Orthomyxoviridae
| Orthomyxoviridae | |
|---|---|

| |
| Influenza A and influenza B viruses genome, mRNA, and virion diagram | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Negarnaviricota
|
| Class: | Insthoviricetes |
| Order: | Articulavirales |
| Family: | Orthomyxoviridae |
| Genera | |
| |
Orthomyxoviridae (from
The four genera of Influenza virus that infect vertebrates, which are identified by antigenic differences in their
- flu pandemics
- Betainfluenzavirus infects humans and seals
- Gammainfluenzavirus infects humans and pigs
- Deltainfluenzavirus infects pigs and cattle.
Structure

The influenzavirus
The viral envelope composed of a
Genome
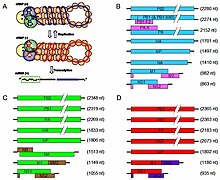
Viruses of the family Orthomyxoviridae contain six to eight segments of linear negative-sense single stranded RNA. They have a total genome length that is 10,000–14,600 nucleotides (nt).[7] The influenza A genome, for instance, has eight pieces of segmented negative-sense RNA (13.5 kilobases total).[8]
The best-characterised of the influenzavirus proteins are hemagglutinin and neuraminidase, two large glycoproteins found on the outside of the viral particles. Hemagglutinin is a lectin that mediates binding of the virus to target cells and entry of the viral genome into the target cell.[9] In contrast, neuraminidase is an enzyme involved in the release of progeny virus from infected cells, by cleaving sugars that bind the mature viral particles. The hemagglutinin (H) and neuraminidase (N) proteins are key targets for antibodies and antiviral drugs,[10][11] and they are used to classify the different serotypes of influenza A viruses, hence the H and N in H5N1.
The genome sequence has terminal repeated sequences; repeated at both ends. Terminal repeats at the 5′-end 12–13 nucleotides long. Nucleotide sequences of 3′-terminus identical; the same in genera of same family; most on RNA (segments), or on all RNA species. Terminal repeats at the 3′-end 9–11 nucleotides long. Encapsidated nucleic acid is solely genomic. Each virion may contain defective interfering copies. In Influenza A (H1N1) PB1-F2 is produced from an alternative reading frame in PB1. The M and NS genes produce two different genes via alternative splicing.[12]
Replication cycle
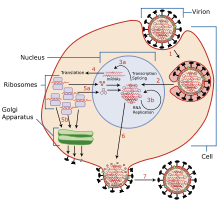
Typically, influenza is transmitted from infected mammals through the air by coughs or sneezes, creating
The viruses bind to a cell through interactions between its
Negative-sense vRNAs that form the genomes of future viruses, RNA-dependent RNA transcriptase, and other viral proteins are assembled into a virion. Hemagglutinin and neuraminidase molecules cluster into a bulge in the cell membrane. The vRNA and viral core proteins leave the nucleus and enter this membrane protrusion (step 6). The mature virus buds off from the cell in a sphere of host phospholipid membrane, acquiring hemagglutinin and neuraminidase with this membrane coat (step 7).[20] As before, the viruses adhere to the cell through hemagglutinin; the mature viruses detach once their neuraminidase has cleaved sialic acid residues from the host cell.[16] After the release of new influenza virus, the host cell dies.

Orthomyxoviridae viruses are one of two RNA viruses that replicate in the nucleus (the other being
Since RNA proofreading enzymes are absent, the RNA-dependent RNA transcriptase makes a single nucleotide insertion error roughly every 10 thousand nucleotides, which is the approximate length of the influenza vRNA. Hence, nearly every newly manufactured influenza virus will contain a mutation in its genome.[23] The separation of the genome into eight separate segments of vRNA allows mixing (reassortment) of the genes if more than one variety of influenza virus has infected the same cell (superinfection). The resulting alteration in the genome segments packaged into viral progeny confers new behavior, sometimes the ability to infect new host species or to overcome protective immunity of host populations to its old genome (in which case it is called an antigenic shift).[10]
Classification
In a
| Genus | Species (* indicates type species) | Serotypes or Subtypes
|
Hosts |
|---|---|---|---|
| Alphainfluenzavirus | Influenza A virus* | H10N7
|
Human, pig, bird, horse, bat |
Betainfluenzavirus
|
Influenza B virus *
|
Victoria, Yamagata[24] | Human, seal |
Gammainfluenzavirus
|
Influenza C virus *
|
Human, pig | |
Deltainfluenzavirus
|
Influenza D virus *
|
Pig, cattle | |
Isavirus
|
Infectious salmon anemia virus *
|
Atlantic salmon | |
| Thogotovirus | Thogotovirus* | Tick, mosquito, mammal (including human) | |
| Dhori virus | Batken virus, Bourbon virus, Jos virus
| ||
| Quaranjavirus[25] | |||
Quaranfil virus,* Johnston Atoll virus
|
Types
There are four genera of influenza virus, each containing only a single species, or type. Influenza A and C infect a variety of species (including humans), while influenza B almost exclusively infects humans, and influenza D infects cattle and pigs.[26][27][28]
Influenza A

Influenza A viruses are further classified, based on the viral surface proteins hemagglutinin (HA or H) and neuraminidase (NA or N). 18 HA subtypes (or serotypes) and 11 NA subtypes of influenza A virus have been isolated in nature. Among these, the HA subtype 1-16 and NA subtype 1-9 are found in wild waterfowl and shorebirds and the HA subtypes 17-18 and NA subtypes 10-11 have only been isolated from bats.[29][30]
Further variation exists; thus, specific influenza strain isolates are identified by a standard nomenclature specifying virus type, geographical location where first isolated, sequential number of isolation, year of isolation, and HA and NA subtype.[31][32]
Examples of the nomenclature are:
- A/Brisbane/59/2007 (H1N1)
- A/Moscow/10/99 (H3N2).
The type A influenza viruses are the most virulent human pathogens among the three influenza types and cause the most severe disease. It is thought that all influenza A viruses causing outbreaks or pandemics originate from wild aquatic birds.
- Swine flu" in 2009.[35]
- H2N2caused "Asian Flu".
- Hong Kong Flu".
- H5N1, "avian" or "bird flu".[36]
- H7N7 has unusual zoonotic potential.[37]
- H1N2 infects pigs and humans.[38]
- H10N7.
| Name of pandemic | Date | Deaths | Case fatality rate | Subtype involved | Pandemic Severity Index
|
|---|---|---|---|---|---|
1889–1890 flu pandemic
(Asiatic or Russian Flu)[41] |
1889–1890 | 1 million | 0.15% | Possibly H2N2 |
— |
1918 flu pandemic
(Spanish flu)[42] |
1918–1920 | 20 to 100 million | 2% | H1N1 |
5 |
Asian Flu
|
1957–1958 | 1 to 1.5 million | 0.13% | H2N2 |
2 |
Hong Kong Flu
|
1968–1969 | 0.75 to 1 million | <0.1% | H3N2 |
2 |
| Russian flu | 1977–1978 | No accurate count | — | H1N1 |
— |
| 2009–2010 | 105,700–395,600[45] | 0.03% | H1N1 |
N/A |
Influenza B

