Heterochromatin
Heterochromatin is a tightly packed form of
Constitutive heterochromatin can affect the genes near itself (e.g.
Facultative heterochromatin is the result of genes that are silenced through a mechanism such as histone deacetylation or Piwi-interacting RNA (piRNA) through RNAi. It is not repetitive and shares the compact structure of constitutive heterochromatin. However, under specific developmental or environmental signaling cues, it can lose its condensed structure and become transcriptionally active.[4]
Heterochromatin has been associated with the di- and tri -methylation of H3K9 in certain portions of the human genome.[5] H3K9me3-related methyltransferases appear to have a pivotal role in modifying heterochromatin during lineage commitment at the onset of organogenesis and in maintaining lineage fidelity.[6]
Structure

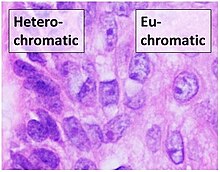
Chromatin is found in two varieties: euchromatin and heterochromatin.[7] Originally, the two forms were distinguished cytologically by how intensely they get stained – the euchromatin is less intense, while heterochromatin stains intensely, indicating tighter packing. Heterochromatin is usually localized to the periphery of the nucleus.
Despite this early dichotomy, recent evidence in both animals
Heterochromatin mainly consists of genetically inactive satellite sequences,[10] and many genes are repressed to various extents, although some cannot be expressed in euchromatin at all.[11] Both centromeres and telomeres are heterochromatic, as is the Barr body of the second, inactivated X-chromosome in a female.
Function

Heterochromatin has been associated with several functions, from gene regulation to the protection of chromosome integrity;[12] some of these roles can be attributed to the dense packing of DNA, which makes it less accessible to protein factors that usually bind DNA or its associated factors. For example, naked double-stranded DNA ends would usually be interpreted by the cell as damaged or viral DNA, triggering cell cycle arrest, DNA repair or destruction of the fragment, such as by endonucleases in bacteria.
Some regions of chromatin are very densely packed with fibers that display a condition comparable to that of the chromosome at
Constitutive heterochromatin
All cells of a given species package the same regions of DNA in
Facultative heterochromatin

The regions of DNA packaged in facultative heterochromatin will not be consistent between the cell types within a species, and thus a sequence in one cell that is packaged in facultative heterochromatin (and the genes within are poorly expressed) may be packaged in euchromatin in another cell (and the genes within are no longer silenced). However, the formation of facultative heterochromatin is regulated, and is often associated with morphogenesis or differentiation. An example of facultative heterochromatin is X chromosome inactivation in female mammals: one X chromosome is packaged as facultative heterochromatin and silenced, while the other X chromosome is packaged as euchromatin and expressed.
Among the molecular components that appear to regulate the spreading of heterochromatin are the
Yeast heterochromatin
Saccharomyces cerevisiae, or budding yeast, is a model eukaryote and its heterochromatin has been defined thoroughly. Although most of its genome can be characterized as euchromatin, S. cerevisiae has regions of DNA that are transcribed very poorly. These loci are the so-called silent mating type loci (HML and HMR), the rDNA (encoding ribosomal RNA), and the sub-telomeric regions.
Fission yeast (
In the fission yeast Schizosaccharomyces pombe, two RNAi complexes, the RITS complex and the RNA-directed RNA polymerase complex (RDRC), are part of an RNAi machinery involved in the initiation, propagation and maintenance of heterochromatin assembly. These two complexes localize in a
See also
References
- S2CID 2613813.
- ^ "What is the current evidence showing active transcription withinin..." www.researchgate.net. Retrieved 2016-04-30.
- PMID 28751582.
- S2CID 15674132.
- PMID 19335899.
- PMID 30606806.
- ^
Elgin, S.C. (1996). "Heterochromatin and gene regulation in Drosophila". PMID 8722176.
- ^
van Steensel B (May 2011). "Chromatin: constructing the big picture". The EMBO Journal. 30 (10): 1885–95. PMID 21527910.
- ^
Roudier F, Ahmed I, Bérard C, Sarazin A, Mary-Huard T, Cortijo S, et al. (May 2011). "Integrative epigenomic mapping defines four main chromatin states in Arabidopsis". The EMBO Journal. 30 (10): 1928–38. PMID 21487388.
- ^
Lohe AR, Hilliker AJ, Roberts PA (August 1993). "Mapping simple repeated DNA sequences in heterochromatin of Drosophila melanogaster". Genetics. 134 (4): 1149–74. PMID 8375654.
- ^
Lu BY, Emtage PC, Duyf BJ, Hilliker AJ, Eissenberg JC (June 2000). "Heterochromatin protein 1 is required for the normal expression of two heterochromatin genes in Drosophila". Genetics. 155 (2): 699–708. PMID 10835392.
- ^
Grewal SI, Jia S (January 2007). "Heterochromatin revisited". Nature Reviews. Genetics. 8 (1): 35–46. S2CID 31811880.
An up-to-date account of the current understanding of repetitive DNA, which usually doesn't contain genetic information. If evolution makes sense only in the context of the regulatory control of genes, we propose that heterochromatin, which is the main form of chromatin in higher eukaryotes, is positioned to be a deeply effective target for evolutionary change. Future investigations into assembly, maintenance and the many other functions of heterochromatin will shed light on the processes of gene and chromosome regulation.
- ^
Fisher AG, Merkenschlager M (April 2002). "Gene silencing, cell fate and nuclear organisation". Current Opinion in Genetics & Development. 12 (2): 193–7. PMID 11893493.
- ^
S2CID 24439936.
- ^
Burgess-Beusse B, Farrell C, Gaszner M, Litt M, Mutskov V, Recillas-Targa F, et al. (December 2002). "The insulation of genes from external enhancers and silencing chromatin". Proceedings of the National Academy of Sciences of the United States of America. 99 (Suppl 4): 16433–7. PMID 12154228.
- ^
Allis CD, Grewal SI (August 2001). "Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries". Science. 293 (5532): 1150–5. S2CID 26350729.
- ^
Donze D, Kamakaka RT (February 2001). "RNA polymerase III and RNA polymerase II promoter complexes are heterochromatin barriers in Saccharomyces cerevisiae". The EMBO Journal. 20 (3): 520–31. PMID 11157758.
- S2CID 1671107.
- PMID 28758948.
- S2CID 22636283.
- PMID 16204182.
- ISBN 978-1-904455-25-7.
- PMID 19942857.
External links
- Histology image: 20102loa – Histology Learning System at Boston University
- Avramova ZV (May 2002). "Heterochromatin in animals and plants. Similarities and differences". Plant Physiology. 129 (1): 40–9. PMID 12011336.
- Caron H, van Schaik B, van der Mee M, Baas F, Riggins G, van Sluis P, et al. (February 2001). "The human transcriptome map: clustering of highly expressed genes in chromosomal domains". Science. 291 (5507): 1289–92. PMID 11181992.
- Cha, Ariana Eunjung; Bernstein, Lenny (April 30, 2015). "Scientists discover an important new driver of aging". New York Times. Retrieved 4 May 2015.
