Gene duplication
Gene duplication (or chromosomal duplication or gene amplification) is a major mechanism through which new genetic material is generated during
Mechanisms of duplication
Ectopic recombination
Duplications arise from an event termed
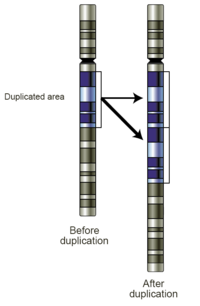
Replication slippage
Retrotransposition
Retrotransposons, mainly L1, can occasionally act on cellular mRNA. Transcripts are reverse transcribed to DNA and inserted into random place in the genome, creating retrogenes. Resulting sequence usually lack introns and often contain poly(A) sequences that are also integrated into the genome. Many retrogenes display changes in gene regulation in comparison to their parental gene sequences, which sometimes results in novel functions. Retrogenes can move between different chromosomes to shape chromosomal evolution.[3]
Aneuploidy
Aneuploidy occurs when nondisjunction at a single chromosome results in an abnormal number of chromosomes. Aneuploidy is often harmful and in mammals regularly leads to spontaneous abortions (miscarriages). Some aneuploid individuals are viable, for example trisomy 21 in humans, which leads to Down syndrome. Aneuploidy often alters gene dosage in ways that are detrimental to the organism; therefore, it is unlikely to spread through populations.
Polyploidy
Polyploidy, or whole genome duplication is a product of nondisjunction during meiosis which results in additional copies of the entire genome. Polyploidy is common in plants, but it has also occurred in animals, with two rounds of whole genome duplication (2R event) in the vertebrate lineage leading to humans.[4] It has also occurred in the hemiascomycete yeasts ~100 mya.[5][6]
After a whole genome duplication, there is a relatively short period of genome instability, extensive gene loss, elevated levels of nucleotide substitution and regulatory network rewiring.[7][8] In addition, gene dosage effects play a significant role.[9] Thus, most duplicates are lost within a short period, however, a considerable fraction of duplicates survive.[10] Interestingly, genes involved in regulation are preferentially retained.[11][12] Furthermore, retention of regulatory genes, most notably the Hox genes, has led to adaptive innovation.
Rapid evolution and functional divergence have been observed at the level of the transcription of duplicated genes, usually by point mutations in short transcription factor binding motifs.[13][14] Furthermore, rapid evolution of protein phosphorylation motifs, usually embedded within rapidly evolving intrinsically disordered regions is another contributing factor for survival and rapid adaptation/neofunctionalization of duplicate genes.[15] Thus, a link seems to exist between gene regulation (at least at the post-translational level) and genome evolution.[15]
Polyploidy is also a well known source of speciation, as offspring, which have different numbers of chromosomes compared to parent species, are often unable to interbreed with non-polyploid organisms. Whole genome duplications are thought to be less detrimental than aneuploidy as the relative dosage of individual genes should be the same.
As an evolutionary event

Rate of gene duplication
Comparisons of genomes demonstrate that gene duplications are common in most species investigated. This is indicated by variable copy numbers (copy number variation) in the genome of humans[16][17] or fruit flies.[18] However, it has been difficult to measure the rate at which such duplications occur. Recent studies yielded a first direct estimate of the genome-wide rate of gene duplication in C. elegans, the first multicellular eukaryote for which such as estimate became available. The gene duplication rate in C. elegans is on the order of 10−7 duplications/gene/generation, that is, in a population of 10 million worms, one will have a gene duplication per generation. This rate is two orders of magnitude greater than the spontaneous rate of point mutation per nucleotide site in this species.[19] Older (indirect) studies reported locus-specific duplication rates in bacteria, Drosophila, and humans ranging from 10−3 to 10−7/gene/generation.[20][21][22]
Neofunctionalization
Gene duplications are an essential source of genetic novelty that can lead to evolutionary innovation. Duplication creates genetic redundancy, where the second copy of the gene is often free from
Gene duplication is believed to play a major role in evolution; this stance has been held by members of the scientific community for over 100 years.[25] Susumu Ohno was one of the most famous developers of this theory in his classic book Evolution by gene duplication (1970).[26] Ohno argued that gene duplication is the most important evolutionary force since the emergence of the universal common ancestor.[27] Major
Subfunctionalization
Another possible fate for duplicate genes is that both copies are equally free to accumulate degenerative mutations, so long as any defects are complemented by the other copy. This leads to a neutral "subfunctionalization" (a process of constructive neutral evolution) or DDC (duplication-degeneration-complementation) model,[29][30] in which the functionality of the original gene is distributed among the two copies. Neither gene can be lost, as both now perform important non-redundant functions, but ultimately neither is able to achieve novel functionality.