Influenza B virus is almost exclusively a human pathogen, and is less common than influenza A. The only other animal known to be susceptible to influenza B infection is the seal.[46] This type of influenza mutates at a rate 2–3 times lower than type A[47] and consequently is less genetically diverse, with only one influenza B serotype.[26] As a result of this lack of antigenic diversity, a degree of immunity to influenza B is usually acquired at an early age. However, influenza B mutates enough that lasting immunity is not possible.[48] This reduced rate of antigenic change, combined with its limited host range (inhibiting cross species antigenic shift), ensures that pandemics of influenza B do not occur.[49]
Influenza C
The influenza C virus infects humans and pigs, and can cause severe illness and local epidemics.[50] However, influenza C is less common than the other types and usually causes mild disease in children.[51][52]
Influenza D
This is a genus that was classified in 2016, the members of which were first isolated in 2011.[53] This genus appears to be most closely related to Influenza C, from which it diverged several hundred years ago.[54] There are at least two extant strains of this genus.[55] The main hosts appear to be cattle, but the virus has been known to infect pigs as well.
Viability and disinfection
Mammalian influenza viruses tend to be labile, but can survive several hours in mucus.[56] Avian influenza virus can survive for 100 days in distilled water at room temperature, and 200 days at 17 °C (63 °F). The avian virus is inactivated more quickly in manure, but can survive for up to two weeks in feces on cages. Avian influenza viruses can survive indefinitely when frozen.[56] Influenza viruses are susceptible to bleach, 70% ethanol, aldehydes, oxidizing agents, and quaternary ammonium compounds. They are inactivated by heat of 133 °F (56 °C) for minimum of 60 minutes, as well as by low pH <2.[56]
Vaccination and prophylaxis

Vaccines and drugs are available for the prophylaxis and treatment of influenza virus infections. Vaccines are composed of either inactivated or live attenuated virions of the H1N1 and H3N2 human influenza A viruses, as well as those of influenza B viruses. Because the antigenicities of the wild viruses evolve, vaccines are reformulated annually by updating the seed strains.[citation needed]
When the antigenicities of the seed strains and wild viruses do not match, vaccines fail to protect the vaccinees.[citation needed] In addition, even when they do match, escape mutants are often generated.[citation needed]
Drugs available for the treatment of influenza include
See also
References
- ^ International Committee on Taxonomy of Viruses Index of Viruses — Orthomyxovirus (2006). In: ICTVdB—The Universal Virus Database, version 4. Büchen-Osmond, C (Ed), Columbia University, New York.
- PMID 2617637.
- ^ Ely B (1999). "Infectious Salmon Anaemia". Mill Hill Essays. National Institute for Medical Research. Archived from the original on 2007-08-24. Retrieved 2007-09-14.
- PMID 11678233.
- PMID 22291683.
- S2CID 220715148.
- ^ "ICTV Ninth Report; 2009 Taxonomy Release: Orthomyxoviridae". ICTV. Retrieved 19 September 2020.
- PMID 16208317.
- PMID 15744059.
- ^ PMID 12163258.
- PMID 12816348.
- PMID 19230160.
- S2CID 8612187.
- ^ "Avian Influenza (Bird Flu) Implications for Human Disease. Physical characteristics of influenza A viruses". CIDRAP - Center for Infectious Disease Research and Policy. University of Minnesota. 12 March 2024.
- ^ "Flu viruses 'can live for decades' on ice". The New Zealand Herald. Reuters. November 30, 2006. Retrieved November 1, 2011.
- ^ S2CID 30876482.
- PMID 12883000.
- PMID 12921991.
- PMID 16630668.
- PMID 15567494.
- ^ "Cap Snatching". ViralZone. expasy. Retrieved 11 September 2014.
- S2CID 4421958.
- PMID 8387212.
- PMID 20107085.
- ^ ICTV Taxonomy History, ICTV, 2014, archived from the original on 2 April 2015, retrieved 6 June 2006
- ^ PMID 11779385.
- ^ "Avian Influenza (Bird Flu)". Centers for Disease Control and Prevention. Retrieved 2007-09-15.
- PMID 29322273.
- PMID 17126960.
- PMID 24582528.
- ^ Atkinson W, Hamborsky J, McIntyre L, Wolfe S, eds. (2007). Epidemiology and Prevention of Vaccine-Preventable Diseases (10th ed.). Washington DC: Centers for Disease Control and Prevention.
- ^ "Avian Influenza (Bird Flu): