Glutamine synthetase
| glutamate—ammonia ligase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Glutamine synthetase, beta-Grasp domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||
| Symbol | Gln-synt_N | ||||||||||
SCOP2 | 2gls / SCOPe / SUPFAM | ||||||||||
| |||||||||||
| Glutamine synthetase, catalytic domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||
| glutamate-ammonia ligase (glutamine synthetase) | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | GLUL | ||||||
| Alt. symbols | GLNS | ||||||
Chr. 1 q31 | |||||||
| |||||||
Glutamine synthetase (GS) (
:Glutamate + ATP + NH3 → Glutamine + ADP + phosphate

Glutamine synthetase uses ammonia produced by nitrate reduction,
Other reactions may take place via GS. Competition between ammonium ion and water, their binding affinities, and the concentration of ammonium ion, influences glutamine synthesis and glutamine hydrolysis. Glutamine is formed if an ammonium ion attacks the acyl-phosphate intermediate, while glutamate is remade if water attacks the intermediate.[6][7] Ammonium ion binds more strongly than water to GS due to electrostatic forces between a cation and a negatively charged pocket.[4] Another possible reaction is upon NH2OH binding to GS, rather than NH4+, yields γ-glutamylhydroxamate.[6][7]
Structure
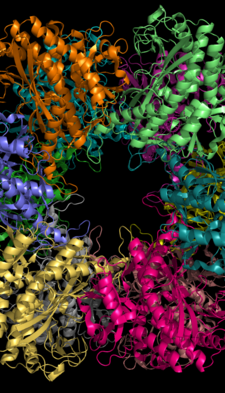
Glutamine synthetase can be composed of 8, 10, or 12 identical subunits separated into two face-to-face rings.
Mechanism
GS catalyzes the ATP-dependent condensation of glutamate with ammonia to yield glutamine.[4] The hydrolysis of ATP drives[8] the first step of a two-part, concerted mechanism.[4][6] ATP phosphorylates glutamate to form ADP and an acyl-phosphate intermediate, γ-glutamyl phosphate, which reacts with ammonia, forming glutamine and inorganic phosphate. ADP and Pi do not dissociate until ammonia binds and glutamine is released.[6]
ATP binds first to the top of the active site near a cation binding site, while glutamate binds near the second cation binding site at the bottom of the active site.[5][7] The presence of ADP causes a conformational shift in GS that stabilizes the γ-glutamyl phosphate moiety. Ammonium binds strongly to GS only if the acyl-phosphate intermediate is present. Ammonium, rather than ammonia, binds to GS because the binding site is polar and exposed to solvent.[7] In the second step, deprotonation of ammonium allows ammonia to attack the intermediate from its nearby site to form glutamine.[12] Phosphate leaves through the top of the active site, while glutamine leaves through the bottom (between two rings).[13][7]

Biological function
GS is present predominantly in the brain, kidneys, and liver.
Classes
There seem to be three different classes of GS:[18][19][20]
- Class I enzymes (GSI) are specific to prokaryotes, and are oligomers of 12 identical subunits.[21] The activity of GSI-type enzyme is controlled by the adenylation of a tyrosine residue. The adenylated enzyme is inactive.[22]
- Class II enzymes (GSII) are found in .
Plants have two or more isozymes of GSII, one of the isozymes is translocated into the chloroplast. Another form is cytosolic. The cytosolic GS gene translation is regulated by its 5' untranslated region (UTR), while its 3' UTR plays role in transcript turnover.[23]
- Class III enzymes (GSIII) have, currently, only been found in Butyrivibrio fibrisolvens. It is a double-ringed dodecamer of identical chains.[24]It is much larger (about 700 amino acids) than the GSI (450 to 470 amino acids) or GSII (350 to 420 amino acids) enzymes.
While the three classes of GSs are clearly structurally related, the sequence similarities are not so extensive.
