Gene
| Part of a series on |
| Genetics |
|---|
 |
base pairs encode genes, which provide functions. A human DNA can have up to 500 million base pairs with thousands of genes. |
In biology, the word gene (Greek: γένος, génos;[1] generation,[2] or birth,[1] or gender) has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA, that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and non-coding genes.[3][4][5][6]
During gene expression, DNA is first copied into RNA. RNA can be directly functional or be the intermediate template for the synthesis of a protein.
The transmission of genes to an organism's
A gene can acquire
The term gene was introduced by Danish botanist, plant physiologist and geneticist Wilhelm Johannsen in 1909.[8] It is inspired by the Ancient Greek: γόνος, gonos, that means offspring and procreation.
Definitions
There are many different ways to use the term "gene" based on different aspects of their inheritance, selection, biological function, or molecular structure but most of these definitions fall into two categories, the Mendelian gene or the molecular gene.[3][9][10][11][12]
The Mendelian gene is the classical gene of genetics and it refers to any heritable trait. This is the gene described in The Selfish Gene.[13] More thorough discussions of this version of a gene can be found in the articles on Genetics and Gene-centered view of evolution.
The molecular gene definition is more commonly used across biochemistry, molecular biology, and most of genetics — the gene that is described in terms of DNA sequence.[3] There are many different definitions of this gene — some of which are misleading or incorrect.[9][14]
Very early work in the field that became
This idea of two kinds of genes is still part of the definition of a gene in most textbooks. For example,
- "The primary function of the genome is to produce RNA molecules. Selected portions of the DNA nucleotide sequence are copied into a corresponding RNA nucleotide sequence, which either encodes a protein (if it is an mRNA) or forms a 'structural' RNA, such as a transfer RNA (tRNA) or ribosomal RNA (rRNA) molecule. Each region of the DNA helix that produces a functional RNA molecule constitutes a gene."[19]
- "We define a gene as a DNA sequence that is transcribed. This definition includes genes that do not encode proteins (not all transcripts are messenger RNA). The definition normally excludes regions of the genome that control transcription but are not themselves transcribed. We will encounter some exceptions to our definition of a gene - surprisingly, there is no definition that is entirely satisfactory."[20]
- "A gene is a DNA sequence that codes for a diffusible product. This product may be protein (as is the case in the majority of genes) or may be RNA (as is the case of genes that code for tRNA and rRNA). The crucial feature is that the product diffuses away from its site of synthesis to act elsewhere."[21]
The important parts of such definitions are: (1) that a gene corresponds to a transcription unit; (2) that genes produce both mRNA and noncoding RNAs; and (3) regulatory sequences control gene expression but are not part of the gene itself. However, there's one other important part of the definition and it is emphasized in Kostas Kampourakis' book Making Sense of Genes.
- "Therefore in this book I will consider genes as DNA sequences encoding information for functional products, be it proteins or RNA molecules. With 'encoding information,' I mean that the DNA sequence is used as a template for the production of an RNA molecule or a protein that performs some function.'[9]
The emphasis on function is essential because there are stretches of DNA that produce non-functional transcripts and they do not qualify as genes. These include obvious examples such as transcribed pseudogenes as well as less obvious examples such as junk RNA produced as noise due to transcription errors. In order to qualify as a true gene, by this definition, one has to prove that the transcript has a biological function.[9]
Early speculations on the size of a typical gene were based on high resolution genetic mapping and on the size of proteins and RNA molecules. A length of 1500 base pairs seemed reasonable at the time (1965).[18] This was based on the idea that the gene was the DNA that was directly responsible for production of the functional product. The discovery of introns in the 1970s meant that many eukaryotic genes were much larger than the size of the functional product would imply. Typical mammalian protein-coding genes, for example, are about 62,000 base pairs in length (transcribed region) and since there are about 20,000 of them they occupy about 35–40% of the mammalian genome (including the human genome).[22][23][24]
In spite of the fact that both protein-coding genes and noncoding genes have been known for more than 50 years, there are still a number of textbooks, websites, and scientific publications that define a gene as a DNA sequence that specifies a protein. In other words, the definition is restricted to protein-coding genes. Here is an example from a recent article in American Scientist.
