Frameshift mutation

A frameshift mutation (also called a framing error or a reading frame shift) is a
Frameshift mutations are apparent in severe genetic diseases such as
Background
The information contained in DNA determines protein function in the cells of all organisms. Transcription and translation allow this information to be communicated into making proteins. However, an error in reading this communication can cause protein function to be incorrect and eventually cause disease even as the cell incorporates a variety of corrective measures.Genetic information is conveyed by DNA for protein synthesis within cells. Misinterpretation can lead to faulty function and disease, despite cellular correction mechanisms.
Central dogma
In 1956
Transcription and translation

After DNA replication, the reading of a selected section of genetic information is accomplished by
Frameshifting may also occur during
There are several biological processes that help to prevent frameshift mutations. Reverse mutations occur which change the mutated sequence back to the original wild type sequence. Another possibility for mutation correction is the use of a suppressor mutation. This offsets the effect of the original mutation by creating a secondary mutation, shifting the sequence to allow for the correct amino acids to be read. Guide RNA can also be used to insert or delete Uridine into the mRNA after transcription, this allows for the correct reading frame.[1]
Codon-triplet importance
A

Mechanism
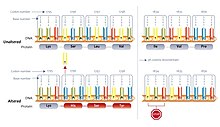
Frameshift mutations can occur randomly or be caused by an external stimulus. The detection of frameshift mutations can occur via several different methods. Frameshifts are just one type of mutation that can lead to incomplete or incorrect proteins, but they account for a significant percentage of errors in DNA.In an unaltered gene, codons (triplets of nucleotides) are sequentially interpreted, with each codon encoding a specific amino acid. This is known as the standard reading frame. However, in cases of frame shift mutations, an extra nucleotide (or more) is inserted into the DNA sequence, disrupting the typical reading frame and causing a shift in the sequence.
This insertion prompts a shift in the reading frame due to the triplet nature of the genetic code. For instance, the addition of an extra "A" leads to a sequence shift, triggering the reading of an entirely different set of codons. This deviation in genetic information causes the ribosome, which reads the mRNA for protein synthesis, to misinterpret the genetic data. Consequently, an entirely different series of amino acids is generated, resulting in the generation of an altered protein sequence. In most instances, the new reading frame results in an early encounter with a stop codon, leading to the formation of a shortened and usually inactive protein. This form of mutation is termed an early stop codon or a nonsense mutation.
Genetic or environmental
This is a genetic mutation at the level of nucleotide bases. Why and how frameshift mutations occur are continually being sought after. An environmental study, specifically the production of
Detection
Fluorescence
The effects of neighboring bases and secondary structure to detect the frequency of frameshift mutations has been investigated in depth using fluorescence. Fluorescently tagged DNA, by means of base analogues, permits one to study the local changes of a DNA sequence.[8] Studies on the effects of the length of the primer strand reveal that an equilibrium mixture of four hybridization conformations was observed when template bases looped-out as a bulge, i.e. a structure flanked on both sides by duplex DNA. In contrast, a double-loop structure with an unusual unstacked DNA conformation at its downstream edge was observed when the extruded bases were positioned at the primer–template junction, showing that misalignments can be modified by neighboring DNA secondary structure.[9]
Sequencing
Sanger sequencing and pyrosequencing are two methods that have been used to detect frameshift mutations, however, it is likely that data generated will not be of the highest quality. Even still, 1.96 million indels have been identified through Sanger sequencing that do not overlap with other databases. When a frameshift mutation is observed it is compared against the Human Genome Mutation Database (HGMD) to determine if the mutation has a damaging effect. This is done by looking at four features. First, the ratio between the affected and conserved DNA, second the location of the mutation relative to the transcript, third the ratio of conserved and affected amino acids and finally the distance of the indel to the end of the exon.[10]
Diagnosis
A US patent (5,958,684) in 1999 by Leeuwen, details the methods and reagents for diagnosis of diseases caused by or associated with a gene having a somatic mutation giving rise to a frameshift mutation. The methods include providing a tissue or fluid sample and conducting gene analysis for frameshift mutation or a protein from this type of mutation. The nucleotide sequence of the suspected gene is provided from published gene sequences or from cloning and sequencing of the suspect gene. The amino acid sequence encoded by the gene is then predicted.[14]NA Sequencing: Sanger sequencing or Next-Generation Sequencing (NGS) can be used to directly sequence the DNA and identify insertions or deletions.Polymerase Chain Reaction (PCR): PCR can be used to amplify the specific region containing the mutation for subsequent analysis.Multiplex Ligation-dependent Probe Amplification (MLPA): MLPA is a technique used to detect copy number variations and small insertions or deletions.Comparative Genomic Hybridization (CGH): CGH is used to detect chromosomal imbalances, which may include large insertions or deletions.
