Hepatitis C virus
| Hepacivirus C | |
|---|---|
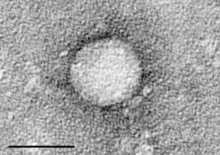
| |
Electron micrograph of Hepacivirus C purified from cell culture. Scale bar = 50 nanometres
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Kitrinoviricota |
| Class: | Flasuviricetes |
| Order: | Amarillovirales |
| Family: | Flaviviridae |
| Genus: | Hepacivirus |
| Species: | Hepacivirus C
|
| Synonyms[1][2] | |
|
Hepatitis C virus | |
The hepatitis C virus (HCV)[3] is a small (55–65 nm in size), enveloped, positive-sense single-stranded RNA virus of the family Flaviviridae. The hepatitis C virus is the cause of hepatitis C and some cancers such as liver cancer (hepatocellular carcinoma, abbreviated HCC) and lymphomas in humans.[4][5]
Taxonomy
The hepatitis C virus belongs to the genus
Structure

The hepatitis C virus particle consists of a
E1 and E2 glycoproteins
E1 and E2 are
These glycoproteins play an important role in the interactions hepatitis C has with the immune system. A hypervariable region, the hypervariable region 1 (HVR1) can be found on the E2 glycoprotein.[10] HVR1 is flexible and quite accessible to surrounding molecules.[13] HVR1 helps E2 shield the virus from the immune system. It prevents CD81 from latching onto its respective receptor on the virus.[13] In addition, E2 can shield E1 from the immune system.[13] Although HVR1 is quite variable in amino acid sequence, this region has similar chemical, physical, and conformational characteristics across many E2 glycoproteins.[14]
Genome

Hepatitis C virus has a
At the 5′ and 3′ ends of the RNA are the
Molecular biology
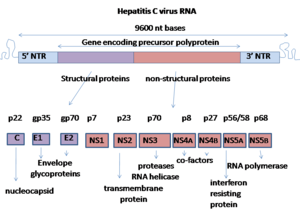
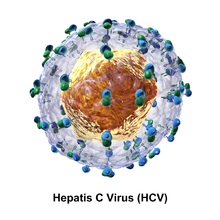
This section needs additional citations for verification. (May 2016) |
The proteins of this virus are arranged along the genome in the following order: N terminal-core-envelope (E1)–E2–p7-nonstructural protein 2 (NS2)–NS3–NS4A–NS4B–NS5A–NS5B–C terminal. The mature nonstructural proteins (NS2 to NS5B) generation relies on the activity of viral proteinases.[19] The NS2/NS3 junction is cleaved by a metal-dependent autocatalytic proteinase encoded within NS2 and the N-terminus of NS3. The remaining cleavages downstream from this site are catalysed by a serine protease also contained within the N-terminal region of NS3.
- The core protein has 191 amino acids and can be divided into three domains on the basis of hydrophobicity: domain 1 (residues 1–117) contains mainly basic residues with two short hydrophobic regions; domain 2 (residues 118–174) is less basic and more hydrophobic and its C-terminus is at the end of p21; domain 3 (residues 175–191) is highly hydrophobic and acts as a signal sequence for E1 envelope protein.
- Both envelope proteins (E1 and E2) are highly glycosylated and important in cell entry. E1 serves as the fusogenic subunit and E2 acts as the receptor binding protein. E1 has 4–5 N-linked glycans and E2 has 11 N-glycosylation sites.
- NS1 (p7) protein is dispensable for viral genome replication but plays a critical role in virus morphogenesis. This protein is a 63 amino acid membrane-spanning protein which locates itself in the endoplasmic reticulum. Cleavage of p7 is mediated by the endoplasmic reticulum's signal peptidases. Two transmembrane domains of p7 are connected by a cytoplasmic loop and are oriented towards the endoplasmic reticulum's lumen.
- NS2 protein is a 21–23 kilodalton(kDa) transmembrane protein with protease activity.
