Gene prediction
In
In its earliest days, "gene finding" was based on painstaking experimentation on living cells and organisms. Statistical analysis of the rates of
Determining that a sequence is functional should be distinguished from determining the function of the gene or its product. Predicting the function of a gene and confirming that the gene prediction is accurate still demands in vivo experimentation[1] through gene knockout and other assays, although frontiers of bioinformatics research [2] are making it increasingly possible to predict the function of a gene based on its sequence alone.
Gene prediction is one of the key steps in
Gene prediction is closely related to the so-called 'target search problem' investigating how
.Empirical methods
In empirical (similarity, homology or evidence-based) gene finding systems, the target genome is searched for sequences that are similar to extrinsic evidence in the form of the known
A high degree of similarity to a known messenger RNA or protein product is strong evidence that a region of a target genome is a protein-coding gene. However, to apply this approach systemically requires extensive sequencing of mRNA and protein products. Not only is this expensive, but in complex organisms, only a subset of all genes in the organism's genome are expressed at any given time, meaning that extrinsic evidence for many genes is not readily accessible in any single cell culture. Thus, to collect extrinsic evidence for most or all of the genes in a complex organism requires the study of many hundreds or thousands of cell types, which presents further difficulties. For example, some human genes may be expressed only during development as an embryo or fetus, which might be difficult to study for ethical reasons.
Despite these difficulties, extensive transcript and protein sequence databases have been generated for human as well as other important model organisms in biology, such as mice and yeast. For example, the
New high-throughput
Major challenges involved in gene prediction involve dealing with sequencing errors in raw DNA data, dependence on the quality of the sequence assembly, handling short reads, frameshift mutations, overlapping genes and incomplete genes.
In prokaryotes it's essential to consider horizontal gene transfer when searching for gene sequence homology. An additional important factor underused in current gene detection tools is existence of gene clusters — operons (which are functioning units of DNA containing a cluster of genes under the control of a single promoter) in both prokaryotes and eukaryotes. Most popular gene detectors treat each gene in isolation, independent of others, which is not biologically accurate.
Ab initio methods
Ab Initio gene prediction is an intrinsic method based on gene content and signal detection. Because of the inherent expense and difficulty in obtaining extrinsic evidence for many genes, it is also necessary to resort to
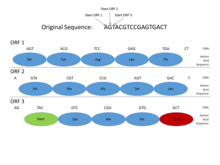
In the genomes of
Ab initio gene finding in
Second,
Advanced gene finders for both prokaryotic and eukaryotic genomes typically use complex
Ab Initio methods have been benchmarked, with some approaching 100% sensitivity,[3] however as the sensitivity increases, accuracy suffers as a result of increased false positives.
Other signals
Among the derived signals used for prediction are statistics resulting from the sub-sequence statistics like
It has been suggested that signals other than those directly detectable in sequences may improve gene prediction. For example, the role of
Neural networks
Combined approaches
Programs such as Maker combine extrinsic and ab initio approaches by mapping protein and EST data to the genome to validate ab initio predictions. Augustus, which may be used as part of the Maker pipeline, can also incorporate hints in the form of EST alignments or protein profiles to increase the accuracy of the gene prediction.
Comparative genomics approaches
As the entire genomes of many different species are sequenced, a promising direction in current research on gene finding is a comparative genomics approach.
This is based on the principle that the forces of natural selection cause genes and other functional elements to undergo mutation at a slower rate than the rest of the genome, since mutations in functional elements are more likely to negatively impact the organism than mutations elsewhere. Genes can thus be detected by comparing the genomes of related species to detect this evolutionary pressure for conservation. This approach was first applied to the mouse and human genomes, using programs such as SLAM, SGP and TWINSCAN/N-SCAN and CONTRAST.[20]
Multiple informants
TWINSCAN examined only human-mouse synteny to look for orthologous genes. Programs such as N-SCAN and CONTRAST allowed the incorporation of alignments from multiple organisms, or in the case of N-SCAN, a single alternate organism from the target. The use of multiple informants can lead to significant improvements in accuracy.[20]
CONTRAST is composed of two elements. The first is a smaller classifier, identifying donor splice sites and acceptor splice sites as well as start and stop codons. The second element involves constructing a full model using machine learning. Breaking the problem into two means that smaller targeted data sets can be used to train the classifiers, and that classifier can operate independently and be trained with smaller windows. The full model can use the independent classifier, and not have to waste computational time or model complexity re-classifying intron-exon boundaries. The paper in which CONTRAST is introduced proposes that their method (and those of TWINSCAN, etc.) be classified as de novo gene assembly, using alternate genomes, and identifying it as distinct from ab initio, which uses a target 'informant' genomes.[20]
Comparative gene finding can also be used to project high quality annotations from one genome to another. Notable examples include Projector, GeneWise, GeneMapper and GeMoMa. Such techniques now play a central role in the annotation of all genomes.
