Human microbiome
The human microbiome is the aggregate of all
The human body hosts many microorganisms, with approximately the same
During early life, the establishment of a diverse and balanced human microbiota plays a critical role in shaping an individual's long-term health.[7] Studies have shown that the composition of the gut microbiota during infancy is influenced by various factors, including mode of delivery, breastfeeding, and exposure to environmental factors.[8] There are several beneficial species of bacteria and potential probiotics present in breast milk.[9] Research has highlighted the beneficial effects of a healthy microbiota in early life, such as the promotion of immune system development, regulation of metabolism, and protection against pathogenic microorganisms.[10] Understanding the complex interplay between the human microbiota and early life health is crucial for developing interventions and strategies to support optimal microbiota development and improve overall health outcomes in individuals.[11]
The Human Microbiome Project (HMP) took on the project of sequencing the genome of the human microbiota, focusing particularly on the microbiota that normally inhabit the skin, mouth, nose, digestive tract, and vagina.[2] It reached a milestone in 2012 when it published its initial results.[12]
Terminology
million years ago) |
Though widely known as flora or microflora, this is a
Relative numbers
The number of
Study

The problem of elucidating the human microbiome is essentially identifying the members of a microbial community, which includes bacteria, eukaryotes, and viruses.
Aside from simply elucidating the composition of the human microbiome, one of the major questions involving the human microbiome is whether there is a "core", that is, whether there is a subset of the community that is shared among most humans.[23][24] If there is a core, then it would be possible to associate certain community compositions with disease states, which is one of the goals of the HMP. It is known that the human microbiome (such as the gut microbiota) is highly variable both within a single subject and among different individuals, a phenomenon which is also observed in mice.[4]
On 13 June 2012, a major milestone of the HMP was announced by the
Analysis after the processing
The statistical analysis is essential to validate the obtained results (
Once a metagenome is assembled, it is possible to infer the functional potential of the microbiome. The computational challenges for this type of analysis are greater than for single genomes, because usually metagenomes assemblers have poorer quality, and many recovered
Marker gene analysis
It is a technique that exploits
Marker gene analysis can be influenced by the primer choice; in this kind of analysis, it is desirable to use a well-validated protocol (such as the one used in the
Another important step in the analysis is to assign a taxonomic name to microbial sequences in the data. This can be done using machine learning approaches that can reach an accuracy at genus-level of about 80%. Other popular analysis packages provide support for taxonomic classification using exact matches to reference databases and should provide greater specificity, but poor sensitivity. Unclassified microorganism should be further checked for organelle sequences.[30]
Phylogenetic analysis
Many methods that exploit phylogenetic inference use the 16SRNA gene for Archea and Bacteria and the 18SRNA gene for Eukaryotes. Phylogenetic comparative methods (PCS) are based on the comparison of multiple traits among microorganisms; the principle is: the closely they are related, the higher number of traits they share. Usually PCS are coupled with phylogenetic generalized least square (PGLS) or other statistical analysis to get more significant results. Ancestral state reconstruction is used in microbiome studies to impute trait values for taxa whose traits are unknown. This is commonly performed with PICRUSt, which relies on available databases. Phylogenetic variables are chosen by researchers according to the type of study: through the selection of some variables with significant biological informations, it is possible to reduce the dimension of the data to analyse.[31]
Phylogenetic aware distance is usually performed with
Ecological Network analysis
Microbial communities develop in a very complex dynamic which can be viewed and analyzed as an ecosystem. The ecological interactions between microbes govern its change, equilibrium and stability, and can be represented by a population dynamic model.[32] The ongoing study of ecological features of the microbiome is growing rapidly and allows to understand the fundamental properties of the microbiome. Understanding the underlying rules of microbial community could help with treating diseases related to unstable microbial communities. A very basic question is if different humans, who share different microbial communities, have the same underlying microbial dynamics.[33] Increasing evidence and indications have found that the dynamics is indeed universal.[34] This question is a basic step that will allow scientists to develop treatment strategies, based on the complex dynamics of human microbial communities. There are more important properties on which considerations should be taken into account for developing interventions strategies for controlling the human microbial dynamics.[35] Controlling the microbial communities could result in solving very bad and harmful diseases.