Frequency
Despite the rules that govern the genetic code and the various mechanisms present in a cell to ensure the correct transfer of genetic information during the process of DNA replication as well as during translation, mutations do occur; frameshift mutation is not the only type. There are at least two other types of recognized point mutations, specifically missense mutation and nonsense mutation.[1] A frameshift mutation can drastically change the coding capacity (genetic information) of the message.[1] Small insertions or deletions (those less than 20 base pairs) make up 24% of mutations that manifest in currently recognized genetic disease.[10]
Frameshift mutations are found to be more common in repeat regions of DNA. A reason for this is because of slipping of the polymerase enzyme in repeat regions, allowing for mutations to enter the sequence.[15] Experiments can be run to determine the frequency of the frameshift mutation by adding or removing a pre-set number of nucleotides. Experiments have been run by adding four basepairs, called the +4 experiments, but a team from Emory University looked at the difference in frequency of the mutation by both adding and deleting a base pair. It was shown that there was no difference in the frequency between the addition and deletion of a base pair. There is however, a difference in the result of the protein.[15]
Huntington's disease is one of the nine codon reiteration disorders caused by polyglutamine expansion mutations that include spino-cerebellar ataxia (SCA) 1, 2, 6, 7 and 3, spinobulbar muscular atrophy and dentatorubal-pallidoluysianatrophy. There may be a link between diseases caused by polyglutamine and polyalanine expansion mutations, as frame shifting of the original SCA3 gene product encoding CAG/polyglutamines to GCA/polyalanines. Ribosomal slippage during translation of the SCA3 protein has been proposed as the mechanism resulting in shifting from the polyglutamine to the polyalanine-encoding frame. A dinucleotide deletion or single nucleotide insertion within the polyglutamine tract of huntingtin exon 1 would shift the CAG, polyglutamineen coding frame by +1 (+1 frame shift) to the GCA, polyalanine-encoding frame and introduce a novel epitope to the C terminus of Htt exon 1 (APAAAPAATRPGCG).[16]
Diseases
Several diseases have frameshift mutations as at least part of the cause. Knowing prevalent mutations can also aid in the diagnosis of the disease. Currently there are attempts to use frameshift mutations beneficially in the treatment of diseases, changing the reading frame of the amino acids.


