Histidine
 Skeletal formula of histidine (zwitterionic form)
| |||
| |||
| Names | |||
|---|---|---|---|
| IUPAC name
Histidine
| |||
| Other names
2-Amino-3-(1H-imidazol-4-yl)propanoic acid
| |||
| Identifiers | |||
3D model (
JSmol ) |
| ||
| 84088 | |||
| ChEBI | |||
| ChEMBL | |||
| ChemSpider | |||
| DrugBank | |||
ECHA InfoCard
|
100.000.678 | ||
| EC Number |
| ||
| 83042 | |||
IUPHAR/BPS |
|||
| KEGG | |||
PubChem CID
|
|||
| UNII | |||
CompTox Dashboard (EPA)
|
|||
| |||
| |||
| Properties | |||
| C6H9N3O2 | |||
| Molar mass | 155.157 g·mol−1 | ||
| 4.19g/100g @ 25 °C [1] | |||
| Hazards | |||
| NFPA 704 (fire diamond) | |||
| Supplementary data page | |||
| Histidine (data page) | |||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |||
Histidine (symbol His or H)
Histidine was first isolated by Albrecht Kossel and Sven Gustaf Hedin in 1896.[4] The name stems from its discovery in tissue, from ἱστός histós "tissue".[2] It is also a precursor to histamine, a vital inflammatory agent in immune responses. The acyl radical is histidyl.
Properties of the imidazole side chain
The conjugate acid (protonated form) of the
The acid-base properties of the imidazole side chain are relevant to the
The tautomerism and acid-base properties of the imidazole side chain has been characterized by 15N NMR spectroscopy. The two 15N chemical shifts are similar (about 200 ppm, relative to
Ligand

Histidine forms
Metabolism
Biosynthesis
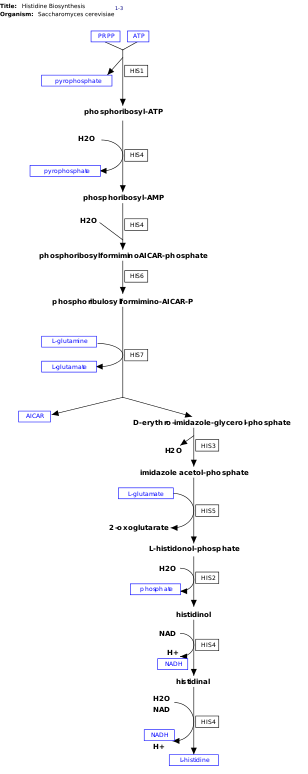
l-Histidine is an essential amino acid that is not synthesized de novo in humans.[14] Humans and other animals must ingest histidine or histidine-containing proteins. The biosynthesis of histidine has been widely studied in prokaryotes such as E. coli. Histidine synthesis in E. coli involves eight gene products (His1, 2, 3, 4, 5, 6, 7, and 8) and it occurs in ten steps. This is possible because a single gene product has the ability to catalyze more than one reaction. For example, as shown in the pathway, His4 catalyzes 4 different steps in the pathway.[15]
Histidine is synthesized from
The histidine biosynthesis pathway has been studied in the fungus Neurospora crassa, and a gene (His-3) encoding a multienzyme complex was found that was similar to the His4 gene of the bacterium E. coli.[18] A genetic study of N. crassa histidine mutants indicated that the individual activities of the multienzyme complex occur in discrete, contiguous sections of the His-3 genetic map, suggesting that the different activities of the multienzyme complex are encoded separately from each other.[18] However, mutants were also found that lacked all three activities simultaneously, suggesting that some mutations cause loss of function of the complex as a whole.
Just like animals and microorganisms, plants need histidine for their growth and development.[10] Microorganisms and plants are similar in that they can synthesize histidine.[19] Both synthesize histidine from the biochemical intermediate phosphoribosyl pyrophosphate. In general, the histidine biosynthesis is very similar in plants and microorganisms.[20]
Regulation of biosynthesis
This pathway requires energy in order to occur therefore, the presence of ATP activates the first enzyme of the pathway, ATP-phosphoribosyl transferase (shown as His1 in the image on the right). ATP-phosphoribosyl transferase is the rate determining enzyme, which is regulated through feedback inhibition meaning that it is inhibited in the presence of the product, histidine.[21]
Degradation
Histidine is one of the amino acids that can be converted to intermediates of the tricarboxylic acid (TCA) cycle (also known as the citric acid cycle).
Conversion to other biologically active amines
- The histidine amino acid is a precursor for histamine, an amine produced in the body necessary for inflammation.[25]
- The enzyme histidine ammonia-lyase converts histidine into ammonia and urocanic acid. A deficiency in this enzyme is present in the rare metabolic disorder histidinemia, producing urocanic aciduria as a key diagnostic finding.
- Histidine can be converted to 3-methylhistidine, which serves as a biomarker for skeletal muscle damage, by certain methyltransferase enzymes.[26]
- Histidine is also a precursor for carnosine biosynthesis, which is a dipeptide found in skeletal muscle.[27]
- In Actinomycetota and filamentous fungi, such as Neurospora crassa, histidine can be converted into the antioxidant ergothioneine.[28]

Requirements
The
See also
- Carnosinemia
- Beta-Alanine
- Diphthamide
- Pauly reaction
References
- ^ http://prowl.rockefeller.edu/aainfo/solub.htm[full citation needed]
- ^ a b "Nomenclature and Symbolism for Amino Acids and Peptides". IUPAC-IUB Joint Commission on Biochemical Nomenclature. 1983. Archived from the original on 9 October 2008. Retrieved 5 March 2018.
- PMID 1123426.
- ISSN 0021-9258.
- ^ PMID 23579613.
- ^ a b "HISTIDINE". ambermd.org. Retrieved 2022-05-12.
- ^ S2CID 88813605.
- .
- S2CID 265572280.
- ^ PMID 22303266.
- ISBN 978-1-891389-18-4.
- PMID 11036646.
- PMID 26214303.
- ^ Roche Biochemical Pathways Map Roche biochemical pathways map
- ^ PMID 8852895.
- ^ PMID 23617600.
- PMID 13271397.
- ^ a b Ahmed A. Organization of the histidine-3 region of Neurospora. Mol Gen Genet. 1968;103(2):185-93. doi: 10.1007/BF00427145. PMID 4306011
- ^ "Understanding Genetics". genetics.thetech.org. Archived from the original on 2016-05-25. Retrieved 2016-05-19.
- S2CID 23733445.
- S2CID 18380727.
- ^ a b c Board review series (BRS)-- Biochemistry, Molecular Biology, and Genetics (fifth edition): Swanson, Kim, Glucksman
- PMID 4146796.
- PMID 13061415.
- PMID 26015312.
- ^ "3-Methylhistidine". HMDB Version 4.0. Human Metabolome Database. 20 December 2017. Retrieved 25 December 2017.
- S2CID 7661250.
- PMID 11544359.
- ISBN 978-0-309-08525-0.
- PMID 32235743.