Types
Bacteria

Populations of microbes (such as
The Human Microbiome Project found that individuals host thousands of bacterial types, different body sites having their own distinctive communities. Skin and vaginal sites showed smaller diversity than the mouth and gut, these showing the greatest richness. The bacterial makeup for a given site on a body varies from person to person, not only in type, but also in abundance. Bacteria of the same species found throughout the mouth are of multiple subtypes, preferring to inhabit distinctly different locations in the mouth. Even the enterotypes in the human gut, previously thought to be well understood, are from a broad spectrum of communities with blurred taxon boundaries.[37][38]
It is estimated that 500 to 1,000
A number of types of bacteria, such as Actinomyces viscosus and A. naeslundii, live in the mouth, where they are part of a sticky substance called plaque. If this is not removed by brushing, it hardens into calculus (also called tartar). The same bacteria also secrete acids that dissolve tooth enamel, causing tooth decay.[citation needed]
The vaginal microflora consist mostly of various lactobacillus species. It was long thought that the most common of these species was Lactobacillus acidophilus, but it has later been shown that L. iners is in fact most common, followed by L. crispatus. Other lactobacilli found in the vagina are L. jensenii, L. delbruekii and L. gasseri. Disturbance of the vaginal flora can lead to infections such as bacterial vaginosis and candiadiasis.[40]
Archaea
Archaea are present in the human gut, but, in contrast to the enormous variety of bacteria in this organ, the numbers of archaeal species are much more limited.[41] The dominant group are the methanogens, particularly Methanobrevibacter smithii and Methanosphaera stadtmanae.[42] However, colonization by methanogens is variable, and only about 50% of humans have easily detectable populations of these organisms.[43]
As of 2007, no clear examples of archaeal pathogens were known,[44][45] although a relationship has been proposed between the presence of some methanogens and human periodontal disease.[46] Methane-dominant small intestinal bacterial overgrowth (SIBO) is also predominently caused by methanogens, and Methanobrevibacter smithii in particular.[47]
Fungi
Fungi, in particular
Viruses
Viruses, especially bacterial viruses (bacteriophages), colonize various body sites. These colonized sites include the skin,[54] gut,[55] lungs,[56] and oral cavity.[57] Virus communities have been associated with some diseases, and do not simply reflect the bacterial communities.[58][59][60]
In January 2024, biologists reported the discovery of "
Anatomical areas
Skin
A study of 20 skin sites on each of ten healthy humans found 205 identified genera in 19 bacterial phyla, with most sequences assigned to four phyla: Actinomycetota (51.8%), Bacillota (24.4%), Pseudomonadota (16.5%), and Bacteroidota (6.3%).[63] A large number of fungal genera are present on healthy human skin, with some variability by region of the body; however, during pathological conditions, certain genera tend to dominate in the affected region.[48] For example, Malassezia is dominant in atopic dermatitis and Acremonium is dominant on dandruff-affected scalps.[48]
The skin acts as a barrier to deter the invasion of pathogenic microbes. The human skin contains microbes that reside either in or on the skin and can be residential or transient. Resident microorganism types vary in relation to skin type on the human body. A majority of microbes reside on superficial cells on the skin or prefer to associate with glands. These glands such as oil or sweat glands provide the microbes with water, amino acids, and fatty acids. In addition, resident bacteria that associated with oil glands are often Gram-positive and can be pathogenic.[2]
Conjunctiva
A small number of bacteria and fungi are normally present in the
Gastrointestinal tract
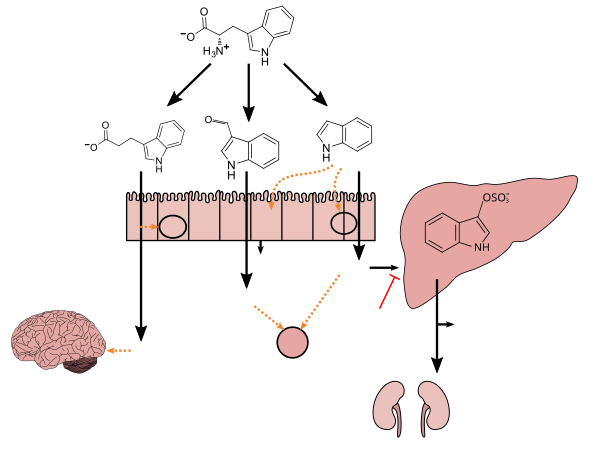
Tryptophan metabolism by
human gastrointestinal microbiota ( )activated charcoal), an intestinal sorbent that is taken by mouth, adsorbs indole, in turn decreasing the concentration of indoxyl sulfate in blood plasma.[65] |
In humans, the composition of the gastrointestinal microbiome is established during birth.
The relationship between some
The composition of human gut microbiota changes over time, when the diet changes, and as overall health changes.
Urethra and bladder
The genitourinary system appears to have a microbiota,[75][76] which is an unexpected finding in light of the long-standing use of standard clinical microbiological culture methods to detect bacteria in urine when people show signs of a urinary tract infection; it is common for these tests to show no bacteria present.[77] It appears that common culture methods do not detect many kinds of bacteria and other microorganisms that are normally present.[77] As of 2017, sequencing methods were used to identify these microorganisms to determine if there are differences in microbiota between people with urinary tract problems and those who are healthy.[75][76][78] To properly assess the microbiome of the bladder as opposed to the genitourinary system, the urine specimen should be collected directly from the bladder, which is often done with a catheter.[79]
Vagina
Vaginal microbiota refers to those species and genera that colonize the vagina. These organisms play an important role in protecting against infections and maintaining vaginal health.
Fungal genera that have been detected in the vagina include Candida, Pichia, Eurotium, Alternaria, Rhodotorula, and Cladosporium, among others.[48]
Placenta
Until recently the placenta was considered to be a sterile organ but commensal, nonpathogenic bacterial species and genera have been identified that reside in the placental tissue.[86][87][88] However, the existence of a microbiome in the placenta is controversial as criticized in several researches. So called "placental microbiome" is likely derived from contamination of regents because low-biomass samples are easily contaminated.[89][90][91]
Uterus
Until recently, the upper reproductive tract of women was considered to be a sterile environment. A variety of microorganisms inhabit the uterus of healthy, asymptomatic women of reproductive age. The microbiome of the uterus differs significantly from that of the vagina and gastrointestinal tract.[92]
Oral cavity
The environment present in the human mouth allows the growth of characteristic microorganisms found there. It provides a source of water and nutrients, as well as a moderate temperature.[2] Resident microbes of the mouth adhere to the teeth and gums to resist mechanical flushing from the mouth to stomach where acid-sensitive microbes are destroyed by hydrochloric acid.[2][50]
Anaerobic bacteria in the oral cavity include:
Bacteria accumulate on both the hard and soft oral tissues in biofilm allowing them to adhere and strive in the oral environment while protected from the environmental factors and antimicrobial agents.[94] Saliva plays a key biofilm homeostatic role allowing recolonization of bacteria for formation and controlling growth by detaching biofilm buildup.[95] It also provides a means of nutrients and temperature regulation. The location of the biofilm determines the type of exposed nutrients it receives.[96]
Oral bacteria have evolved mechanisms to sense their environment and evade or modify the host. However, a highly efficient innate host defense system constantly monitors the bacterial colonization and prevents bacterial invasion of local tissues. A dynamic equilibrium exists between dental plaque bacteria and the innate host defense system.[97]
This dynamic between host oral cavity and oral microbes plays a key role in health and disease as it provides entry into the body.[98]
A healthy equilibrium presents a symbiotic relationship where oral microbes limit growth and adherence of pathogens while the host provides an environment for them to flourish.
Persistent proper oral hygiene is the primary method for preventing oral and systemic disease.[98] It reduces the density of biofilm and overgrowth of potential pathogenic bacteria resulting in disease.[96] However, proper oral hygiene may not be enough as the oral microbiome, genetics, and changes to immune response play a factor in developing chronic infections.[96] Use of antibiotics could treat already spreading infection but ineffective against bacteria within biofilms.[96]
Nasal cavity
The healthy nasal microbiome is dominated by Corynebacterium, and Staphylococcus. The mucosal microbiome plays a critical role in modulating viral infection.[99]
Lung
Much like the oral cavity, the upper and lower respiratory system possess mechanical deterrents to remove microbes. Goblet cells produce mucous which traps microbes and moves them out of the respiratory system via continuously moving ciliated epithelial cells.[2] In addition, a bactericidal effect is generated by nasal mucus which contains the enzyme lysozyme.[2] The upper and lower respiratory tract appears to have its own set of microbiota.[100] Pulmonary bacterial microbiota belong to 9 major bacterial genera: Prevotella, Sphingomonas, Pseudomonas, Acinetobacter, Fusobacterium, Megasphaera, Veillonella, Staphylococcus, and Streptococcus. Some of the bacteria considered "normal biota" in the respiratory tract can cause serious disease especially in immunocompromised individuals; these include Streptococcus pyogenes, Haemophilus influenzae, Streptococcus pneumoniae, Neisseria meningitidis, and Staphylococcus aureus.[citation needed] Fungal genera that compose the pulmonary mycobiome include Candida, Malassezia, Neosartorya, Saccharomyces, and Aspergillus, among others.[48]
Unusual distributions of bacterial and fungal genera in the respiratory tract is observed in people with cystic fibrosis.[48][101] Their bacterial flora often contains antibiotic-resistant and slow-growing bacteria, and the frequency of these pathogens changes in relation to age.[101]
Biliary tract
Traditionally the biliary tract has been considered to be normally sterile, and the presence of microorganisms in bile is a marker of pathological process. This assumption was confirmed by failure in allocation of bacterial strains from the normal bile duct. Papers began emerging in 2013 showing that the normal biliary microbiota is a separate functional layer which protects a biliary tract from colonization by exogenous microorganisms.[102]
Disease and death
Human bodies rely on the innumerable bacterial genes as the source of essential nutrients.[103] Both metagenomic and epidemiological studies indicate vital roles for the human microbiome in preventing a wide range of diseases, from type 2 diabetes and obesity to inflammatory bowel disease, Parkinson's disease, and even mental health conditions like depression.[104] A symbiotic relationship between the gut microbiota and different bacteria may influence an individual's immune response.[105] Metabolites generated by gut microbes appear to be causative factors in type 2 diabetes.[106] Although in its infancy, microbiome-based treatment is also showing promise, most notably for treating drug-resistant C. difficile[dead link] infection[107] and in diabetes treatment.[108]
Clostridioides difficile infection
An overwhelming presence of the bacteria, C. difficile, leads to an infection of the gastrointestinal tract, normally associated to dysbiosis with the microbiota believed to have been caused by the administration of antibiotics. Use of antibiotics eradicates the beneficial gut flora within the gastrointestinal tract, which normally prevents pathogenic bacteria from establishing dominance.[109] Traditional treatment for C. difficile infections includes an additional regime of antibiotics, however, efficacy rates average between 20 and 30%.[110] Recognizing the importance of healthy gut bacteria, researchers turned to a procedure known as fecal microbiota transplant (FMT), where patients experiencing gastrointestinal diseases, such as C. difficile infection (CDI), receive fecal content from a healthy individual in hopes of restoring a normal functioning intestinal microbiota.[111] Fecal microbiota transplant is approximately 85–90% effective in people with CDI for whom antibiotics have not worked or in whom the disease recurs following antibiotics.[112][113] Most people with CDI recover with one FMT treatment.[114][109][115]
Cancer
Although cancer is generally a disease of host genetics and environmental factors, microorganisms are implicated in some 20% of human cancers.
The microbiota may affect carcinogenesis in three broad ways: (i) altering the balance of tumor cell proliferation and death, (ii) regulating immune system function, and (iii) influencing metabolism of host-produced factors, foods and pharmaceuticals.
Concerning the relationship of immune function and development of inflammation, mucosal surface barriers are subject to environmental risks and must rapidly repair to maintain homeostasis. Compromised host or microbiota resiliency also reduce resistance to malignancy, possibly inducing inflammation and cancer. Once barriers are breached, microbes can elicit proinflammatory or immunosuppressive programs through various pathways.[116] For example, cancer-associated microbes appear to activate NF-κΒ signaling within the tumor microenvironment. Other pattern recognition receptors, such as nucleotide-binding oligomerization domain–like receptor (NLR) family members NOD-2, NLRP3, NLRP6 and NLRP12, may play a role in mediating colorectal cancer.[116] Likewise Helicobacter pylori appears to increase the risk of gastric cancer, due to its driving a chronic inflammatory response in the stomach.[116][117]
Inflammatory bowel disease
Inflammatory bowel disease consists of two different diseases: ulcerative colitis and Crohn's disease and both of these diseases present with disruptions in the gut microbiota (also known as dysbiosis). This dysbiosis presents itself in the form of decreased microbial diversity in the gut,[118][119] and is correlated to defects in host genes that changes the innate immune response in individuals.[118]
Human immunodeficiency virus
The
Vaginal microbiota plays a role in the infectivity of HIV, with an increased risk of infection and transmission when the woman has
Gut microbiome of centenarians
Humans who are 100 years old or older, called centenarians, have a distinct gut microbiome. This microbiome is characteristically enriched in microorganisms that are able to synthesize novel secondary bile acids.[122] These secondary bile acids include various isoforms of lithocholic acid that may contribute to healthy aging.[122]
Death
With death, the microbiome of the living body collapses and a different composition of microorganisms named necrobiome establishes itself as an important active constituent of the complex physical decomposition process. Its predictable changes over time are thought to be useful to help determine the time of death.[123][124]
Environmental health
Studies in 2009 questioned whether the decline in
Changes, modulation and transmission

Person-to-person transmission
The human microbiome is transmitted between a
Research
Migration
Cellulose digestion
A 2024 study suggests that
See also
Bibliography
- Ed Yong. I Contain Multitudes: The Microbes Within Us and a Grander View of Life. 368 pages, Published 9 August 2016 by Ecco, ISBN 0062368591.
References
- ^ PMID 26229597., which combined with the environment constitutes the microbiome.
Microbiome
This term refers to the entire habitat, including the microorganisms (bacteria, archaea, lower and higher eurkaryotes, and viruses), their genomes (i.e., genes), and the surrounding environmental conditions. This definition is based on that of "biome," the biotic and abiotic factors of given environments. Others in the field limit the definition of microbiome to the collection of genes and genomes of members of a microbiota. It is argued that this is the definition of metagenome - ^ OCLC 886600661.
- PMID 26824647.
- ^ PMID 24729765.
- PMID 26274026.
we review literature on trimethylamine (TMA), a microbiota-generated metabolite linked to atherosclerosis development.
- PMID 25473158.
Trimethylamine is exclusively a microbiota-derived product of nutrients (lecithin, choline, TMAO, L-carnitine) from normal diet, from which seems originate two diseases, trimethylaminuria (or Fish-Odor Syndrome) and cardiovascular disease through the proatherogenic property of its oxidized liver-derived form.
- ^ Smith A, et al. (2019). "The role of the microbiota in the development of allergies and asthma." Current Allergy and Asthma Reports, 19(8), 38.
- ^ Jackson KD, et al. (2016). "Gut microbiota associations with common diseases and prescription medications in a population-based cohort." Nature Communications, 7, 11622.
- PMID 34578971.
- ^ Yatsunenko T, et al. (2012). "Human gut microbiome viewed across age and geography." Nature, 486(7402), 222-227.
- ^ Sjögren YM, et al. (2009). "Influence of early gut microbiota on the maturation of childhood mucosal and systemic immune responses." Clinical and Experimental Allergy, 39(12), 1842-1851.
- ^ a b "NIH Human Microbiome Project defines normal bacterial makeup of the body". NIH News. 13 June 2012.
- PMID 27541692.
- ISSN 0002-9270.
- PMID 37769422.
- S2CID 190879263.
- PMID 29634682.
- PMID 27327243.
- PMID 20203603.
- ^ PMID 19819907.
- PMID 22179717.
- PMID 18638418.
- PMID 19601958.
- PMID 19383763.
- PMID 22699610.
- PMID 22699609.
- ^ "NIH Human Microbiome Project defines normal bacterial makeup of the body". National Institutes of Health (NIH). 31 August 2015. Retrieved 7 July 2023.
- S2CID 19041044.
- S2CID 24644894.
- ^ S2CID 43936002.
- ^ S2CID 43962376.
- PMID 32605663.
- S2CID 205089271.
- PMID 27279224.
- PMID 36796332.
- S2CID 256268291.
- ^ PLoS Human Microbiome Project Collection Manuscript Summaries Archived 4 March 2014 at the Wayback Machine 13 June 2012
- ^ "Consortium of Scientists Map the Human Body's Bacterial Ecosystem". ucsf.edu. 13 June 2012.
- S2CID 22798964.
- PMID 22553250.
- PMID 15831718.
- S2CID 43706882.
- S2CID 1087298.
- PMID 12540534.
- PMID 14579252.
- PMID 15067114.
- PMID 32754068.
- ^
- ^ S2CID 795450.
Candida species and other microorganisms are involved in this complicated fungal infection, but Candida albicans continues to be the most prevalent. In the past two decades, it has been observed an abnormal overgrowth in the gastrointestinal, urinary and respiratory tracts, not only in immunocompromised patients, but also related to nosocomial infections and even in healthy individuals. There is a widely variety of causal factors that contribute to yeast infection which means that candidiasis is a good example of a multifactorial syndrome.
- ^ S2CID 22101484.
In addition, GI fungal infection is reported even among those patients with normal immune status. Digestive system-related fungal infections may be induced by both commensal opportunistic fungi and exogenous pathogenic fungi. ... Candida sp. is also the most frequently identified species among patients with gastric IFI. ... It was once believed that gastric acid could kill microbes entering the stomach and that the unique ecological environment of the stomach was not suitable for microbial colonisation or infection. However, several studies using culture-independent methods confirmed that large numbers of acid-resistant bacteria belonging to eight phyla and up to 120 species exist in the stomach, such as Streptococcus sp., Neisseria sp. and Lactobacillus sp. etc.26, 27 Furthermore, Candida albicans can grow well in highly acidic environments,28 and some genotypes may increase the severity of gastric mucosal lesions.29
- ^ S2CID 3098136.
Small intestinal fungal overgrowth (SIFO) is characterized by the presence of excessive number of fungal organisms in the small intestine associated with gastrointestinal (GI) symptoms. Candidiasis is known to cause GI symptoms particularly in immunocompromised patients or those receiving steroids or antibiotics. However, only recently, there is emerging literature that an overgrowth of fungus in the small intestine of non-immunocompromised subjects may cause unexplained GI symptoms. Two recent studies showed that 26 % (24/94) and 25.3 % (38/150) of a series of patients with unexplained GI symptoms had SIFO. The most common symptoms observed in these patients were belching, bloating, indigestion, nausea, diarrhea, and gas. ... Fungal-bacterial interaction may act in different ways and may either be synergistic or antagonistic or symbiotic [29]. Some bacteria such as Lactobacillus species can interact and inhibit both the virulence and growth of Candida species in the gut by producing hydrogen peroxide [30]. Any damage to the mucosal barrier or disruption of GI microbiota with chemotherapy or antibiotic use, inflammatory processes, activation of immune molecules and disruption of epithelial repair may all cause fungal overgrowth [27].
- PMID 1576583.
- PMID 3144238.
- PMID 26489866.
- PMID 21880779.
- PMID 25403800.
- PMID 24646696.
- PMID 24846382.
- PMID 26962942.
- PMID 25619688.
- ^ Koumoundouros, Tessa (29 January 2024). "'Obelisks': Entirely New Class of Life Has Been Found in The Human Digestive System". ScienceAlert. Archived from the original on 29 January 2024. Retrieved 29 January 2024.
- from the original on 29 January 2024. Retrieved 29 January 2024.
- PMID 19478181.
- ^ a b "The Normal Bacterial Flora of Humans". textbookofbacteriology.net.
- ^ PMID 27102537.
Lactobacillus spp. convert tryptophan to indole-3-aldehyde (I3A) through unidentified enzymes [125]. Clostridium sporogenes convert tryptophan to IPA [6], likely via a tryptophan deaminase. ... IPA also potently scavenges hydroxyl radicals
Table 2: Microbial metabolites: their synthesis, mechanisms of action, and effects on health and disease
Figure 1: Molecular mechanisms of action of indole and its metabolites on host physiology and disease - PMID 19234110.
Production of IPA was shown to be completely dependent on the presence of gut microflora and could be established by colonization with the bacterium Clostridium sporogenes.
IPA metabolism diagram - ^ "3-Indolepropionic acid". Human Metabolome Database. University of Alberta. Retrieved 12 June 2018.
- S2CID 6630247.
[Indole-3-propionic acid (IPA)] has previously been identified in the plasma and cerebrospinal fluid of humans, but its functions are not known. ... In kinetic competition experiments using free radical-trapping agents, the capacity of IPA to scavenge hydroxyl radicals exceeded that of melatonin, an indoleamine considered to be the most potent naturally occurring scavenger of free radicals. In contrast with other antioxidants, IPA was not converted to reactive intermediates with pro-oxidant activity.
- PMID 26657483.
- PMID 25578246.
- PMID 23818794.
- ^ PMID 24892638.
- ^ PMID 27195115.
- PMID 27413138.
These probiotics showed efficacy in improving psychiatric disorder-related behaviors including anxiety, depression, autism spectrum disorder (ASD), obsessive-compulsive disorder, and memory abilities, including spatial and non-spatial memory. Because many of the basic science studies showed some efficacy of probiotics on central nervous system function, this background may guide and promote further preclinical and clinical studies. ... According to the qualitative analyses of current studies, we can provisionally draw the conclusion that B. longum, B. breve, B. infantis, L. helveticus, L. rhamnosus, L. plantarum, and L. casei were most effective in improving CNS function, including psychiatric disease-associated functions (anxiety, depression, mood, stress response) and memory abilities
- ^ S2CID 27636043.