- NS3 is 67 kDa protein whose N-terminal has serine protease activity and whose C-terminal has of the proteinase.
- NS4A—a 54 amino acid membrane protein that acts as a cofactor of the proteinase.
- NS4B is a small (27 kDa) hydrophobic integral membrane protein with four transmembrane domains. It is located within the endoplasmic reticulum and plays an important role for recruitment of other viral proteins. It induces morphological changes to the endoplasmic reticulum forming a structure termed the membranous web.
- NS5A is a hydrophilic phosphoprotein which plays an important role in viral replication, modulation of cell signaling pathways and the interferon response. It is known to bind to endoplasmic reticulum-anchored human VAP proteins.[20]
- The NS5B protein (65 kDa) is the viral RNA-dependent RNA replication.[21][22][23] Several crystal structures of NS5B polymerase in several crystalline forms have been determined based on the same consensus sequence BK (HCV-BK, genotype 1).[24] The structure can be represented by a right hand shape with fingers, palm, and thumb. The encircled active site, unique to NS5B, is contained within the palm structure of the protein. Recent studies on NS5B protein genotype 1b strain J4's (HC-J4) structure indicate a presence of an active site where possible control of nucleotide binding occurs and initiation of de-novo RNA synthesis. De-novo adds necessary primers for initiation of RNA replication.[25] Current research attempts to bind structures to this active site to alter its functionality in order to prevent further viral RNA replication.[26]
An 11th protein has also been described.
Replication

Replication of HCV involves several steps. The virus replicates mainly in the
HCV has a wide variety of genotypes and mutates rapidly due to a high error rate on the part of the virus' RNA-dependent RNA polymerase. The mutation rate produces so many variants of the virus it is considered a quasispecies rather than a conventional virus species.[29] Entry into host cells occur through complex interactions between virions, especially through their glycoproteins, and cell-surface molecules CD81, LDL receptor, SR-BI, DC-SIGN, Claudin-1, and Occludin.[30][31]
The envelope of HCV is similar to
These interactions lead to the endocytosis of the viral particle. This process is aided by clathrin proteins. Once inside an early endosome, the endosome and the viral envelope fuse and the RNA is allowed into the cytoplasm.[10]
HCV takes over portions of the intracellular machinery to replicate.[32] The HCV genome is translated to produce a single protein of around 3,011 amino acids. The polyprotein is then proteolytically processed by viral and cellular proteases to produce three structural (virion-associated) and seven nonstructural (NS) proteins. Alternatively, a frameshift may occur in the Core region to produce an alternate reading frame protein (ARFP).[33] HCV encodes two proteases, the NS2 cysteine autoprotease and the NS3-4A serine protease. The NS proteins then recruit the viral genome into an RNA replication complex, which is associated with rearranged cytoplasmic membranes. RNA replication takes place via the viral RNA-dependent RNA polymerase NS5B, which produces a negative strand RNA intermediate. The negative strand RNA then serves as a template for the production of new positive strand viral genomes. Nascent genomes can then be translated, further replicated or packaged within new virus particles.[citation needed]
The virus replicates on intracellular lipid membranes.
Genotypes
Based on genetic differences between HCV isolates, the hepatitis C virus species is classified into six
Clinical importance
Parts of this article (those related to direct-acting antiviral medications) need to be updated. (January 2024) |
Genotype is clinically important in determining potential response to
Infection with one genotype does not confer immunity against others, and concurrent infection with two strains is possible. In most of these cases, one of the strains outcompetes the other in a short time. This finding may be useful in treatment, in replacing strains non-responsive to medication with others easier to treat.[46]
Recombination
When two viruses infect the same cell, genetic recombination may occur.[47] Although infrequent, HCV recombination has been observed between different genotypes, between subtypes of the same genotype and even between strains of the same subtype.[47]
Epidemiology
Hepatitis C virus is predominantly a
Evolution
Identification of the origin of this virus has been difficult but genotypes 1 and 4 appear to share a common origin.[51] A
A study of genotype 6 strains suggests an earlier date of evolution: approximately 1,100 to 1,350 years Before Present.[53] The estimated rate of mutation was 1.8 × 10−4. An experimental study estimated the mutation rate at 2.5–2.9 × 10−3 base substitutions per site per year.[54] This genotype may be the ancestor of the other genotypes.[53]
A study of European, US and Japanese isolates suggested that the date of origin of genotype 1b was approximately in the year 1925.[55] The estimated dates of origin of types 2a and 3a were 1917 and 1943 respectively. The time of divergence of types 1a and 1b was estimated to be 200–300 years.[citation needed]
A study of genotype 1a and 1b estimated the dates of origin to be 1914–1930 for type 1a and 1911–1944 for type 1b.[56] Both types 1a and 1b underwent massive expansions in their effective population size between 1940 and 1960. The expansion of HCV subtype 1b preceded that of subtype 1a by at least 16 years. Both types appear to have spread from the developed world to the developing world.