Regulation and inhibition
GS is subject to reversible covalent modification. Tyr397 of all 12 subunits can undergo
Inhibition of GS has largely focused on amino site ligands.[6] Other inhibitors are the result of glutamine metabolism: tryptophan, histidine, carbamoyl phosphate, glucosamine-6-phosphate, cytidine triphosphate (CTP), and adenosine monophosphate (AMP).[5][8][27] Other inhibitors/regulators are glycine and alanine. Alanine, glycine, and serine bind to the glutamate substrate site. GDP, AMP, ADP bind to the ATP site.[6] L-serine, L-alanine, and glycine bind to the site for L-glutamate in unadenylated GS. The four amino acids bind to the site by their common atoms, “the main chain” of amino acids.[5] Glutamate is another product of glutamine metabolism; however, glutamate is a substrate for GS inhibiting it to act as a regulator to GS.2 Each inhibitor can reduce the activity of the enzyme; once all final glutamine metabolites are bound to GS, the activity of GS is almost completely inhibited.[8] Many inhibitory input signals allows for fine tuning of GS by reflecting nitrogen levels in the organism.
Feedback regulation distinguishes the difference between two eukaryotic types of GS: brain and non-brain tissues. Non-brain GS responds to end-product feedback inhibition, while brain GS does not.[6] High concentrations of glutamine-dependent metabolites should inhibit GS activity, while low concentrations should activate GS activity.[6]
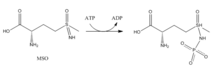
Inhibitors:
- Methionine sulfoximine (MSO): MSO is an inhibitor that binds to the glutamate site. Bound to GS, MSO is phosphorylated by ATP that results in an irreversible, non-covalent inhibition of GS. The S-isomer configuration is more inhibitory.[6] Glutamate entry is blocked into the active site by a stabilization of the flexible loop in the active site by MSO.[7]
- Many synthetic inhibitors are available today.[6]
Research on E. coli revealed that GS is regulated through gene expression. The gene that encodes the GS subunit is designated glnA. Transcription of glnA is dependent on NRI (a specific transcriptional enhancer). Active transcription occurs if NRI is in its phosphorylated form, designated NRI-P. Phosphorylation of NRI is catalyzed by NRII, a protein kinase. If NRII is complexed with PIIA then it will function as a phosphatase and NRI-P is converted back to NRI. In this case, transcription of glnA ceases.[25]
GS is subject to completely different regulatory mechanisms in
References
- ^ PMID 11329256.
- PMID 2572586.
- PMID 2900091.
- ^ PMID 8563633.
- ^ PMID 8099447.
- ^ PMID 10708854.
- ^ PMID 7904828.
- ^ ISBN 978-0-7167-8724-2.
- ^ Goodsell DS (June 2002). "Glutamine Synthetase". Molecule of the month. RCSB Protein Data Bank. Archived from the original on 2008-05-31. Retrieved 2010-05-08.
- ^ PMID 18005987.
- PMID 4906326.
- PMID 235885.
- ^ Goodsell, DS (June 2002). "Glutamine Synthetase". RCSB Protein Data Bank. Archived from the original on 31 May 2008. Retrieved 8 May 2010.
- ^ S2CID 24661063.
- PMID 24005729.
- ^ S2CID 31674240.
- PMID 15708485.
- PMID 8096645.
- S2CID 36704558.
- S2CID 21493521.
- ^ "GSI structure". Archived from the original on 2008-12-17. Retrieved 2009-03-31.
- ^ InterPro:IPR001637 Glutamine synthetase class-I, adenylation site
- PMID 23080263.
- PMID 16879836.
- ^ ISBN 978-1-305-57720-6.
- .
- PMID 2871863.
- PMID 30373240.
- PMID 8531888.
- ^ PMID 6115380.
- S2CID 32757978.
- PMID 9098067.
- PMID 10377385.
- S2CID 23804565.
- PMID 26494284.
- PMID 30085248.
External links
- InterPro entry
- RCSB PDB molecule of the month
- PDBe-KB provides an overview of all the structure information available in the PDB for human glutamine synthetase