Pseudogene prediction
Sequence similarity methods can be customised for pseudogene prediction using additional filtering to find candidate pseudogenes. This could use disablement detection, which looks for nonsense or frameshift mutations that would truncate or collapse an otherwise functional coding sequence.[22] Additionally, translating DNA into proteins sequences can be more effective than just straight DNA homology.[21]
Content sensors can be filtered according to the differences in statistical properties between pseudogenes and genes, such as a reduced count of CpG islands in pseudogenes, or the differences in G-C content between pseudogenes and their neighbours. Signal sensors also can be honed to pseudogenes, looking for the absence of introns or polyadenine tails. [23]
Metagenomic gene prediction
Metagenomics is the study of genetic material recovered from the environment, resulting in sequence information from a pool of organisms. Predicting genes is useful for comparative metagenomics.
Metagenomics tools also fall into the basic categories of using either sequence similarity approaches (MEGAN4) and ab initio techniques (GLIMMER-MG).
Glimmer-MG[24] is an extension to GLIMMER that relies mostly on an ab initio approach for gene finding and by using training sets from related organisms. The prediction strategy is augmented by classification and clustering gene data sets prior to applying ab initio gene prediction methods. The data is clustered by species. This classification method leverages techniques from metagenomic phylogenetic classification. An example of software for this purpose is, Phymm, which uses interpolated markov models—and PhymmBL, which integrates BLAST into the classification routines.
MEGAN4[25] uses a sequence similarity approach, using local alignment against databases of known sequences, but also attempts to classify using additional information on functional roles, biological pathways and enzymes. As in single organism gene prediction, sequence similarity approaches are limited by the size of the database.
FragGeneScan and MetaGeneAnnotator are popular gene prediction programs based on Hidden Markov model. These predictors account for sequencing errors, partial genes and work for short reads.
Another fast and accurate tool for gene prediction in metagenomes is MetaGeneMark.[26] This tool is used by the DOE Joint Genome Institute to annotate IMG/M, the largest metagenome collection to date.
See also
- List of gene prediction software
- Phylogenetic footprinting
- Protein function prediction
- Protein structure prediction
- Protein–protein interaction prediction
- Pseudogene (database)
- Sequence mining
- Sequence similarity (homology)
References
- PMID 20430068.
- PMID 32962098.
- ^ S2CID 3352427.
- PMID 24187380.
- PMID 15908574.
- ISBN 9780321897398.
- ^ "GeneMark-ES".
- PMID 15144565.
- PMID 17319737.
- PMID 18096039.
- PMID 19494180.
- PMID 17204465.
- PMID 16987907.
- PMID 11928478.
- PMID 16386465.
- PMID 16772025.
- ^ Rogic, S (2006). The role of pre-mRNA secondary structure in gene splicing in Saccharomyces cerevisiae (PDF) (PhD thesis). University of British Columbia. Archived from the original (PDF) on 2009-05-30. Retrieved 2007-04-01.
- PMID 23529114.
- ISBN 978-3-642-02503-7.
- ^ PMID 18096039.
- ^ S2CID 6617359.
- PMID 16680195.
- PMID 15261647.
- PMID 22102569.
- PMID 21690186.
- PMID 20403810.
External links
- Augustus
- FGENESH
- GeMoMa - Homology-based gene prediction based on amino acid and intron position conservation as well as RNA-Seq data
- geneid, SGP2
- Glimmer Archived 2011-08-26 at the Wayback Machine, GlimmerHMM Archived 2011-08-18 at the Wayback Machine
- GenomeThreader
- ChemGenome
- GeneMark
- Gismo
- mGene
- StarORF — A multi-platform and web tool for predicting ORFs and obtaining reverse complement sequence
- Maker - A portable and easily configurable genome annotation pipeline