The genotype 2 strains from Africa can be divided into four clades that correlate with their country of origin: (1) Cameroon and Central African Republic (2) Benin, Ghana and Burkina Faso (3) Gambia, Guinea, Guinea-Bissau and Senegal (4) Madagascar.[57] There is also strong evidence for the dissemination of HCV genotype 2 from West Africa to the Caribbean by the
Genotype 3 is thought to have its origin in South East Asia.[59]
These dates from these various countries suggests that this virus may have evolved in South East Asia and was spread to West Africa by traders from Western Europe.[60] It was later introduced into Japan once that country's self-imposed isolation was lifted. Once introduced to a country its spread has been influenced by many local factors including blood transfusions, vaccination programmes, intravenous drug use and treatment regimes. Given the reduction in the rate of spread once screening for HCV in blood products was implemented in the 1990s, it would seem that previously blood transfusion was an important method of spread. Additional work is required to determine the dates of evolution of the various genotypes and the timing of their spread across the globe.[citation needed]
Vaccination
Unlike hepatitis A and B, there is currently no vaccine to prevent hepatitis C infection.[61]
Current research
This article's factual accuracy may be compromised due to out-of-date information. The reason given is: HCV is now curable as a result of medical research: https://web.archive.org/web/20180919174730/http://www.who.int/hepatitis/news-events/direct-acting-antiviral-cure-hepatitis-c/en/. (November 2019) |
The study of HCV has been hampered by the narrow host range of HCV.[62] The use of replicons has been successful but these have only been recently discovered.[63] HCV, as with most RNA viruses, exists as a viral quasispecies, making it very difficult to isolate a single strain or receptor type for study.[64][65]
Current research is focused on small-molecule inhibitors of the viral
A possible association between low Vitamin D levels and a poor response to treatment has been reported.[67][68][69][70] In vitro work has shown that vitamin D may be able to reduce viral replication.[71] While this work looks promising[72][73] the results of clinical trials are pending.[74][75] However, it has been proposed that vitamin D supplementation is important in addition to standard treatment, in order to enhance treatment response.[76]
Naringenin, a flavonoid found in grapefruit and other fruits and herbs, has been shown to block the assembly of intracellular infectious viral particles without affecting intracellular levels of the viral RNA or protein.[76]
Other agents that are under investigation include
Sofosbuvir for use against chronic hepatitis C infection was approved by the FDA on December 6, 2013. It has been reported to be the first drug that has demonstrated safety and efficacy to treat certain types of HCV infection without the need for co-administration of interferon.[78] On November 22, the FDA approved simeprevir for use in combination with peginterferon-alfa and ribavirin.[79] Simeprevir has been approved in Japan for the treatment of chronic hepatitis C infection, genotype 1.[80]
There is also current experimental research on non drug related therapies. Oxymatrine, for example, is a root extract found in the continent of Asia that has been reported to have antiviral activity against HCV in cell cultures and animal studies. Small and promising human trials have shown beneficial results and no serious side effects, but they were too small to generalize conclusions.[76]
On October 5, 2020, it was announced that
See also
- Blood-borne disease
- Cancer virus
- Discovery and development of NS5A inhibitors
- HCV IRES
- Hepatitis C virus stem-loop VII
- Hepatitis C virus 3′X element
- Hepatitis C virus (HCV) cis-acting replication element (CRE)
- Sexually transmitted infection
References
- ^ "Hepacivirus C". National Center for Biotechnology Information (NCBI). Retrieved 3 January 2022.
