Glutathione reductase
| GSR | |||
|---|---|---|---|
Gene ontology | |||
| Molecular function | |||
| Cellular component | |||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 8: 30.68 – 30.73 Mb | Chr 8: 34.14 – 34.19 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Glutathione reductase (GR) also known as glutathione-disulfide reductase (GSR) is an enzyme that in humans is encoded by the GSR gene. Glutathione reductase (EC 1.8.1.7) catalyzes the reduction of glutathione disulfide (GSSG) to the sulfhydryl form glutathione (GSH), which is a critical molecule in resisting oxidative stress and maintaining the reducing environment of the cell.[5][6][7] Glutathione reductase functions as dimeric disulfide oxidoreductase and utilizes an FAD prosthetic group and NADPH to reduce one molar equivalent of GSSG to two molar equivalents of GSH:

The glutathione reductase is conserved between all
Function
| glutathione-disulfide reductase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Glutathione plays a key role in maintaining proper function and preventing oxidative stress in human cells. It can act as a scavenger for hydroxyl radicals, singlet oxygen, and various electrophiles. Reduced glutathione reduces the oxidized form of the enzyme glutathione peroxidase, which in turn reduces hydrogen peroxide (H2O2), a dangerously reactive species within the cell. [In the following illustration of redox reeactions, the rightmost arrow is reversed; it should be pointing up not down.] In addition, it plays a key role in the metabolism and clearance of xenobiotics, acts as a cofactor in certain detoxifying enzymes, participates in transport, and regenerates antioxidants such and Vitamins E and C to their reactive forms. The ratio of GSSG/GSH present in the cell is a key factor in properly maintaining the oxidative balance of the cell, that is, it is critical that the cell maintains high levels of the reduced glutathione and a low level of the oxidized glutathione disulfide. This narrow balance is maintained by glutathione reductase, which catalyzes the reduction of GSSG to GSH.[5]

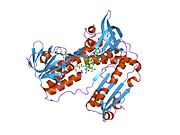
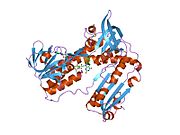
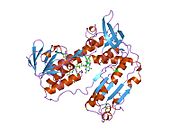
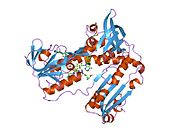
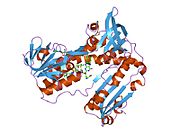
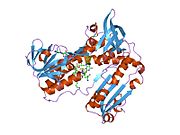
Structure
Glutathione reductase from human erythrocytes is a homodimer consisting of 52Kd monomers, each containing 3 domains. GR exhibits single sheet, double layered topology where an anti-parallel beta-sheet is largely exposed to the solvent on one face while being covered by random coils on the other face.[10] This includes and NADPH-binding Domain, FAD-binding domain(s) and a dimerization domain. Each monomer contains 478 residues and one FAD molecule. GR is a thermostable protein, retaining function up to 65 °C.[11][12]
Reaction mechanism


Steps:
| 1 | NADPH binding to the oxidized enzyme |
| 2 | Reduction of FAD to FADH− anion by NADPH |
| 3 | Reduced FADH− anion collapses into a charge relay complex and reduces Cys58-Cys63 disulfide |
| 4 | Oxidized Glutathione disulfide binds to the reduced enzyme and forms a mixed disulfide with Cys58 and releases one reduced glutathione |
| 5 | Cys63 attacks the mixed disulfide on Cys58 to release a reduced glutathione and reform the redox active disulfide |
Reductive half
The action of GR proceeds through two distinct half reactions, a reductive half mechanism followed by an oxidative half. In the first half, NADPH reduces FAD present in GSR to produce a transient FADH− anion. This anion then quickly breaks a disulfide bond of Cys58 - Cys63, forming a short lived covalent bond a stable charge-transfer complex between the flavin and Cys63. The now oxidized NADP+ is released and is subsequently replaced by a new molecule of NADPH. This is the end of the so-called reductive half of the mechanism.
Oxidative half
In the oxidative half of the mechanism, Cys63 nucleophilically attacks the nearest sulfide unit in the GSSG molecule (promoted by His467), which creates a mixed disulfide bond (GS-Cys58) and a GS− anion. His467 of GSR then protonates the GS- anion to release the first molecule of GSH. Next, Cys63 nucleophilically attacks the sulfide of Cys58, releasing a GS− anion, which, in turn, picks up a solvent
Inhibition
In vitro, glutathione reductase is inhibited by low concentrations of
Clinical significance
GSH is a key cellular antioxidant and plays a major role in the phase 2 metabolic clearance of electrophilic xenobiotics. The importance of the GSH pathway and enzymes that affect this delicate balance is gaining an increased level of attention in recent years. Although glutathione reductase has been an attractive target for many pharmaceuticals, there have been no successful glutathione reductase related therapeutic compounds created to date. In particular, glutathione reductase appears to be a good target for anti-malarials, as the glutathione reductase of the malaria parasite
There are two main classes of GR targeting compounds:[17][18][19][20]
- Inhibitors of GSSG binding, or dimerization: Reactive electrophilessuch as gold compounds, and fluoronaphthoquinones.