Subfunctionalization can occur through neutral processes in which mutations accumulate with no detrimental or beneficial effects. However, in some cases subfunctionalization can occur with clear adaptive benefits. If an ancestral gene is pleiotropic and performs two functions, often neither one of these two functions can be changed without affecting the other function. In this way, partitioning the ancestral functions into two separate genes can allow for adaptive specialization of subfunctions, thereby providing an adaptive benefit.[31]
Loss
Often the resulting genomic variation leads to gene dosage dependent neurological disorders such as Rett-like syndrome and Pelizaeus–Merzbacher disease.[32] Such detrimental mutations are likely to be lost from the population and will not be preserved or develop novel functions. However, many duplications are, in fact, not detrimental or beneficial, and these neutral sequences may be lost or may spread through the population through random fluctuations via genetic drift.
Identifying duplications in sequenced genomes
Criteria and single genome scans
The two genes that exist after a gene duplication event are called
It is important (but often difficult) to differentiate between paralogs and orthologs in biological research. Experiments on human gene function can often be carried out on other
Paralogs can be identified in single genomes through a sequence comparison of all annotated gene models to one another. Such a comparison can be performed on translated amino acid sequences (e.g. BLASTp, tBLASTx) to identify ancient duplications or on DNA nucleotide sequences (e.g. BLASTn, megablast) to identify more recent duplications. Most studies to identify gene duplications require reciprocal-best-hits or fuzzy reciprocal-best-hits, where each paralog must be the other's single best match in a sequence comparison.[33]
Most gene duplications exist as
Genomic microarrays detect duplications
Technologies such as genomic
Next generation sequencing
Gene duplications can also be identified through the use of next-generation sequencing platforms. The simplest means to identify duplications in genomic resequencing data is through the use of paired-end sequencing reads. Tandem duplications are indicated by sequencing read pairs which map in abnormal orientations. Through a combination of increased sequence coverage and abnormal mapping orientation, it is possible to identify duplications in genomic sequencing data.
Nomenclature
The
As amplification
Gene duplication does not necessarily constitute a lasting change in a species' genome. In fact, such changes often don't last past the initial host organism. From the perspective of
Role in cancer
Duplications of
| Cancer type | Associated gene amplifications |
Prevalence of amplification in cancer type (percent) |
|---|---|---|
| Breast cancer | MYC | 20%[39] |
ERBB2 (HER2 ) |
20%[39] | |
CCND1 (Cyclin D1 ) |
15–20%[39] | |
FGFR1 |
12%[39] | |
FGFR2 |
12%[39] | |
| Cervical cancer | MYC | 25–50%[39] |
ERBB2 |
20%[39] | |
| Colorectal cancer | HRAS | 30%[39] |
| KRAS | 20%[39] | |
| MYB | 15–20%[39] | |
| Esophageal cancer | MYC | 40%[39] |
CCND1 |
25%[39] | |
MDM2 |
13%[39] | |
Gastric cancer |
CCNE (Cyclin E) | 15%[39] |
| KRAS | 10%[39] | |
MET |
10%[39] | |
| Glioblastoma | ERBB1 (EGFR) | 33–50%[39] |
CDK4 |
15%[39] | |
| Head and neck cancer | CCND1 |
50%[39] |
| ERBB1 | 10%[39] | |
| MYC | 7–10%[39] | |
Hepatocellular cancer |
CCND1 |
13%[39] |
| Neuroblastoma | MYCN |
20–25%[39] |
| Ovarian cancer | MYC | 20–30%[39] |
ERBB2 |
15–30%[39] | |
| AKT2 | 12%[39] | |
| Sarcoma | MDM2 |
10–30%[39] |
CDK4 |
10%[39] | |
Small cell lung cancer |
MYC | 15–20%[39] |
Whole-genome duplications are also frequent in cancers, detected in 30% to 36% of tumors from the most common cancer types.[40][41] Their exact role in carcinogenesis is unclear, but they in some cases lead to loss of chromatin segregation leading to chromatin conformation changes that in turn lead to oncogenic epigenetic and transcriptional modifications.[42]
See also
References
- .
- ^ "Definition of Gene duplication". medterms medical dictionary. MedicineNet. 2012-03-19.
- PMID 35741730.
- PMID 16128622.
- S2CID 4307263.
- S2CID 4422074.
- S2CID 10054182.
- PMID 16555924.
- S2CID 4382441.
- PMID 11073452.
- PMID 16818725.
- PMID 16098632.
- PMID 16507168.
- PMID 16140417.
- ^ PMID 20080574.
- S2CID 20357402.
- PMID 15286789.
- S2CID 206512885.
- PMID 21295484.
- PMID 6789329.
- PMID 19114461.
- PMID 18059269.
- PMID 17233905.
- S2CID 1240225.
- PMID 15568988.
- ISBN 978-0-04-575015-3.
- ISBN 978-91-554-5776-1.
- S2CID 4422074.
- PMID 10101175.
- S2CID 1743092.
- S2CID 418964.
- S2CID 22412305.
- PMID 17997610.
- PMID 16240409.
- PMID 15647348.
- ^ "ISCN Symbols and Abbreviated Terms". Coriell Institute for Medical Research. Retrieved 2022-10-27.
- OMIM. Updated : 4/23/2014
- PMID 35030162.
- ^ ISBN 978-0-07-137050-9.
- PMID 30013179.
- PMID 33505027.
- PMID 36922594.