- ^ Smith, Donald B.; et al. (23 June 2016). "Create 13 new species in the genus Hepacivirusand rename 1 species (family Flaviviridae)" (PDF). International Committee on Taxonomy of Viruses (ICTV). Retrieved 13 March 2019.
- ^ "Genus: Hepacivirus". International Committee on Taxonomy of Viruses. July 2018. Archived from the original on February 19, 2020.
- PMID 25848462.
- PMID 23871966.
- PMID 21610165.
- PMID 22491452.
- PMID 23610427.
- PMID 23572554.
- ^ PMID 25443344.
- ^ S2CID 24668769.
- S2CID 22280227.
- ^ PMID 28303031.
- PMID 15078928.
- PMID 11252351.
- PMID 11497352.
- PMID 22000514.
- PMID 17552023.
- PMID 10622560.
- PMID 22720086.
- PMID 22303022.
- S2CID 13176201.
- PMID 20813137.
- PMID 15746101.
- PMID 12589751.
- PMID 16828488.
- PMID 11350035.
- S2CID 7379944.
- PMID 10859368. Archived from the original on 2009-12-03. Retrieved 2010-07-16.)
{{cite journal}}: CS1 maint: DOI inactive as of February 2024 (link - PMID 19273272.
- PMID 20463075.
- PMID 16107832.
- S2CID 260316882.
- PMID 12446113.
- PMID 12021330.
- S2CID 20609887.
- PMID 19854061.
- S2CID 17597460. Retrieved 10 July 2020.
- S2CID 23271017.
- PMID 8968908.
- S2CID 21393716.
- S2CID 27333980.
- S2CID 1707096.
- PMID 24140473.
- PMID 11160710.
- ^ PMID 22069526.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - ^ PMID 16122679.
- S2CID 260310609.
- PMID 17521655.
- S2CID 35899454.
- PMID 21696641.
- ^ PMID 18971279.
- ^ Kato N, Ueda Y, Sejima H, Gu W, Satoh S, Dansako H, Ikeda M, Shimotohno K (2019) Study of multiple genetic variations caused by persistent hepatitis C virus replication in long-term cell culture. Arch Virol
- S2CID 41594303.
- PMID 20041120.
- PMID 19474244.
- PMID 22573865.
- PMID 15483230.
- PMID 11516379.
- PMID 20625493.
- PMID 19891557.
- S2CID 38411966.
- S2CID 52874660.
- S2CID 3829214.
- ^ "FDA approves Victrelis for Hepatitis C" (press release). FDA. May 13, 2011.
- PMID 22189978.
- PMID 21145801. Retrieved 10 July 2020.
- S2CID 32175340.
- S2CID 5329252.
- S2CID 10090454.
- PMID 22215943.
- S2CID 22124427.
- S2CID 25534747.
- PMID 22201953.
- ^ PMID 24653790.
- PMID 22300469.
- ^ "Press announcement, FDA, December 6 2013". Food and Drug Administration.
- ^ "FDA approves new treatment for hepatitis C virus". Food and Drug Administration. Nov 22, 2013.
- ^ "Medivir: Simeprevir has been approved in Japan for the treatment of genotype 1 chronic hepatitis C infection". The Wall Street Journal. September 27, 2013.
- ^ "The Nobel Prize in Physiology or Medicine 2020". NobelPrize.org.
External links
- Academic articles about the HCV six genotypes Clodovero Ferri
- HCV Sequence and Immunology Databases at Los Alamos National Laboratory
- Virus Pathogen Database and Analysis Resource (ViPR): Flaviviridae
