Gene expression
| Part of a series on |
| Biochemistry |
|---|
 |
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, proteins or non-coding RNA, and ultimately affect a phenotype. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. The process of gene expression is used by all known life—
machinery for life.In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, i.e. observable trait. The genetic information stored in DNA represents the genotype, whereas the phenotype results from the "interpretation" of that information. Such phenotypes are often displayed by the synthesis of proteins that control the organism's structure and development, or that act as enzymes catalyzing specific metabolic pathways.
All steps in the gene expression process may be modulated (regulated), including the
Mechanism
Transcription

The production of a RNA copy from a DNA strand is called transcription, and is performed by RNA polymerases, which add one ribonucleotide at a time to a growing RNA strand as per the complementarity law of the nucleotide bases. This RNA is complementary to the template 3′ → 5′ DNA strand,[1] with the exception that thymines (T) are replaced with uracils (U) in the RNA and possible errors.
mRNA processing
While transcription of prokaryotic protein-coding genes creates
The processing of pre-mRNA include 5′ capping, which is set of enzymatic reactions that add
Another modification is 3′ cleavage and polyadenylation. They occur if polyadenylation signal sequence (5′- AAUAAA-3′) is present in pre-mRNA, which is usually between protein-coding sequence and terminator. The pre-mRNA is first cleaved and then a series of ~200 adenines (A) are added to form poly(A) tail, which protects the RNA from degradation. The poly(A) tail is bound by multiple poly(A)-binding proteins (PABPs) necessary for mRNA export and translation re-initiation. In the inverse process of deadenylation, poly(A) tails are shortened by the CCR4-Not 3′-5′ exonuclease, which often leads to full transcript decay.
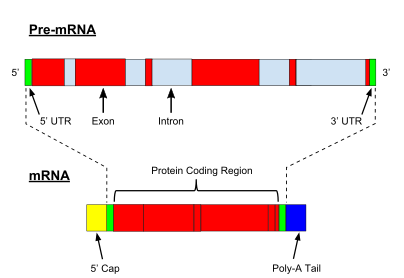
A very important modification of eukaryotic pre-mRNA is
Extensive RNA processing may be an
Non-coding RNA maturation
In most organisms
In the case of transfer RNA (tRNA), for example, the 5′ sequence is removed by
Even snRNAs and snoRNAs themselves undergo series of modification before they become part of functional RNP complex. This is done either in the nucleoplasm or in the specialized compartments called Cajal bodies. Their bases are methylated or pseudouridinilated by a group of small Cajal body-specific RNAs (scaRNAs), which are structurally similar to snoRNAs.
RNA export
In eukaryotes most mature RNA must be exported to the cytoplasm from the nucleus. While some RNAs function in the nucleus, many RNAs are transported through the nuclear pores and into the cytosol.[7] Export of RNAs requires association with specific proteins known as exportins. Specific exportin molecules are responsible for the export of a given RNA type. mRNA transport also requires the correct association with Exon Junction Complex (EJC), which ensures that correct processing of the mRNA is completed before export. In some cases RNAs are additionally transported to a specific part of the cytoplasm, such as a synapse; they are then towed by motor proteins that bind through linker proteins to specific sequences (called "zipcodes") on the RNA.[8]
Translation
For some non-coding RNA, the mature RNA is the final gene product.
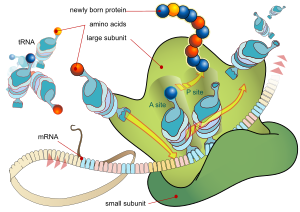
Every mRNA consists of three parts: a 5′ untranslated region (5′UTR), a protein-coding region or
In prokaryotes translation generally occurs at the point of transcription (co-transcriptionally), often using a messenger RNA that is still in the process of being created. In eukaryotes translation can occur in a variety of regions of the cell depending on where the protein being written is supposed to be. Major locations are the cytoplasm for soluble cytoplasmic proteins and the membrane of the endoplasmic reticulum for proteins that are for export from the cell or insertion into a cell membrane. Proteins that are supposed to be produced at the endoplasmic reticulum are recognised part-way through the translation process. This is governed by the signal recognition particle—a protein that binds to the ribosome and directs it to the endoplasmic reticulum when it finds a signal peptide on the growing (nascent) amino acid chain.[14]
Folding
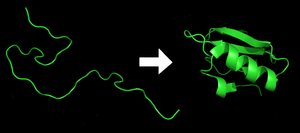
Each
The correct three-dimensional structure is essential to function, although some parts of functional proteins
Enzymes called chaperones assist the newly formed protein to attain (fold into) the 3-dimensional structure it needs to function.[20] Similarly, RNA chaperones help RNAs attain their functional shapes.[21] Assisting protein folding is one of the main roles of the endoplasmic reticulum in eukaryotes.
Translocation
Secretory proteins of eukaryotes or prokaryotes must be translocated to enter the secretory pathway. Newly synthesized proteins are directed to the eukaryotic Sec61 or prokaryotic SecYEG translocation channel by signal peptides. The efficiency of protein secretion in eukaryotes is very dependent on the signal peptide which has been used.[22]
Protein transport
Many proteins are destined for other parts of the cell than the cytosol and a wide range of signalling sequences or (signal peptides) are used to direct proteins to where they are supposed to be. In prokaryotes this is normally a simple process due to limited compartmentalisation of the cell. However, in eukaryotes there is a great variety of different targeting processes to ensure the protein arrives at the correct organelle.
Not all proteins remain within the cell and many are exported, for example,
Regulation of gene expression

Regulation of gene expression is the control of the amount and timing of appearance of the functional product of a gene. Control of expression is vital to allow a cell to produce the gene products it needs when it needs them; in turn, this gives cells the flexibility to adapt to a variable environment, external signals, damage to the cell, and other stimuli. More generally, gene regulation gives the cell control over all structure and function, and is the basis for cellular differentiation, morphogenesis and the versatility and adaptability of any organism.
Numerous terms are used to describe types of genes depending on how they are regulated; these include:
- A constitutive gene is a gene that is transcribed continually as opposed to a facultative gene, which is only transcribed when needed.
- A GAPDH and ubiquitin. Some housekeeping genes are transcribed at a relatively constant rate and these genes can be used as a reference point in experiments to measure the expression rates of other genes.
- A facultative gene is a gene only transcribed when needed as opposed to a constitutive gene.
- An inducible gene is a gene whose expression is either responsive to environmental change or dependent on the position in the cell cycle.
Any step of gene expression may be modulated, from the DNA-RNA transcription step to post-translational modification of a protein. The stability of the final gene product, whether it is RNA or protein, also contributes to the expression level of the gene—an unstable product results in a low expression level. In general gene expression is regulated through changes[25] in the number and type of interactions between molecules[26] that collectively influence transcription of DNA[27] and translation of RNA.[28]
Some simple examples of where gene expression is important are:
- Control of blood glucose regulation.
- mammalsto prevent an "overdose" of the genes it contains.
- Cyclin expression levels control progression through the eukaryotic cell cycle.
Transcriptional regulation

Regulation of transcription can be broken down into three main routes of influence; genetic (direct interaction of a control factor with the gene), modulation interaction of a control factor with the transcription machinery and epigenetic (non-sequence changes in DNA structure that influence transcription).

Direct interaction with DNA is the simplest and the most direct method by which a protein changes transcription levels. Genes often have several protein binding sites around the coding region with the specific function of regulating transcription. There are many classes of regulatory DNA binding sites known as enhancers, insulators and silencers. The mechanisms for regulating transcription are varied, from blocking key binding sites on the DNA for RNA polymerase to acting as an activator and promoting transcription by assisting RNA polymerase binding.
The activity of transcription factors is further modulated by intracellular signals causing protein post-translational modification including phosphorylation, acetylation, or glycosylation. These changes influence a transcription factor's ability to bind, directly or indirectly, to promoter DNA, to recruit RNA polymerase, or to favor elongation of a newly synthesized RNA molecule.
The nuclear membrane in eukaryotes allows further regulation of transcription factors by the duration of their presence in the nucleus, which is regulated by reversible changes in their structure and by binding of other proteins.[29] Environmental stimuli or endocrine signals[30] may cause modification of regulatory proteins[31] eliciting cascades of intracellular signals,[32] which result in regulation of gene expression.
It has become apparent that there is a significant influence of non-DNA-sequence specific effects on transcription. These effects are referred to as epigenetic and involve the higher order structure of DNA, non-sequence specific DNA binding proteins and chemical modification of DNA. In general epigenetic effects alter the accessibility of DNA to proteins and so modulate transcription.
In eukaryotes the structure of chromatin, controlled by the histone code, regulates access to DNA with significant impacts on the expression of genes in euchromatin and heterochromatin areas.
Enhancers, transcription factors, mediator complex and DNA loops in mammalian transcription
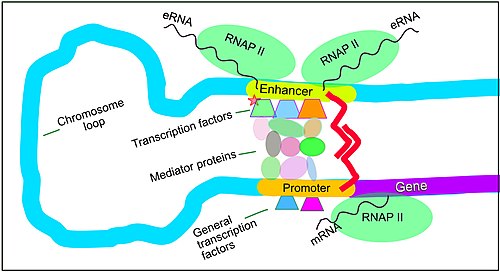
Gene expression in mammals is regulated by many
Enhancers are genome regions that regulate genes. Enhancers control cell-type-specific gene expression programs, most often by looping through long distances to come in physical proximity with the promoters of their target genes.[35] Multiple enhancers, each often tens or hundred of thousands of nucleotides distant from their target genes, loop to their target gene promoters and coordinate with each other to control gene expression.[35]
The illustration shows an enhancer looping around to come into proximity with the promoter of a target gene. The loop is stabilized by a dimer of a connector protein (e.g. dimer of CTCF or YY1). One member of the dimer is anchored to its binding motif on the enhancer and the other member is anchored to its binding motif on the promoter (represented by the red zigzags in the illustration).[36] Several cell function-specific transcription factors (among the about 1,600 transcription factors in a human cell)[37] generally bind to specific motifs on an enhancer.[38] A small combination of these enhancer-bound transcription factors, when brought close to a promoter by a DNA loop, govern transcription level of the target gene. Mediator (a complex usually consisting of about 26 proteins in an interacting structure) communicates regulatory signals from enhancer DNA-bound transcription factors directly to the RNA polymerase II (pol II) enzyme bound to the promoter.[39]
Enhancers, when active, are generally transcribed from both strands of DNA with RNA polymerases acting in two different directions, producing two eRNAs as illustrated in the figure.[40] An inactive enhancer may be bound by an inactive transcription factor. Phosphorylation of the transcription factor may activate it and that activated transcription factor may then activate the enhancer to which it is bound (see small red star representing phosphorylation of transcription factor bound to enhancer in the illustration).[41] An activated enhancer begins transcription of its RNA before activating transcription of messenger RNA from its target gene.[42]
DNA methylation and demethylation in transcriptional regulation
Methylation of cytosine in DNA has a major role in regulating gene expression. Methylation of CpGs in a promoter region of a gene usually represses gene transcription[45] while methylation of CpGs in the body of a gene increases expression.[46] TET enzymes play a central role in demethylation of methylated cytosines. Demethylation of CpGs in a gene promoter by TET enzyme activity increases transcription of the gene.[47]
Transcriptional regulation in learning and memory
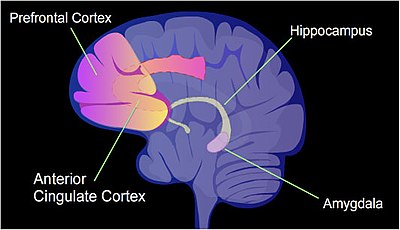
In a rat, contextual fear conditioning (CFC) is a painful learning experience. Just one episode of CFC can result in a life-long fearful memory.[48] After an episode of CFC, cytosine methylation is altered in the promoter regions of about 9.17% of all genes in the hippocampus neuron DNA of a rat.[49] The hippocampus is where new memories are initially stored. After CFC about 500 genes have increased transcription (often due to demethylation of CpG sites in a promoter region) and about 1,000 genes have decreased transcription (often due to newly formed 5-methylcytosine at CpG sites in a promoter region). The pattern of induced and repressed genes within neurons appears to provide a molecular basis for forming the first transient memory of this training event in the hippocampus of the rat brain.[49]
Some specific mechanisms guiding new DNA methylations and new DNA demethylations in the hippocampus during memory establishment have been established (see [50] for summary). One mechanism includes guiding the short isoform of the TET1 DNA demethylation enzyme, TET1s, to about 600 locations on the genome. The guidance is performed by association of TET1s with EGR1 protein, a transcription factor important in memory formation. Bringing TET1s to these locations initiates DNA demethylation at those sites, up-regulating associated genes. A second mechanism involves DNMT3A2, a splice-isoform of DNA methyltransferase DNMT3A, which adds methyl groups to cytosines in DNA. This isoform is induced by synaptic activity, and its location of action appears to be determined by histone post-translational modifications (a histone code). The resulting new messenger RNAs are then transported by messenger RNP particles (neuronal granules) to synapses of the neurons, where they can be translated into proteins affecting the activities of synapses.[50]
