Human coronavirus OC43
| Human coronavirus OC43 | |
|---|---|
| |
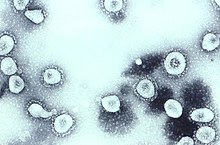
Transmission electron micrograph of human coronavirus OC43
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Genus: | Betacoronavirus |
| Subgenus: | Embecovirus |
| Species: | |
| Virus: | Human coronavirus OC43
|
Human coronavirus OC43[1] (HCoV-OC43) is a member of the species Betacoronavirus 1, which infects humans and cattle.[2][3] The infecting coronavirus is an enveloped, positive-sense, single-stranded RNA virus that enters its host cell by binding to the N-acetyl-9-O-acetylneuraminic acid receptor.[4] OC43 is one of seven coronaviruses known to infect humans. It is one of the viruses responsible for the common cold[5][6] and may have been responsible for the 1889–1890 pandemic.[7] It has, like other coronaviruses from genus Betacoronavirus, subgenus Embecovirus, an additional shorter spike protein called hemagglutinin-esterase (HE).[8][2]
Virology
Four HCoV-OC43
Comparison of HCoV-OC43 with the most closely related strain of Betacoronavirus 1 species,
The origin of HCoV-OC43 is uncertain, but it is thought that it may have originated in rodents, then passed through cattle as intermediate hosts.[12] A deletion from BCoV to HCoV-OC43 may have taken place for the interspecies transmission event from bovines to humans.[9]
Pathogenesis
Along with
If HCoV-OC43 was indeed the pathogen responsible for the 1889–1890 pandemic, which resembled the COVID-19 pandemic, severe disease was much more common and mortality much higher in populations that had not previously been exposed.[16]
Epidemiology
Coronaviruses have a worldwide distribution, causing 10–15% of common cold cases (the virus most commonly implicated in the common cold is a rhinovirus, found in 30–50% of cases).[17] Infections show a seasonal pattern with most cases occurring in the winter months in temperate climates, and summer and spring in warm climates.[18][17][19][20]
See also
References
- hdl:10722/131538.)
{{cite thesis}}: CS1 maint: DOI inactive as of April 2024 (link - ^ a b "Taxonomy browser (Betacoronavirus 1)". www.ncbi.nlm.nih.gov. Retrieved 2020-02-29.
- PMID 28933406.
See Table 1.
- PMID 27578435.
BCoV S1-NTD does not recognize galactose as galectins do. Instead, it recognizes 5-N-acetyl-9-O-acetylneuraminic acid (Neu5,9Ac2) (30, 43). The same sugar receptor is also recognized by human coronavirus OC43 (43, 99). OC43 and BCoV are closely related genetically, and OC43 might have resulted from zoonotic spillover of BCoV (100, 101).
- ^ PMID 21849456.
- PMID 20554810.
- PMID 34254725.
- PMID 21994708.
In all members of Betacoronavirus subgroup A, a haemagglutinin esterase (HE) gene, which encodes a glycoprotein with neuraminate O-acetyl-esterase activity and the active site FGDS, is present downstream to ORF1ab and upstream to S gene (Figure 1).
- ^ PMID 15650185.
- ^ Knudsen, Jeppe Kyhne (13 August 2020). "Overraskende opdagelse: Coronavirus har tidligere lagt verden ned" [Surprising discovery: Coronavirus has previously brought down the world]. DR (in Danish). Retrieved 13 August 2020.
A presumed influenza pandemic in 1889 was actually caused by coronavirus, Danish research shows.
- PMID 34464023.
- PMID 27743750.
- PMID 19892230.
- ISBN 978-1-55581-371-0.
- PMID 17346212.
- PMID 34254725.
- ^ PMID 15172021.
- S2CID 31867379.
- PMID 32291278.
- PMID 25757061.