- Drugs which use glutathione reductase to regenerate, such as redox cyclers. Two examples of these types of compounds are Methylene blue and Naphthoquinone.
Clinical trials performed in
In cells exposed to high levels of
Lupus is an autoimmune disorder in which patients produce an elevated quantity of antibodies that attack DNA and other cell components. In a recent study, a single nucleotide polymorphism (SNP) in the Glutathione Reductase gene was found to be highly associated with lupus in African Americans in the study.[22] African Americans with lupus have also been shown to express less reduced glutathione in their T cells.[23] The study's authors believe that reduced glutathione reductase activity may contribute to the increased production of reactive oxygen in African Americans with lupus.[22]
In mice, glutathione reductase has been implicated in the oxidative burst, a component of the immune response.[24] The oxidative burst is a defense mechanism in which neutrophils produce and release reactive oxidative species in the vicinity of bacteria or fungi to destroy the foreign cells. Glutathione Reductase deficient neutrophils were shown to produce a more transient oxidative burst in response to bacteria than neutrophils that express GR at ordinary levels.[24] The mechanism of Glutathione Reductase in sustaining the oxidative burst is still unknown.[24]
Deficiency
Glutathione reductase deficiency is a rare disorder in which the glutathione reductase activity is absent from
Some patients exhibit deficient levels of glutathione activity as a result of not consuming enough riboflavin in their diets. Riboflavin is a precursor for FAD, whose reduced form donates two electron to the disulfide bond which is present in the oxidized form of glutathione reductase in order to begin the enzyme's catalytic cycle. In 1999, a study found that 17.8% of males and 22.4% of females examined in Saudi Arabia suffered from low glutathione reductase activity due to riboflavin deficiency.[27]
Connection to favism
In
Monitoring glutathione reductase activity
The activity of glutathione reductase is used as indicator for
In plants
As it does in human cells, glutathione reductase helps to protect plant cells from reactive oxygen species. In plants, reduced glutathione participates in the glutathione-ascorbate cycle in which reduced glutathione reduces dehydroascorbate, a reactive byproduct of the reduction of hydrogen peroxide. In particular, glutathione reductase contributes to plants' response to abiotic stress.[31] The enzyme's activity has been shown to be modulated in response to metals, metalloids, salinity, drought, UV radiation and heat induced stress.[31]
History
Glutathione reductase was first purified in 1955 at Yale University by P. Janmeda.[32] Janmeda also identified NADPH as the primary electron donor for the enzyme. Later groups confirmed the presence of FAD and the thiol group, and an initial mechanism was suggested for the mechanism in 1965.[33][34] The initial (low resolution) structure of glutathione reductase was solved in 1977. This was quickly followed by a 3Å structure by Shulze et al. in 1978.[35] Glutathione reductase has been studied exhaustively since these early experiments, and is subsequently one of the most well characterized enzymes to date.
Interactive pathway map
Interactive pathway can be found here: pathway map
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000104687 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031584 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ PMID 23036594.
- PMID 3053703.
- PMID 3315772.
- ^ PMID 11158675.
- PMID 18395526.
- ISBN 0534490336.)
{{cite book}}: CS1 maint: multiple names: authors list (link - PMID 16111877.
- PMID 11514662.
- PMID 18638483.
- PMID 15601678.
- PMID 1329770.
- PMID 12729762.
- PMID 17967916.
- PMID 21682307.
- PMID 15792952.
- PMID 23036594.
- ISBN 978-0-7817-6960-0.
- ^ PMID 23637325.
- PMID 11817589.
- ^ PMID 22279102.
- ^ PMID 17185460.
- PMID 435643.
- PMID 11924113.
- S2CID 29165746.
- PMID 3239770.
- PMID 19451637.
- ^ PMID 23792825.
- PMID 13271446.
- PMID 4378936.
- PMID 13932735.
- S2CID 4153363.
Further reading
- Sinet PM, Bresson JL, Couturier J, Laurent C, Prieur M, Rethoré MO, Taillemite JL, Toudic D, Jérome H, Lejeune J (1977). "[Possible localization of the glutathione reductase (EC 1.6.4.2) on the 8p21 band]". Ann. Genet. (in French). 20 (1): 13–7. PMID 302667.
