GRIA2
| GRIA2 | |||
|---|---|---|---|
Gene ontology | |||
| Molecular function | |||
| Cellular component |
| ||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 4: 157.2 – 157.37 Mb | Chr 3: 80.59 – 80.71 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
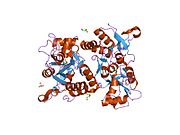
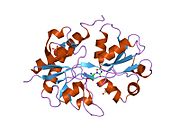
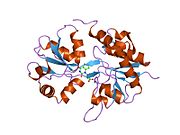
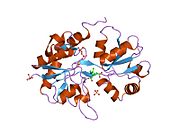
Glutamate ionotropic receptor AMPA type subunit 2 (Glutamate receptor 2, or GluR-2) is a protein that in humans is encoded by the GRIA2 (or GLUR2) gene and it is a subunit found in the AMPA receptors.[5][6][7]
Function
Interactions
GRIA2 has been shown to
RNA editing
Several ion channels and neurotransmitters receptors pre-mRNA as substrates for ADARs. This includes 5 subunits of the glutamate receptor ionotropic AMPA glutamate receptor subunits (Glur2, Glur3, Glur4) and kainate receptor subunits (Glur5, Glur6). Glutamate-gated ion channels are made up of four subunits per channel, with each subunit contributing to the pore loop structure. The pore loop structure is related to that found in K+ channels (e.g., human Kv1.1 channel).[11] The human Kv1.1 channel pre mRNA is also subject to A to I RNA editing.[12] The function of the glutamate receptors is in the mediation of fast neurotransmission to the brain. The diversity of the subunits is determined, as well as RNA splicing by RNA editing events of the individual subunits. This give rise to the necessarily high diversity of these receptors. Glur2 is a gene product of the pre-mRNA of the GRIA2 gene and subject to RNA editing.
Type
The type of RNA editing that occurs in the pre-mRNA of GluR-2 is Adenosine-to-Inosine (A-to-I) editing. [11] A-to-I RNA editing is catalyzed by a family of adenosine deaminases acting on RNA (ADARs) that specifically recognize adenosines within double-stranded regions of pre-mRNAs and deaminate them to inosine. Inosines are recognised as guanosine by the cells translational machinery. There are three members of the ADAR family ADARs 1-3, with ADAR1 and ADAR2 being the only enzymatically active members. ADAR3 is thought to have a regulatory role in the brain. ADAR1 and ADAR2 are widely expressed in tissues, while ADAR3 is restricted to the brain. The double-stranded regions of RNA are formed by base-pairing between residues in the close to region of the editing site, with residues usually in a neighboring intron, but can be an exonic sequence. The region that base pairs with the editing region is known as an Editing Complementary Sequence (ECS). ADARs bind interact directly with the dsRNA substrate via their double-stranded RNA binding domains. If an editing site occurs within a coding sequence, it can result in a codon change. This can lead to translation of a protein isoform due to a change in its primary protein structure. Therefore, editing can also alter protein function. A-to-I editing occurs in a non coding RNA sequences such as introns, untranslated regions (UTRs), LINEs, SINEs (especially Alu repeats). The function of A to I editing in these regions is thought to involve creation of splice sites and retention of RNAs in the nucleus amongst others.
Location
In the pre-mRNA of GluR-2 the editing site Q/R is found at amino acid position 607. This location is in the pore loop region deep within the ion channel in the proteins membrane segment 2. Editing results in a change from a glutamine(Q) codon to an Arginine (R) codon. Editing at the R/G site, located at amino acid position 764 results in a codon change from arginine to glycine. All editing in glutamate receptors occurs in double-stranded RNAs (dsRNAs), which form due to complementary base pairing between the region of the editing site within the exon and an ECS within an intron sequence.[13] R/G site
Conservation
Regulation
Editing occurs at the Q/R site at a frequency of 100% of GluR2 transcripts in the brain. It is the only known editing site to be edited at a frequency of 100%.[11] However some striatal and cortical neurons are edited less frequently. This has been suggested as a reason for the higher level of excitotoxicity of these particular neurons.[14] The R/G site is developmentally regulated, being largely unedited in the embryonic brain with levels rising after birth. (ref 53)
Consequences
Structure
Editing results in a codon change from a glutamine codon (CAG) to an arginine codon (CIG).[15] Editing at R/G results in a codon change. The region of the editing site is known to be the region that controls divalent cation permeability. The other ionotropic AMPA glutamate receptors have a genomically encoded have a glutamine residue, while GluR2 has an arginine.