Cancer
Frameshift mutations are known to be a factor in
Crohn's disease
Crohn's disease has an association with the NOD2 gene. The mutation is an insertion of a Cytosine at position 3020. This leads to a premature stop codon, shortening the protein that is supposed to be transcribed. When the protein is able to form normally, it responds to bacterial liposaccharides, where the 3020insC mutation prevents the protein from being responsive.[20]
Cystic fibrosis
HIV
CCR5 is one of the cell entry co-factors associated with HIV, most frequently involved with nonsyncytium-inducing strains, is most apparent in HIV patients as opposed to AIDS patients. A 32 base pair deletion in CCR5 has been identified as a mutation that negates the likelihood of an HIV infection. This region on the open reading frame ORF contains a frameshift mutation leading to a premature stop codon. This leads to the loss of the HIV-coreceptor function in vitro. CCR5-1 is considered the wild type and CCR5-2 is considered to be the mutant allele. Those with a heterozygous mutation for the CCR5 were less susceptible to the development of HIV. In a study, despite high exposure to the HIV virus, there was no one homozygous for the CCR5 mutation that tested positive for HIV.[3]
Tay–Sachs disease
Smith–Magenis syndrome
Smith–Magenis syndrome (SMS) is a complex syndrome involving intellectual disabilities, sleep disturbance, behavioural problems, and a variety of craniofacial, skeletal, and visceral anomalies. The majority of SMS cases harbor an ~3.5 Mb common deletion that encompasses the retinoic acid induced-1 (RAI1) gene. Other cases illustrate variability in the SMS phenotype not previously shown for RAI1 mutation, including hearing loss, absence of self-abusive behaviours, and mild global delays. Sequencing of RAI1 revealed mutation of a heptamericC-tract (CCCCCCC) in exon 3 resulting in frameshift mutations. Of the seven reported frameshift mutations occurring in poly C-tracts in RAI1, four cases (~57%) occur at this heptameric C-tract. The results indicate that this heptameric C-tract is a preferential recombination hotspot insertion/deletions (SNindels) and therefore a primary target for analysis in patients suspected for mutations in RAI1.[25]
Hypertrophic cardiomyopathy
Cures
Finding a cure for the diseases caused by frameshift mutations is rare. Research into this is ongoing. One example is a
A European patent (EP1369126A1) in 2003 by Bork records a method used for prevention of cancers and for the curative treatment of cancers and precancers such as DNA-mismatch repair deficient (MMR) sporadic tumours and HNPCC associated tumours. The idea is to use immunotherapy with combinatorial mixtures of tumour-specific frameshift mutation-derived peptides to elicit a cytotoxic T-cell response specifically directed against tumour cells.[28]
See also
- Translational frameshift
- Mutation
- Transcription (genetics)
- Translation (biology)
- codon
- protein
- reading frame
- point mutation
- Crohn's disease
- Tay–Sachs disease
References
- ^ ISBN 978-0-8053-9592-1.
- ^ "DNA Is Constantly Changing through the Process of Mutation". Nature. Retrieved 17 May 2019.
- ^ PMID 9132277.
- PMID 16162506.
- PMID 15269381.
- ISBN 978-0-7167-7108-1.
- PMID 8028006.
- PMID 14993592.
- PMID 19155277.
- ^ PMID 22322200.
- PMID 19679224.
- PMID 22006311.
- PMID 20616022.
- ^ US Patent 5,958,684 (September 28, 1999) "Diagnosis of Neurodegenerative Disease" by Leeuwen et al
- ^ PMID 10373526.
- PMID 16801344.
- PMID 10.
- PMID 23434521.
- ^ "Cancer Genomics". National Cancer Institute at the National Institute of Health. Archived from the original on 18 March 2013. Retrieved 24 March 2013.
- S2CID 205017657.
- PMID 18639722.
- PMID 1990834.
- ^ "Learning About Tay-Sachs Disease". National Human Genome Research Institute. Retrieved 24 March 2013.
- PMID 9090523.
- PMID 20932317.
- S2CID 46682245.
- PMID 18978469.
- ^ European Patent [1] (December 10, 2003) "Use of coding microsatellite region frameshift mutation-derived peptides for treating cancer" by Bork et al
Further reading
- Farabaugh PJ (1996). "Programmed translational frameshifting". Annu. Rev. Genet. 30 (1): 507–28. PMID 8982463.
- Lewis, Ricki (2005). Human Genetics: Concepts and Applications (6th ed.). Boston MA: McGraw Hill. pp. 227–8. ISBN 978-0-07-111156-0.
- "Nylonase Enzymes". 20 April 2004. Retrieved 2 June 2009.
External links
- Frameshift+Mutation at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- NCBI dbSNP database — "a central repository for both single base nucleotide substitutions and short deletion and insertion polymorphisms"
- Wise2 - aligns a frameshifts and introns
- FastY - compare a DNA sequence to a protein sequence database, allowing gaps and frameshifts
- Path - tool that compares two translationprinciple)
- HGMD - Human Genome Mutation Database