- ... to truly assess the potential significance of de novo genes, we relied on a strict definition of the word "gene" with which nearly every expert can agree. First, in order for a nucleotide sequence to be considered a true gene, an open reading frame (ORF) must be present. The ORF can be thought of as the "gene itself"; it begins with a starting mark common for every gene and ends with one of three possible finish line signals. One of the key enzymes in this process, the RNA polymerase, zips along the strand of DNA like a train on a monorail, transcribing it into its messenger RNA form. This point brings us to our second important criterion: A true gene is one that is both transcribed and translated. That is, a true gene is first used as a template to make transient messenger RNA, which is then translated into a protein.[25]
This restricted definition is so common that it has spawned many recent articles that criticize this "standard definition" and call for a new expanded definition that includes noncoding genes.
Although some definitions can be more broadly applicable than others, the fundamental complexity of biology means that no definition of a gene can capture all aspects perfectly. Not all genomes are DNA (e.g. RNA viruses),[29] bacterial operons are multiple protein-coding regions transcribed into single large mRNAs, alternative splicing enables a single genomic region to encode multiple district products and trans-splicing concatenates mRNAs from shorter coding sequence across the genome.[30][31][32] Since molecular definitions exclude elements such as introns, promotors and other regulatory regions, these are instead thought of as 'associated' with the gene and affect its function.
An even broader operational definition is sometimes used to encompass the complexity of these diverse phenomena, where a gene is defined as a union of genomic sequences encoding a coherent set of potentially overlapping functional products.[33] This definition categorizes genes by their functional products (proteins or RNA) rather than their specific DNA loci, with regulatory elements classified as gene-associated regions.[33]
History
Discovery of discrete inherited units

The existence of discrete inheritable units was first suggested by
Prior to Mendel's work, the dominant theory of heredity was one of
Mendel's work went largely unnoticed after its first publication in 1866, but was rediscovered in the late 19th century by Hugo de Vries, Carl Correns, and Erich von Tschermak, who (claimed to have) reached similar conclusions in their own research.[38] Specifically, in 1889, Hugo de Vries published his book Intracellular Pangenesis,[39] in which he postulated that different characters have individual hereditary carriers and that inheritance of specific traits in organisms comes in particles. De Vries called these units "pangenes" (Pangens in German), after Darwin's 1868 pangenesis theory.
Twenty years later, in 1909, Wilhelm Johannsen introduced the term 'gene'[8] and in 1906, William Bateson, that of 'genetics'[40][33] while Eduard Strasburger, amongst others, still used the term 'pangene' for the fundamental physical and functional unit of heredity.[39]: Translator's preface, viii
Discovery of DNA
Advances in understanding genes and inheritance continued throughout the 20th century.
In the early 1950s the prevailing view was that the genes in a chromosome acted like discrete entities arranged like beads on a string. The experiments of Benzer using mutants defective in the rII region of bacteriophage T4 (1955–1959) showed that individual genes have a simple linear structure and are likely to be equivalent to a linear section of DNA.[45][46]
Collectively, this body of research established the
In 1972, Walter Fiers and his team were the first to determine the sequence of a gene: that of Bacteriophage MS2 coat protein.[47] The subsequent development of chain-termination DNA sequencing in 1977 by Frederick Sanger improved the efficiency of sequencing and turned it into a routine laboratory tool.[48] An automated version of the Sanger method was used in early phases of the Human Genome Project.[49]
Modern synthesis and its successors
The theories developed in the early 20th century to integrate
This view of evolution was emphasized by
The development of the neutral theory of evolution in the late 1960s led to the recognition that random genetic drift is a major player in evolution and that neutral theory should be the null hypothesis of molecular evolution.[53] This led to the construction of phylogenetic trees and the development of the molecular clock, which is the basis of all dating techniques using DNA sequences. These techniques are not confined to molecular gene sequences but can be used on all DNA segments in the genome.
Molecular basis
DNA
The vast majority of organisms encode their genes in long strands of DNA (deoxyribonucleic acid). DNA consists of a chain made from four types of nucleotide subunits, each composed of: a five-carbon sugar (2-deoxyribose), a phosphate group, and one of the four bases adenine, cytosine, guanine, and thymine.[54]: 2.1
Two chains of DNA twist around each other to form a DNA
Due to the chemical composition of the
The
Chromosomes


The total complement of genes in an organism or cell is known as its genome, which may be stored on one or more chromosomes. A chromosome consists of a single, very long DNA helix on which thousands of genes are encoded.[54]: 4.2 The region of the chromosome at which a particular gene is located is called its locus. Each locus contains one allele of a gene; however, members of a population may have different alleles at the locus, each with a slightly different gene sequence.
The majority of
Whereas the chromosomes of prokaryotes are relatively gene-dense, those of eukaryotes often contain regions of DNA that serve no obvious function. Simple single-celled eukaryotes have relatively small amounts of such DNA, whereas the genomes of complex multicellular organisms, including humans, contain an absolute majority of DNA without an identified function.[59] This DNA has often been referred to as "junk DNA". However, more recent analyses suggest that, although protein-coding DNA makes up barely 2% of the human genome, about 80% of the bases in the genome may be expressed, so the term "junk DNA" may be a misnomer.[30]
Structure and function
Structure
|
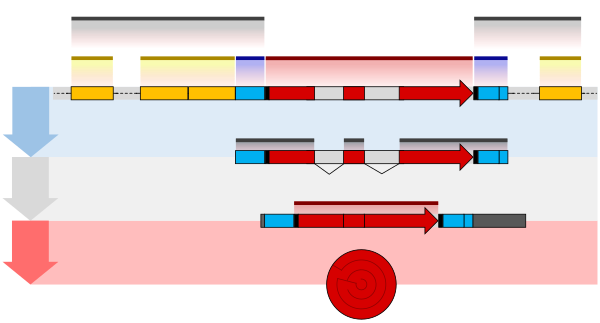
The structure of a protein-coding gene consists of many elements of which the actual protein coding sequence is often only a small part. These include introns and untranslated regions of the mature mRNA. Noncoding genes can also contain introns that are removed during processing to produce the mature functional RNA.
All genes are associated with
Additionally, genes can have regulatory regions many kilobases upstream or downstream of the gene that alter expression. These act by
The mature messenger RNA produced from protein-coding genes contains
Many noncoding genes in eukaryotes have different transcription termination mechanisms and they do not have poly(A) tails.
Many prokaryotic genes are organized into
Complexity
Though many genes have simple structures, as with much of biology, others can be quite complex or represent unusual edge-cases. Eukaryotic genes often have introns are often much larger than their exons,[70][71] and those introns can even have other genes nested inside them.[72] Associated enhancers may be many kilobase away, or even on entirely different chromosomes operating via physical contact between two chromosomes.[73][74] A single gene can encode multiple different functional products by alternative splicing, and conversely gene may be split across chromosomes but those transcripts are concatenated back together into a functional sequence by trans-splicing.[75] It is also possible for overlapping genes to share some of their DNA sequence, either on opposite strands or the same strand (in a different reading frame, or even the same reading frame).[76]
Gene expression
In all organisms, two steps are required to read the information encoded in a gene's DNA and produce the protein it specifies. First, the gene's DNA is
Genetic code
The nucleotide sequence of a gene's DNA specifies the amino acid sequence of a protein through the
Additionally, a "start codon", and three "stop codons" indicate the beginning and end of the protein coding region. There are 64 possible codons (four possible nucleotides at each of three positions, hence 43 possible codons) and only 20 standard amino acids; hence the code is redundant and multiple codons can specify the same amino acid. The correspondence between codons and amino acids is nearly universal among all known living organisms.[79]
Transcription
In
Translation
Regulation
RNA genes
A typical protein-coding gene is first copied into RNA as an intermediate in the manufacture of the final protein product.[54]: 6.1 In other cases, the RNA molecules are the actual functional products, as in the synthesis of ribosomal RNA and transfer RNA. Some RNAs known as ribozymes are capable of enzymatic function, while others such as microRNAs and riboswitches have regulatory roles. The DNA sequences from which such RNAs are transcribed are known as non-coding RNA genes.[77]
Some
Inheritance
Organisms inherit their genes from their parents. Asexual organisms simply inherit a complete copy of their parent's genome. Sexual organisms have two copies of each chromosome because they inherit one complete set from each parent.[54]
Mendelian inheritance
According to
Alleles at a locus may be
DNA replication and cell division
The growth, development, and reproduction of organisms relies on
The rate of DNA replication in living cells was first measured as the rate of phage T4 DNA elongation in phage-infected E. coli and found to be impressively rapid.[87] During the period of exponential DNA increase at 37 °C, the rate of elongation was 749 nucleotides per second.