- Krohne-Ehrich G, Schirmer RH, Untucht-Grau R (1978). "Glutathione reductase from human erythrocytes. Isolation of the enzyme and sequence analysis of the redox-active peptide". Eur. J. Biochem. 80 (1): 65–71. PMID 923580.
- Loos H, Roos D, Weening R, Houwerzijl J (1976). "Familial deficiency of glutathione reductase in human blood cells". Blood. 48 (1): 53–62. PMID 947404.
- Tutic M, Lu XA, Schirmer RH, Werner D (1990). "Cloning and sequencing of mammalian glutathione reductase cDNA". Eur. J. Biochem. 188 (3): 523–8. PMID 2185014.
- Palmer EJ, MacManus JP, Mutus B (1990). "Inhibition of glutathione reductase by oncomodulin". Arch. Biochem. Biophys. 277 (1): 149–54. PMID 2306116.
- Arnold HH, Heinze H (1990). "Treatment of human peripheral lymphocytes with concanavalin A activates expression of glutathione reductase". FEBS Lett. 267 (2): 189–92. S2CID 40084640.
- Karplus PA, Schulz GE (1987). "Refined structure of glutathione reductase at 1.54 A resolution". J. Mol. Biol. 195 (3): 701–29. PMID 3656429.
- Pai EF, Schulz GE (1983). "The catalytic mechanism of glutathione reductase as derived from x-ray diffraction analyses of reaction intermediates". J. Biol. Chem. 258 (3): 1752–7. PMID 6822532.
- Krauth-Siegel RL, Blatterspiel R, Saleh M, Schiltz E, Schirmer RH, Untucht-Grau R (1982). "Glutathione reductase from human erythrocytes. The sequences of the NADPH domain and of the interface domain". Eur. J. Biochem. 121 (2): 259–67. PMID 7060551.
- Thieme R, Pai EF, Schirmer RH, Schulz GE (1982). "Three-dimensional structure of glutathione reductase at 2 A resolution". J. Mol. Biol. 152 (4): 763–82. PMID 7334521.
- Huang J, Philbert MA (1995). "Distribution of glutathione and glutathione-related enzyme systems in mitochondria and cytosol of cultured cerebellar astrocytes and granule cells". Brain Res. 680 (1–2): 16–22. S2CID 39710661.
- Savvides SN, Karplus PA (1996). "Kinetics and crystallographic analysis of human glutathione reductase in complex with a xanthene inhibitor". J. Biol. Chem. 271 (14): 8101–7. PMID 8626496.
- Nordhoff A, Tziatzios C, van den Broek JA, Schott MK, Kalbitzer HR, Becker K, Schubert D, Schirmer RH (1997). "Denaturation and reactivation of dimeric human glutathione reductase--an assay for folding inhibitors". Eur. J. Biochem. 245 (2): 273–82. PMID 9151953.
- Stoll VS, Simpson SJ, Krauth-Siegel RL, Walsh CT, Pai E (1997). "Glutathione reductase turned into trypanothione reductase: structural analysis of an engineered change in substrate specificity". Biochemistry. 36 (21): 6437–47. PMID 9174360.
- Becker K, Savvides SN, Keese M, Schirmer RH, Karplus PA (1998). "Enzyme inactivation through sulfhydryl oxidation by physiologic NO-carriers". Nat. Struct. Biol. 5 (4): 267–71. S2CID 20607289.
- Kelner MJ, Montoya MA (2000). "Structural organization of the human glutathione reductase gene: determination of correct cDNA sequence and identification of a mitochondrial leader sequence". Biochem. Biophys. Res. Commun. 269 (2): 366–8. PMID 10708558.
- Qanungo S, Mukherjea M (2001). "Ontogenic profile of some antioxidants and lipid peroxidation in human placental and fetal tissues". Mol. Cell. Biochem. 215 (1–2): 11–9. S2CID 22048227.
- Berry Y, Truscott RJ (2001). "The presence of a human UV filter within the lens represents an oxidative stress". Exp. Eye Res. 72 (4): 411–21. PMID 11273669.
- Rhie G, Shin MH, Seo JY, Choi WW, Cho KH, Kim KH, Park KC, Eun HC, Chung JH (2001). "Aging- and photoaging-dependent changes of enzymic and nonenzymic antioxidants in the epidermis and dermis of human skin in vivo". J. Invest. Dermatol. 117 (5): 1212–7. PMID 11710935.
- Zatorska A, Józwiak Z (2003). "Involvement of glutathione and glutathione-related enzymes in the protection of normal and trisomic human fibroblasts against daunorubicin". Cell Biol. Int. 26 (5): 383–91. S2CID 31321422.