- ^ PMID 28753805.
- ^ PMID 20539810.
- PMID 33205072.
- PMID 30510275.
- ^ PMID 25859220.
- ^ PMID 17215167.
- ^ Todar K (2012). "The Normal Bacterial Flora of Humans". Todar's Online Textbook of Bacteriology. Madison, WI: Kenneth Todar. Retrieved 6 April 2012.
- PMID 3954346.
- PMID 10558952.
- PMID 32214382.
- PMID 26493119.
- S2CID 206169539.
- ISBN 978-3-319-31248-4.
- S2CID 2462790.
- PMID 31367035.
- S2CID 54166123.
- PMID 26597628.
- PMID 6372039.
- ^ S2CID 40735283.
- PMID 27161350.
- ^ PMID 19485767.
- ^ ISBN 978-1-904455-24-0.
- ^ S2CID 24411104.
- PMID 34433082.
- Wikidata: Q34553608
- ^ S2CID 845977. Archived from the original(PDF) on 16 October 2013.
- PMID 26468308.
- PMID 32255860.
- ^ Copeland CS. The World Within Us: Health and the Human Microbiome. Healthcare Journal of New Orleans, Sept-Oct 2017.
- S2CID 4461492.
- ^ "Serum metabolites reflecting gut microbiome alpha diversity predict type 2 diabetes". Metabolon. Retrieved 3 November 2022.
- PMID 27959316.
- PMID 25802471.
- ^ PMID 21871249.
- PMID 22002980.
- S2CID 44651256.
- S2CID 34998497.
- S2CID 1307726.
- S2CID 2508836.
- S2CID 30849491.
- ^ PMID 25838377.
- ^ PMID 26811603.
- ^ .
- PMID 24574795.
- ^ PMID 27256449.
- S2CID 32045698.
- ^ a b Sato Y, Atarashi K, Plichta DR, Arai Y, Sasajima S, Kearney SM, Suda W, Takeshita K, Sasaki T, Okamoto S, Skelly AN, Okamura Y, Vlamakis H, Li Y, Tanoue T, Takei H, Nittono H, Narushima S, Irie J, Itoh H, Moriya K, Sugiura Y, Suematsu M, Moritoki N, Shibata S, Littman DR, Fischbach MA, Uwamino Y, Inoue T, Honda A, Hattori M, Murai T, Xavier RJ, Hirose N, Honda K. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature. 2021 Nov;599(7885):458-464. doi: 10.1038/s41586-021-03832-5. Epub 2021 Jul 29. PMID 34325466
- ^ Intagliata C (22 December 2016). ""Necrobiome" Reveals a Corpse's Time of Death". Scientific American. Retrieved 26 March 2018.
- ^ Young E (10 December 2015). "Meet the Necrobiome: The Waves of Microbes That Will Eat Your Corpse". The Atlantic. Retrieved 26 March 2018.
- ^ Harmon K (16 December 2009). "Bugs Inside: What Happens When the Microbes That Keep Us Healthy Disappear?". Scientific American. Retrieved 27 December 2008.
- ^ PMID 36180583.
- PMID 32255860.
- PMID 20920376.
- PMID 35562885.
- PMID 36882658.
- PMID 33510866.
- PMID 33651651.
- PMID 28357027.
- S2CID 83461620.
- PMID 29757343.
- PMID 30301893.
- PMID 23861384.
- ^ "Comprehensive profiling of human-to-human oral and intestinal microbial transmission". News-Medical.net. 20 January 2023. Archived from the original on 16 February 2023. Retrieved 16 February 2023.
- PMID 36653448.
- ^ PMID 30388453.
- ^ PMID 31672155.
- PMC 7615765.
- ^ Stewart2024-03-14T18:00:00+00:00, Linda. "Cellulose-degrading gut bacteria found in the human gut, although at lower levels in industrialized countries". The Microbiologist. Retrieved 24 March 2024.
{{cite web}}: CS1 maint: numeric names: authors list (link)
External links
- The Secret World Inside You Exhibit 2015–2016, American Museum of Natural History
- FAQ: Human Microbiome, January 2014 American Society For Microbiology