After DNA replication is complete, the cell must physically separate the two copies of the genome and divide into two distinct membrane-bound cells.
Molecular inheritance
The duplication and transmission of genetic material from one generation of cells to the next is the basis for molecular inheritance and the link between the classical and molecular pictures of genes. Organisms inherit the characteristics of their parents because the cells of the offspring contain copies of the genes in their parents' cells. In
During the process of meiotic cell division, an event called genetic recombination or crossing-over can sometimes occur, in which a length of DNA on one chromatid is swapped with a length of DNA on the corresponding homologous non-sister chromatid. This can result in reassortment of otherwise linked alleles.[54]: 5.5 The Mendelian principle of independent assortment asserts that each of a parent's two genes for each trait will sort independently into gametes; which allele an organism inherits for one trait is unrelated to which allele it inherits for another trait. This is in fact only true for genes that do not reside on the same chromosome or are located very far from one another on the same chromosome. The closer two genes lie on the same chromosome, the more closely they will be associated in gametes and the more often they will appear together (known as genetic linkage).[88] Genes that are very close are essentially never separated because it is extremely unlikely that a crossover point will occur between them.[88]
Molecular evolution
Mutation
DNA replication is for the most part extremely accurate, however errors (
When multiple different alleles for a gene are present in a species's population it is called polymorphic. Most different alleles are functionally equivalent, however some alleles can give rise to different phenotypic traits. A gene's most common allele is called the wild type, and rare alleles are called mutants. The genetic variation in relative frequencies of different alleles in a population is due to both natural selection and genetic drift.[94] The wild-type allele is not necessarily the ancestor of less common alleles, nor is it necessarily fitter.
Most mutations within genes are
Sequence homology
The relationship between genes can be measured by comparing the sequences of their DNA. If the level of similarity exceeds a minimum value, one can conclude that the genes descend from a common ancestor; they are homologous.[95][96] Genes that are related by direct descent from a common ancestor are orthologous genes - they are usually found at the same locus in different species. Genes that are related as a result of a gene duplication event are parologous genes.[97][98]
It is often assumed that the functions of orthologous genes are more similar than those of paralogous genes, although the difference is minimal.[99][100]
Origins of new genes
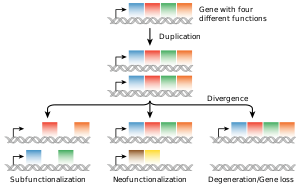
The most common source of new genes in eukaryotic lineages is gene duplication, which creates copy number variation of an existing gene in the genome.[101][102] The resulting genes (paralogs) may then diverge in sequence and in function. Sets of genes formed in this way compose a gene family. Gene duplications and losses within a family are common and represent a major source of evolutionary biodiversity.[103] Sometimes, gene duplication may result in a nonfunctional copy of a gene, or a functional copy may be subject to mutations that result in loss of function; such nonfunctional genes are called pseudogenes.[54]: 7.6
Genome
The
Number of genes
The genome size, and the number of genes it encodes varies widely between organisms. The smallest genomes occur in viruses,[122] and viroids (which act as a single non-coding RNA gene).[123] Conversely, plants can have extremely large genomes,[124] with rice containing >46,000 protein-coding genes.[118] The total number of protein-coding genes (the Earth's proteome) is estimated to be 5 million sequences.[125]
Although the number of base-pairs of DNA in the human genome has been known since the 1950s, the estimated number of genes has changed over time as definitions of genes, and methods of detecting them have been refined. Initial theoretical predictions of the number of human genes in the 1960s and 1970s were based on mutation load estimates and the numbers of mRNAs and these estimates tended to be about 30,000 protein-coding genes.[126][127][128] During the 1990s there were guesstimates of up to 100,000 genes and early data on detection of mRNAs (expressed sequence tags) suggested more than the traditional value of 30,000 genes that had been reported in the textbooks during the 1980s.[129]
The initial draft sequences of the human genome confirmed the earlier predictions of about 30,000 protein-coding genes however that estimate has fallen to about 19,000 with the ongoing GENCODE annotation project .[130] The number of noncoding genes is not known with certainty but the latest estimates from Ensembl suggest 26,000 noncoding genes.[131]
Essential genes
Essential genes are the set of genes thought to be critical for an organism's survival.