Function
RNA editing of the GluR-2 (GluR-B) pre-mRNA is the best-characterised example of A-to-I editing. Activated by L-Glutamate, a major excitatory neurotransmitter in vertebrates central nervous systems, it acts as an agonist at NMDA, AMPA, and kainate neurotransmitters.(103) Activation results in neuronal cation entry (CA2+), causing membrane depolarisation required for the process of excitatory neurotransmission. The calcium permeability of these receptor channels is required for many important events in the CNS, including long-term potentiation.(104) Since editing occurs in nearly 100% of transcripts and is necessary for life, it is often wondered why edited GluR-B is not genomically encoded instead of being derived by RNA editing. The answer is unknown.
RNA editing at the Q/R site is thought to alter the permeability of the channel rendering it impermeable to Ca2+. The Q/R site also occurs in the Kainate receptors GluR5 and GluR6. Editing at the Q/R site determines the calcium permeability of the channel,[11] with channels containing the edited form being less permeable to calcium. This differs from GluR6 where editing of the Q/R site may increase calcium permeability of the channel especially if the I/V and Y/C sites are also edited. Therefore, the main function of editing is therefore in regulation of electrophysiology of the channel.[16]
Editing in some striatal and cortical neurons is more likely to be subject to excitotoxicity, thought to be due to less than 100% editing of these particular neurons.[14] Editing also has several other function effects. Editing alters the maturation and assembly of the channel, with the unedited form having a tendency to tetramerize and then is transported to the synapse. However, the edited version is assembled as a monomer and resides mainly in the endoplasmic reticulum. The arginine residue in the pore loop of GluR-2 receptor is thought to belong to a retention signal for the endoplasmic reticulum. Therefore, editing - since it occurs at 100% frequency - inhibits the availability of the channel at the synapse. This process occurs before assembly of the channels, thereby preventing glur-2-forming homeric channels, which could interfere with synaptic signalling.
Editing also occurs at the R/G site. Editing at the R/G sites results in variation in the rate that the receptor recovers from desensitisation. Editing at these sites results in faster recovery time from desensitisation [17]
Dysregulation
Amyotrophic Lateral Sclerosis
Many human and animal studies have determined that RNA editing of the Q/R site in GluR2 pre-mRNA is necessary for normal brain function. Defective editing has been linked to several conditions such as
Epilepsy
In mouse models, failure of editing leads to epileptic seizures and death within 3 weeks of birth. [11] Why editing exists at this site instead of a genomically encoded arginine is unknown since nearly 100% of transcripts are edited.
Cancer
Decreased editing at the Q/R site is also found in some human brain tumors. Reduction of ADAR2 expression is thought to be associated with epileptic seizures in malignant glioma.[24]
Use in diagnostic immunochemistry
GRIA2 is a diagnostic immunochemical marker for
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000120251 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000033981 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- HGNC. "Symbol Report: GRIA2". Retrieved 29 December 2017.
- PMID 1311100.
- ^ a b "Entrez Gene: GRIA2 glutamate receptor, ionotropic, AMPA 2".
- PMID 22102833.
- S2CID 45794233.
- ^ PMID 11891216.
- ^ S2CID 11969068.
- S2CID 34081059.
- PMID 7937939.
- ^ S2CID 15351461.
- S2CID 22029384.
- PMID 7678465.
- S2CID 15936250.
- S2CID 2050462.
- S2CID 25556626.
- S2CID 2255590.
- S2CID 4310256.
- S2CID 5997020.
- ^ S2CID 6640926.
- PMID 11717408.
- S2CID 42812062.
Further reading
- Soundarapandian MM, Tu WH, Peng PL, et al. (2007). S2CID 21618951.
- McNamara JO, Eubanks JH, McPherson JD, et al. (1992). "Chromosomal localization of human glutamate receptor genes". J. Neurosci. 12 (7): 2555–62. PMID 1319477.
