Hypervariable region
It has been suggested that this article should be split into articles titled Hypervariable region (nucleic acid) and Hypervariable region (antibody). (discuss) (June 2022) |
A hypervariable region (HVR) is a location within nuclear DNA or the D-loop of mitochondrial DNA in which base pairs of nucleotides repeat (in the case of nuclear DNA) or have substitutions (in the case of mitochondrial DNA). Changes or repeats in the hypervariable region are highly polymorphic.
Mitochondrial
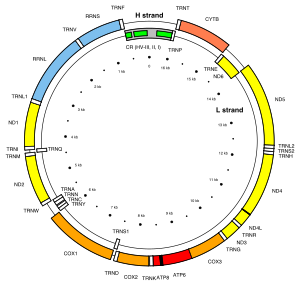
There are two mitochondrial hypervariable regions used in human mitochondrial genealogical DNA testing. HVR1 is considered a "low resolution" region and HVR2 is considered a "high resolution" region. Getting HVR1 and HVR2 DNA tests can help determine one's haplogroup. In the revised Cambridge Reference Sequence of the human mitogenome, the most variable sites of HVR1 are numbered 16024-16383 (this subsequence is called HVR-I), and the most variable sites of HVR2 are numbered 57-372 (i.e., HVR-II) and 438-574 (i.e., HVR-III).[1][2]
In some
Antibodies
In
See also
- Cambridge Reference Sequence
- Genealogical DNA test
- Human mitochondrial DNA haplogroup
- mtDNA control region
References
- PMID 18853457.
- ^ PhyloTree mt. "Annotated mtDNA reference sequences: revised Cambridge Reference Sequence (rCRS)". Retrieved on 4 February 2016.
- (HTML abstract)
- ^ ISBN 9780375761638. Retrieved 5 September 2011.
Antibodies are remarkably specific, thanks to hypervariable regions in both light and heavy chains. The hyperbariable regions for the antigen-binding site.
External links
- Hypervariable+region at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- DNA: Forensic and Legal Applications, Explanation of Hypervariable Regions