Essential genes include
Genetic and genomic nomenclature
Gene nomenclature has been established by the HUGO Gene Nomenclature Committee (HGNC), a committee of the Human Genome Organisation, for each known human gene in the form of an approved gene name and symbol (short-form abbreviation), which can be accessed through a database maintained by HGNC. Symbols are chosen to be unique, and each gene has only one symbol (although approved symbols sometimes change). Symbols are preferably kept consistent with other members of a gene family and with homologs in other species, particularly the mouse due to its role as a common model organism.[141]
Genetic engineering
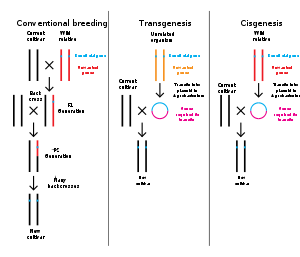
Genetic engineering is the modification of an organism's
Genetic engineering is now a routine research tool with
For multicellular organisms, typically the embryo is engineered which grows into the adult genetically modified organism.[151] However, the genomes of cells in an adult organism can be edited using gene therapy techniques to treat genetic diseases.
See also
- Epigenetics
- Full genome sequencing
- Gene-centric view of evolution
- Gene dosage
- Gene patent
- Gene pool
- Gene redundancy
- Gene silencing
- Genetic algorithm
- Haplotype
- List of gene prediction software
- List of notable genes
- Predictive medicine
- Quantitative trait locus
- Selfish genetic element
References
Citations
- ^ a b "1909: The Word Gene Coined". genome.gov. Retrieved 8 March 2021. "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..."
- PMID 31258451.
- ^ S2CID 24583286.
- ^ "What is a gene?: MedlinePlus Genetics". MedlinePlus. 17 September 2020. Retrieved 4 January 2021.
- OCLC 50166721.
- ^ "Studying Genes". nigms.nih.gov. Archived from the original on 17 January 2021. Retrieved 15 January 2021.
- PMID 22307690.
- ^ a b Johannsen W (1909). Elemente der exakten Erblichkeitslehre [Elements of the exact theory of heredity] (in German). Jena, Germany: Gustav Fischer. p. 124. From p. 124: "Dieses "etwas" in den Gameten bezw. in der Zygote, … – kurz, was wir eben Gene nennen wollen – bedingt sind." (This "something" in the gametes or in the zygote, which has crucial importance for the character of the organism, is usually called by the quite ambiguous term Anlagen [primordium, from the German word Anlage for "plan, arrangement ; rough sketch"]. Many other terms have been suggested, mostly unfortunately in closer connection with certain hypothetical opinions. The word "pangene", which was introduced by Darwin, is perhaps used most frequently in place of Anlagen. However, the word "pangene" was not well chosen, as it is a compound word containing the roots pan (the neuter form of Πας all, every) and gen (from γί-γ(ε)ν-ομαι, to become). Only the meaning of this latter [i.e., gen] comes into consideration here ; just the basic idea – [namely,] that a trait in the developing organism can be determined or is influenced by "something" in the gametes – should find expression. No hypothesis about the nature of this "something" should be postulated or supported by it. For that reason it seems simplest to use in isolation the last syllable gen from Darwin's well-known word, which alone is of interest to us, in order to replace, with it, the poor, ambiguous word Anlage. Thus we will say simply "gene" and "genes" for "pangene" and "pangenes". The word gene is completely free of any hypothesis ; it expresses only the established fact that in any case many traits of the organism are determined by specific, separable, and thus independent "conditions", "foundations", "plans" – in short, precisely what we want to call genes.)
- ^ a b c d Kampourakis K (2017). Making Sense of Genes. Cambridge, UK: Cambridge University Press.
- S2CID 144613322.
- ^ Meunier R (2022). "Stanford Encyclopedia of Philosophy: Gene". Stanford Encyclopedia of Philosophy. Retrieved 28 February 2023.
- PMID 24753594.
- ^ a b Dawkins R (1976). The selfish gene. Oxford, UK: Oxford University Press.
- PMID 15791804.
- PMID 16588492.
- PMID 15020400.
- ^ Judson HF (1996). The Eight Day of Creation (Expanded ed.). Plainview, NY (US): Cold Spring Harbor Laboratory Press.
- ^ a b Watson JD (1965). Molecular Biology of the Gene. New York, NY, US: W.A. Benjamin, Inc.
- ISBN 0-8153-1619-4.
- ^ Moran LA, Horton HR, Scrimgeour KG, Perry MD (2012). Principles of Biochemistry: Fifth Edition. Upper Saddle River, NJ, US: Pearson.
- ^ Lewin B (2004). Genes VIII. Upper Saddle River, NJ, US: Pearson/Prentice Hall.
- PMID 30813969.
- PMID 25710723.
- PMID 28633296.
- ^ Mortola E, Long M (2021). "Turning Junk into Us: How Genes Are Born". American Scientist. 109: 174–182.
- S2CID 88157272.
- S2CID 4420674.
- S2CID 36463252.
- PMID 30482837.
- ^ S2CID 36463252.
- S2CID 30948765.
- PMID 16344564.
- ^ PMID 17567988.
- PMID 18559318.
- ^ "Blending Inheritance - an overview | ScienceDirect Topics".
- ^ "genesis". Oxford English Dictionary (Online ed.). Oxford University Press. (Subscription or participating institution membership required.)
- ISBN 978-0-203-91100-6.
- ISBN 978-0395-97765-1.
- ^ a b de Vries H (1889). Intracellulare Pangenese [Intracellular Pangenesis] (in German). Translated by Gager CS. Jena: Verlag von Gustav Fischer. Translated in 1908 from German to English by Open Court Publishing Co., Chicago, 1910
- ^ Bateson W (1906). "The progress of genetic research". In Wilks W (ed.). Report of the Third International Conference 1906 on Genetics. London, England: Royal Horticultural Society. pp. 90–97.
… the science itself [i.e. the study of the breeding and hybridisation of plants] is still nameless, and we can only describe our pursuit by cumbrous and often misleading periphrasis. To meet this difficulty I suggest for the consideration of this Congress the term Genetics, which sufficiently indicates that our labors are devoted to the elucidation of the phenomena of heredity and variation: in other words, to the physiology of Descent, with implied bearing on the theoretical problems of the evolutionist and the systematist, and application to the practical problems of breeders, whether of animals or plants.
- PMID 33226.
- PMID 12981234.
- ISBN 978-0-87969-477-7.
- S2CID 4253007.
- PMID 16589677.
- PMID 16590553.
- S2CID 4153893.
- PMID 271968.
- ^ Adams JU (2008). "DNA Sequencing Technologies". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 193.
- ISBN 978-0262513661.
- ISBN 9781400820108.
- ISBN 978-0-19-286088-0.
- ^ Duret L (2008). "Neutral Theory: The Null Hypothesis of Molecular Evolution". Nature Education. 1: 218.
- ^ ISBN 978-0-8153-3218-3.
- ISBN 978-0-7167-4955-4.
- PMID 15839726.

- PMID 16540631.
- ^ PMID 18193080.
- PMID 15496913.
- ^ ISSN 2002-4436.
- S2CID 205418589.
- PMID 23503198.
- PMID 16719718.
- PMID 11897027.
- S2CID 5808466.
- PMID 37035540.
- PMID 10823905.
- PMID 15642184.
- S2CID 19804795.
- PMID 17070957.
- PMID 15781704.
- PMID 19542305..
- S2CID 1755326.
- PMID 20236724.
- PMID 26966239.
- PMID 34611352.
- ^ S2CID 18347629.
- S2CID 4276146.
- PMID 13882204.
- PMID 9573040.
- S2CID 19804795.
- PMID 8269709.
- ISBN 978-0470016176.
- S2CID 20865732.
- ^ Miko I (2008). "Gregor Mendel and the Principles of Inheritance". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 134.
- ^ Chial H (2008). "Mendelian Genetics: Patterns of Inheritance and Single-Gene Disorders". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 63.
- PMID 789903.
- ^ a b Lobo I, Shaw K (2008). "Discovery and Types of Genetic Linkage". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 139.
- PMID 10978293.
- PMID 20220176.
- PMID 9560386.
- ^ Pyeritz, Reed E., Bruce R. Korf, and Wayne W. Grody, eds. Emery and Rimoin’s principles and practice of medical genetics and genomics: foundations. Academic Press, 2018.
- ^ "What kinds of gene mutations are possible?". Genetics Home Reference. United States National Library of Medicine. 11 May 2015. Retrieved 19 May 2015.
- ^ Andrews CA (2010). "Natural Selection, Genetic Drift, and Gene Flow Do Not Act in Isolation in Natural Populations". Nature Education Knowledge. SciTable. 3 (10). Nature Publishing Group: 5.
- PMID 3065587.
- ISBN 9781605354699.
- ISBN 9781605354699.
- PMID 11532207.
- PMID 19368988.
- PMID 22615551.

- ^ PMID 22102832.

- PMID 25646380.
- PMID 17183716.

- PMID 19726446.
- PMID 22102831.

- PMID 26323763.
- PMID 23433480.
- PMID 21298028.

- S2CID 85739173.
- S2CID 213613.
- S2CID 5502148.
- ISBN 0-06-019497-9
- PMID 33879057.
- ^ Watson, JD, Baker TA, Bell SP, Gann A, Levine M, Losick R. (2004). "Ch9-10", Molecular Biology of the Gene, 5th ed., Peason Benjamin Cummings; CSHL Press.
- ^ "Integr8 – A.thaliana Genome Statistics".
- ^ "Understanding the Basics". The Human Genome Project. Retrieved 26 April 2015.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Archived from the original on 28 November 2013. Retrieved 19 November 2013.
- ^ S2CID 208529258.
- S2CID 4355527.
- PMID 10731132.
- PMID 20441615.
- PMID 20861255.
- .
- .
- PMID 18059312.
- S2CID 84202145.
- PMID 5065367.
- S2CID 202732556.
- S2CID 22619.
- PMID 27734873.
- ^ "Human assembly and gene annotation". Ensembl. 2022. Retrieved 28 February 2023.
- ^ PMID 27013737.
- PMID 16407165.
- PMID 13129938.
- PMID 16738554.
- ^ PMID 25092907.
- PMID 16504025.

- PMID 23675308.

- PMID 23810203.
- PMID 16844256.
- ^ "About the HGNC". HGNC Database of Human Gene Names. HUGO Gene Nomenclature Committee. Archived from the original on 26 March 2023. Retrieved 14 May 2015.
- PMID 4576014.
- PMID 23340847.
- PMID 23084873.
- PMID 24166445.
- PMID 24157548.
- PMID 22818777.
- PMID 20061565.
- PMID 15340423.
- S2CID 34470273.
- PMID 17998949.
Sources
- Main textbook
- ISBN 978-0-8153-3218-3. – A molecular biology textbook available free online through NCBI Bookshelf.

Further reading
- ISBN 978-0-321-90537-6.
- ISBN 978-0-19-286092-7.
- ISBN 978-0-00-763573-3.
- Brown T (2002). Genomes (2nd ed.). New York: Wiley-Liss. PMID 20821850.
External links
- Comparative Toxicogenomics Database
- DNA From The Beginning – a primer on genes and DNA
- Entrez Gene – a searchable database of genes
- Genes – an Open Access journal
- IDconverter – converts gene IDs between public databases
- iHOP – Information Hyperlinked over Proteins
- TranscriptomeBrowser – Gene expression profile analysis
- The Protein Naming Utility, a database to identify and correct deficient gene names
- IMPC (International Mouse Phenotyping Consortium) – Encyclopedia of mammalian gene function
- Global Genes Project – Leading non-profit organization supporting people living with genetic diseases
- ENCODE threads Explorer Characterization of intergenic regions and gene definition. Nature