In particular, the brain-derived neurotrophic factor gene (BDNF) is known as a "learning gene".[51] After CFC there was upregulation of BDNF gene expression, related to decreased CpG methylation of certain internal promoters of the gene, and this was correlated with learning.[51]
Transcriptional regulation in cancer
The majority of gene promoters contain a CpG island with numerous CpG sites.[52] When many of a gene's promoter CpG sites are methylated the gene becomes silenced.[53] Colorectal cancers typically have 3 to 6 driver mutations and 33 to 66 hitchhiker or passenger mutations.[54] However, transcriptional silencing may be of more importance than mutation in causing progression to cancer. For example, in colorectal cancers about 600 to 800 genes are transcriptionally silenced by CpG island methylation (see regulation of transcription in cancer). Transcriptional repression in cancer can also occur by other epigenetic mechanisms, such as altered expression of microRNAs.[55] In breast cancer, transcriptional repression of BRCA1 may occur more frequently by over-transcribed microRNA-182 than by hypermethylation of the BRCA1 promoter (see Low expression of BRCA1 in breast and ovarian cancers).
Post-transcriptional regulation
In eukaryotes, where export of RNA is required before translation is possible, nuclear export is thought to provide additional control over gene expression. All transport in and out of the nucleus is via the
Expression of a gene coding for a protein is only possible if the messenger RNA carrying the code survives long enough to be translated. In a typical cell, an RNA molecule is only stable if specifically protected from degradation. RNA degradation has particular importance in regulation of expression in eukaryotic cells where mRNA has to travel significant distances before being translated. In eukaryotes, RNA is stabilised by certain post-transcriptional modifications, particularly the
Intentional degradation of mRNA is used not just as a defence mechanism from foreign RNA (normally from viruses) but also as a route of mRNA destabilisation. If an mRNA molecule has a complementary sequence to a small interfering RNA then it is targeted for destruction via the RNA interference pathway.
Three prime untranslated regions and microRNAs
Three prime untranslated regions (3′UTRs) of messenger RNAs (mRNAs) often contain regulatory sequences that post-transcriptionally influence gene expression. Such 3′-UTRs often contain both binding sites for microRNAs (miRNAs) as well as for regulatory proteins. By binding to specific sites within the 3′-UTR, miRNAs can decrease gene expression of various mRNAs by either inhibiting translation or directly causing degradation of the transcript. The 3′-UTR also may have silencer regions that bind repressor proteins that inhibit the expression of a mRNA.
The 3′-UTR often contains microRNA response elements (MREs). MREs are sequences to which miRNAs bind. These are prevalent motifs within 3′-UTRs. Among all regulatory motifs within the 3′-UTRs (e.g. including silencer regions), MREs make up about half of the motifs.
As of 2014, the miRBase web site,[56] an archive of miRNA sequences and annotations, listed 28,645 entries in 233 biologic species. Of these, 1,881 miRNAs were in annotated human miRNA loci. miRNAs were predicted to have an average of about four hundred target mRNAs (affecting expression of several hundred genes).[57] Friedman et al.[57] estimate that >45,000 miRNA target sites within human mRNA 3′UTRs are conserved above background levels, and >60% of human protein-coding genes have been under selective pressure to maintain pairing to miRNAs.
Direct experiments show that a single miRNA can reduce the stability of hundreds of unique mRNAs.[58] Other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).[59][60]
The effects of miRNA dysregulation of gene expression seem to be important in cancer.[61] For instance, in gastrointestinal cancers, nine miRNAs have been identified as epigenetically altered and effective in down regulating DNA repair enzymes.[62]
The effects of miRNA dysregulation of gene expression also seem to be important in neuropsychiatric disorders, such as schizophrenia, bipolar disorder, major depression, Parkinson's disease, Alzheimer's disease and autism spectrum disorders.[63][64]
Translational regulation
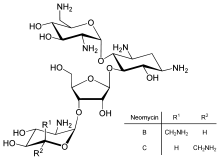
Direct regulation of translation is less prevalent than control of transcription or mRNA stability but is occasionally used. Inhibition of protein translation is a major target for toxins and antibiotics, so they can kill a cell by overriding its normal gene expression control. Protein synthesis inhibitors include the antibiotic neomycin and the toxin ricin.
Post-translational modifications
Post-translational modifications (PTMs) are
PTMs play many important roles in the cell.
Measurement

Measuring gene expression is an important part of many
- Identify viral infection of a cell (viral protein expression).
- Determine an individual's susceptibility to cancer (oncogene expression).
- Find if a bacterium is resistant to penicillin (beta-lactamase expression).
- Gene expression profiling evaluates a panel of genes to help understand the fundamental mechanism of a cell. This is increasingly used in cancer therapy to target specific chemotherapy. (See RNA-Seq and DNA_microarray for details.)
Similarly, the analysis of the location of protein expression is a powerful tool, and this can be done on an organismal or cellular scale. Investigation of localization is particularly important for the study of
mRNA quantification
Levels of mRNA can be quantitatively measured by
Another approach for measuring mRNA abundance is RT-qPCR. In this technique,
For
RNA profiles in Wikipedia
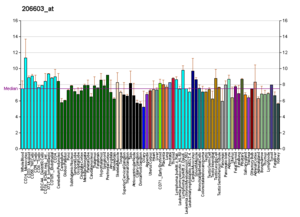
Profiles like these are found for almost all proteins listed in Wikipedia. They are generated by organizations such as the
Protein quantification
For genes encoding proteins, the expression level can be directly assessed by a number of methods with some clear analogies to the techniques for mRNA quantification.
One of the most commonly used methods is to perform a
mRNA-protein correlation
While transcription directly reflects gene expression, the copy number of mRNA molecules does not directly correlate with the number of protein molecules translated from mRNA. Quantification of both protein and mRNA permits a correlation of the two levels. Regulation on each step of gene expression can impact the correlation, as shown for regulation of translation[13] or protein stability.[72] Post-translational factors, such as protein transport in highly polar cells,[73] can influence the measured mRNA-protein correlation as well.
Localization
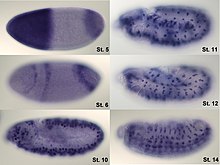
Analysis of expression is not limited to quantification; localization can also be determined. mRNA can be detected with a suitably labelled complementary mRNA strand and protein can be detected via labelled antibodies. The probed sample is then observed by microscopy to identify where the mRNA or protein is.
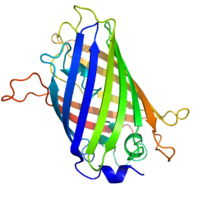
By replacing the gene with a new version fused to a green fluorescent protein marker or similar, expression may be directly quantified in live cells. This is done by imaging using a fluorescence microscope. It is very difficult to clone a GFP-fused protein into its native location in the genome without affecting expression levels, so this method often cannot be used to measure endogenous gene expression. It is, however, widely used to measure the expression of a gene artificially introduced into the cell, for example via an expression vector. By fusing a target protein to a fluorescent reporter, the protein's behavior, including its cellular localization and expression level, can be significantly changed.
The
Expression system
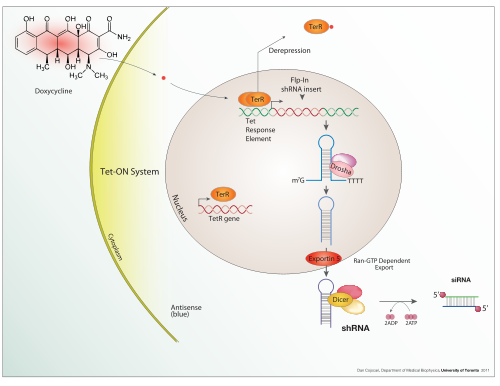
An expression system is a system specifically designed for the production of a gene product of choice. This is normally a protein although may also be RNA, such as
Inducible expression
In nature
In addition to these biological tools, certain naturally observed configurations of DNA (genes, promoters, enhancers, repressors) and the associated machinery itself are referred to as an expression system. This term is normally used in the case where a gene or set of genes is switched on under well defined conditions, for example, the simple repressor switch expression system in
Gene networks
Genes have sometimes been regarded as nodes in a network, with inputs being proteins such as transcription factors, and outputs being the level of gene expression. The node itself performs a function, and the operation of these functions have been interpreted as performing a kind of information processing within cells and determines cellular behavior.
Gene networks can also be constructed without formulating an explicit causal model. This is often the case when assembling networks from large expression data sets.[74] Covariation and correlation of expression is computed across a large sample of cases and measurements (often transcriptome or proteome data). The source of variation can be either experimental or natural (observational). There are several ways to construct gene expression networks, but one common approach is to compute a matrix of all pair-wise correlations of expression across conditions, time points, or individuals and convert the matrix (after thresholding at some cut-off value) into a graphical representation in which nodes represent genes, transcripts, or proteins and edges connecting these nodes represent the strength of association (see GeneNetwork GeneNetwork 2).[75]
Techniques and tools
The following experimental techniques are used to measure gene expression and are listed in roughly chronological order, starting with the older, more established technologies. They are divided into two groups based on their degree of multiplexity.
- Low-to-mid-plex techniques:
- Reporter gene
- Northern blot
- Western blot[76]
- Fluorescent in situ hybridization
- Reverse transcription PCR
- Higher-plex techniques:
Gene expression databases
- Gene expression omnibus (GEO) at NCBI[81]
- Expression Atlas at the EBI
- Bgee SIB Swiss Institute of Bioinformatics
- Mouse Gene Expression Database at the Jackson Laboratory
- CollecTF: a database of experimentally validated transcription factor-binding sites in Bacteria.[82]
- COLOMBOS: collection of bacterial expression compendia.[83]
- Many Microbe Microarrays Database: microbial Affymetrix data[84]
See also
- AlloMap molecular expression testing
- Bookmarking
- EPIC-Seq
- Expressed sequence tag
- Expression Atlas
- Expression profiling
- Gene structure
- Genetic engineering
- Genetically modified organism
- List of biological databases
- List of human genes
- Oscillating gene
- Paramutation
- Protein production
- Protein purification
- Ribonomics
- Ridge
- Sequence profiling tool
- Transcriptional bursting
- Transcriptional noise
- Transcript of unknown function
References
- PMID 19171965.
- )
- PMID 18046571.
- PMID 9759486.
- PMID 17305600.
- PMID 15498478.
- S2CID 10836137.
- PMID 17449729.
- S2CID 206511756.
- PMID 12717454.
- PMID 17574829.
- S2CID 205224972.
- ^ PMID 23407496.
- PMID 18644895.
- ISBN 978-0-8153-3218-3.
- PMID 4565129.
- ISBN 978-0-7167-4684-3.
- S2CID 6451881.
- ISBN 978-0-8153-4129-1.
- PMID 17928587.
- PMID 17981525.
- S2CID 449870.
- PMID 16990376.
- PMID 17786931.
- PMID 15277516.
- PMID 19154003.
- PMID 19274664.
- PMID 18348251.
- PMID 18937349.
- PMID 19182219.
- S2CID 29422790.
- PMID 19319914.
- PMID 33102493.
- S2CID 205485256.
- ^ S2CID 152283312.
- PMID 29224777.
- PMID 29425488.
- PMID 29987030.
- PMID 25693131.
- PMID 29378788.
- PMID 12514134.
- PMID 32810208.
- PMID 26932361.
- PMID 15177689.
- S2CID 22446734.
- PMID 25263941.
- PMID 24108092.
- PMID 16120461.
- ^ PMID 28620075.
- ^ PMID 35098021.
- ^ PMID 28208656.
- PMID 16432200.
- PMID 11782440.
- PMID 23539594.
- PMID 24616890.
- ^ miRBase.org
- ^ PMID 18955434.
- S2CID 4430576.
- S2CID 4429008.
- PMID 18668037.
- PMID 21931505.
- PMID 25987950.
- PMID 22539927.
- PMID 25636176.
- ^ S2CID 32157563.
- PMID 22034591.
- S2CID 205266061.
- PMID 14761301.
- PMID 23887885.
- ^ "Search results < Expression Atlas < EMBL-EBI". www.ebi.ac.uk.
- PMID 31706026.
- PMID 22869793.
- S2CID 58667771.
- PMID 27641093.
- S2CID 20241963.
- PMID 19293442.
- PMID 18796152.
- PMID 18563246.
- PMID 19134166.
- PMID 19087247.
- PMID 27008011.
- PMID 24234444.
- PMID 26586805.
- PMID 17932051.