- Sommer B, Keinänen K, Verdoorn TA, et al. (1990). "Flip and flop: a cell-specific functional switch in glutamate-operated channels of the CNS". Science. 249 (4976): 1580–5. PMID 1699275.
- Sommer B, Köhler M, Sprengel R, Seeburg PH (1991). "RNA editing in brain controls a determinant of ion flow in glutamate-gated channels". Cell. 67 (1): 11–9. S2CID 22029384.
- Paschen W, Hedreen JC, Ross CA (1994). "RNA editing of the glutamate receptor subunits GluR2 and GluR6 in human brain tissue". J. Neurochem. 63 (5): 1596–602. S2CID 25226376.
- Köhler M, Kornau HC, Seeburg PH (1994). "The organization of the gene for the functionally dominant alpha-amino-3-hydroxy-5-methylisoxazole-4-propionic acid receptor subunit GluR-B". J. Biol. Chem. 269 (26): 17367–70. PMID 7545935.
- Eastwood SL, Burnet PW, Beckwith J, et al. (1994). "AMPA glutamate receptors and their flip and flop mRNAs in human hippocampus". NeuroReport. 5 (11): 1325–8. PMID 7919190.
- Sun W, Ferrer-Montiel AV, Montal M (1994). "Primary structure and functional expression of the AMPA/kainate receptor subunit 2 from human brain". NeuroReport. 5 (4): 441–4. PMID 8003671.
- Higuchi M, Single FN, Köhler M, et al. (1994). "RNA editing of AMPA receptor subunit GluR-B: a base-paired intron-exon structure determines position and efficiency". Cell. 75 (7): 1361–70. S2CID 25420811.
- McLaughlin DP, Cheetham ME, Kerwin RW (1993). "Expression of alternatively-spliced glutamate receptors in human hippocampus". Eur. J. Pharmacol. 244 (1): 89–92. PMID 8420792.
- Srivastava S, Osten P, Vilim FS, et al. (1998). "Novel anchorage of GluR2/3 to the postsynaptic density by the AMPA receptor-binding protein ABP". Neuron. 21 (3): 581–91. S2CID 14448034.
- Matsuda S, Mikawa S, Hirai H (1999). "Phosphorylation of serine-880 in GluR2 by protein kinase C prevents its C terminus from binding with glutamate receptor-interacting protein". J. Neurochem. 73 (4): 1765–8. S2CID 39402443.
- Hirai H, Matsuda S (2000). "Interaction of the C-terminal domain of delta glutamate receptor with spectrin in the dendritic spines of cultured Purkinje cells". Neurosci. Res. 34 (4): 281–7. S2CID 45794233.
- Aruscavage PJ, Bass BL (2000). "A phylogenetic analysis reveals an unusual sequence conservation within introns involved in RNA editing". RNA. 6 (2): 257–69. PMID 10688364.
- Osten P, Khatri L, Perez JL, et al. (2000). "Mutagenesis reveals a role for ABP/GRIP binding to GluR2 in synaptic surface accumulation of the AMPA receptor". Neuron. 27 (2): 313–25. S2CID 16213962.
- Chung HJ, Xia J, Scannevin RH, et al. (2001). "Phosphorylation of the AMPA receptor subunit GluR2 differentially regulates its interaction with PDZ domain-containing proteins". J. Neurosci. 20 (19): 7258–67. PMID 11007883.
- Armstrong N, Gouaux E (2000). "Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: crystal structures of the GluR2 ligand binding core". Neuron. 28 (1): 165–81. S2CID 3128719.
- Krampfl K, Schlesinger F, Zörner A, et al. (2002). "Control of kinetic properties of GluR2 flop AMPA-type channels: impact of R/G nuclear editing". Eur. J. Neurosci. 15 (1): 51–62. S2CID 35601416.
- Hirbec H, Perestenko O, Nishimune A, et al. (2002). "The PDZ proteins PICK1, GRIP, and syntenin bind multiple glutamate receptor subtypes. Analysis of PDZ binding motifs". J. Biol. Chem. 277 (18): 15221–4. PMID 11891216.
External links
- GRIA2+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- http://darned.ucc.ie
This article incorporates text from the United States National Library of Medicine, which is in the public domain.