Identity by descent
A

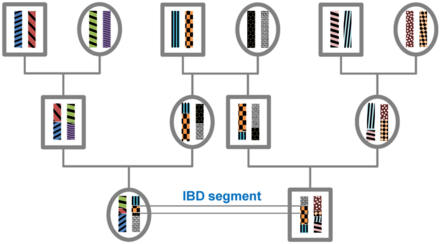
Theory
All individuals in a finite population are related if traced back long enough and will, therefore, share segments of their
Applications
Identified IBD segments can be used for a wide range of purposes. As noted above the amount (length and number) of IBD sharing depends on the familial relationships between the tested individuals. Therefore, one application of IBD segment detection is to quantify relatedness.
IBD mapping
IBD mapping
Using simulated data, Browning and

IBD in population genetics
Detection of natural selection in the human genome is also possible via detected IBD segments. Selection will usually tend to increase the number of IBD segments among individuals in a population. By scanning for regions with excess IBD sharing, regions in the human genome that have been under strong, very recent selection can be identified.[21][22]
In addition to that, IBD segments can be useful for measuring and identifying other influences on population structure.
Methods and software
Programs for the detection of IBD segments in unrelated individuals:
- RAPID: Ultra-fast Identity by Descent Detection in Biobank-Scale Cohorts using Positional Burrows–Wheeler Transform [32]
- Parente: identifies IBD segments between pairs of individuals in unphased genotype data[33]
- BEAGLE/fastIBD: finds segments of IBD between pairs of individuals in genome-wide SNP data[34]
- BEAGLE/RefinedIBD: finds IBD segments in pairs of individuals using a hashing method and evaluates their significance via a likelihood ratio[35]
- IBDseq: detects pairwise IBD segments in sequencing data[36]
- GERMLINE: discovers in linear-time IBD segments in pairs of individuals[5]
- DASH: builds upon pairwise IBD segments to infer clusters of individuals likely to be sharing a single haplotype[15]
- PLINK: is a tool set for whole genome association and population-based linkage analyses including a method for pairwise IBD segment detection[6]
- Relate: estimates the probability of IBD between pairs of individuals at a specific locus using SNPs[3]
- MCMC_IBDfinder: is based on Markov chain Monte Carlo (MCMC) for finding IBD segments in multiple individuals[37]
- IBD-Groupon: detects group-wise IBD segments based on pairwise IBD relationships[38]
- HapFABIA: identifies very short IBD segments characterized by rare variants in large sequencing data simultaneously in multiple individuals[13]
See also
- Association mapping
- Genetic association
- Genetic linkage
- Genome-wide association study
- Identity by type
- Linkage disequilibrium
- Population genetics
References
- PMID 18430938.
- PMID 18282591.
- ^ S2CID 12029712.
- PMID 20303063.
- ^ PMID 18971310.
- ^ PMID 17701901.
- ISBN 978-0-87893-155-2.
- PMID 12900793.
- PMID 16151517.
- PMID 19165921.
- ^ PMID 22135348.
- PMID 19200528.
- ^ PMID 24174545.
- ^ PMID 22267498.
- ^ PMID 21620352.
- S2CID 8131209.
- PMID 19667188.
- PMID 18794890.
- PMID 23472070.
- PMID 22567117.
- PMID 20592267.
- PMID 21769932.
- PMID 6857551.
- ^ PMID 21984068.
- ^ PMID 23103233.
- ^ PMID 23812983.
- PMID 23267057.
- PMID 23733930.
- PMID 23667324.
- PMID 24385924.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - PMID 28108588.
- ^ Naseri A, Liu X, Zhang S, Zhi D. Ultra-fast Identity by Descent Detection in Biobank-Scale Cohorts using Positional Burrows–Wheeler Transform BioRxiv 2017.
- ^ Rodriguez JM, Batzoglou S, Bercovici S. An accurate method for inferring relatedness in large datasets of unphased genotypes via an embedded likelihood-ratio test. RECOMB 2013, LNBI 7821:212-229.
- PMID 21310274.
- PMID 23535385.
- PMID 24207118.
- PMID 21493780.
- PMID 23812980.
