.[1]
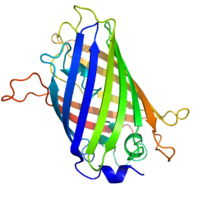
Green fluorescent protein is a naturally occurring fluorescent protein from the jellyfish Aequorea victoria that is widely used to tag proteins of interest. GFP emits a photon in the green region of the light spectrum when excited by the absorption of light. The chromophore consists of an oxidized tripeptide -Ser^65-Tyr^66-Gly^67 located within a β barrel. GFP catalyzes the oxidation and only requires molecular oxygen. GFP has been modified by changing the wavelength of light absorbed to include other colors of fluorescence. YFP or
Synthetic fluorescent probes can also be used as fluorescent labels. Advantages of these labels include a smaller size with more variety in color. They can be used to tag proteins of interest more selectively by various methods including chemical recognition-based labeling, such as utilizing metal-chelating peptide tags, and biological recognition-based labeling utilizing enzymatic reactions.[16] However, despite their wide array of excitation and emission wavelengths as well as better stability, synthetic probes tend to be toxic to the cell and so are not generally used in cell imaging studies.[1]
Fluorescent labels can be hybridized to mRNA to help visualize interaction and activity, such as mRNA localization. An antisense strand labeled with the fluorescent probe is attached to a single mRNA strand, and can then be viewed during cell development to see the movement of mRNA within the cell.[17]
Fluorogenic labels
A fluorogen is a ligand (fluorogenic ligand) which is not itself fluorescent, but when it is bound by a specific protein or RNA structure becomes fluorescent.[18]
For instance, FAST is a variant of photoactive yellow protein which was engineered to bind chemical mimics of the GFP tripeptide chromophore.[19] Likewise, the spinach aptamer is an engineered RNA sequence which can bind GFP chromophore chemical mimics, thereby conferring conditional and reversible fluorescence on RNA molecules containing the sequence.[20]
Use of tags in fluorescent labeling



Fluorescent labeling is known for its non-destructive nature and high sensitivity. This has made it one of the most widely used methods for labeling and tracking biomolecules.[1] Several techniques of fluorescent labeling can be utilized depending on the nature of the target.
Enzymatic labeling
In enzymatic labeling, a DNA construct is first formed, using a gene and the DNA of a fluorescent protein.[21] After transcription, a hybrid RNA + fluorescent is formed. The object of interest is attached to an enzyme that can recognize this hybrid DNA. Usually fluorescein is used as the fluorophore.
Chemical labeling
Chemical labeling or the use of chemical tags utilizes the interaction between a small molecule and a specific genetic amino acid sequence.[22] Chemical labeling is sometimes used as an alternative for GFP. Synthetic proteins that function as fluorescent probes are smaller than GFP's, and therefore can function as probes in a wider variety of situations. Moreover, they offer a wider range of colors and photochemical properties.[23] With recent advancements in chemical labeling, Chemical tags are preferred over fluorescent proteins due to the architectural and size limitations of the fluorescent protein's characteristic β-barrel. Alterations of fluorescent proteins would lead to loss of fluorescent properties.[22]
Protein labeling
Protein labeling use a short tag to minimize disruption of protein folding and function. Transition metals are used to link specific residues in the tags to site-specific targets such as the N-termini, C-termini, or internal sites within the protein. Examples of tags used for protein labeling include biarsenical tags, Histidine tags, and FLAG tags.[1]
Genetic labeling
Cell imaging
Chemical tags have been tailored for imaging technologies more so than fluorescent proteins because chemical tags can localize photosensitizers closer to the target proteins.
Advantages
Although fluorescent dyes may not have the same sensitivity as radioactive probes, they are able to show real-time activity of molecules in action.[27] Moreover, radiation and appropriate handling is no longer a concern.
With the development of fluorescent tagging,
Latest advances in methods involving fluorescent tags have led to the visualization of
See also
- Molecular tagging velocimetry
- Spectrophotometer for Nucleic Acid Measurements
- Protein tags
Notes
- ^ .
- ^ "Fluorescent labeling of biomolecules with organic probes - Presentations - PharmaXChange.info". 29 January 2011.
- ^ Gwynne and Page, Peter and Guy. "Laboratory Technology Trends: Fluorescence + Labeling". Science. Retrieved 10 March 2013.
- ^ PMID 19233914.
- ^ PMID 21879706.
- ^ "Green Fluorescent Protein - GFP History - Osamu Shimomura".
- ^ Shimomura, Osamu. "The Nobel Prize in Chemistry". Retrieved 5 April 2013.
- PMID 10740850.
- ^ a b "Colorimetric Assays". Retrieved 3 April 2013.
- ^ Halevy, Revital; Sofiya Kolusheval; Robert E.W. Hancock; Raz Jelinek (2002). "Colorimetric Biosensor Vesicles for Biotechnological Applications" (PDF). Materials Research Society Symposium Proceedings. 724. Biological and Biomimetic Materials - Properties to Function. Archived (PDF) from the original on October 14, 2013. Retrieved 4 April 2013.
- ^ PMID 23434876.
- PMID 22668009.
- PMID 23710326.
- ^ "bioee.ee.columbia.edu" (PDF). Archived from the original (PDF) on 2012-12-20.
- ISBN 978-0-7167-7108-1.
- PMID 23318293.
- ^ PMID 20444605.
- S2CID 21815631.
We report here the development of protein reporters that generate fluorescence from otherwise dark molecules (fluorogens).
- PMID 26711992.
- PMID 21798953.
- PMID 12238772.
- ^ PMID 21567974.
- PMID 23318293.
- ^ ISBN 978-1-4292-3413-9.
- PMID 24974023.
- ^ PMID 23115610.
- PMID 8948646.
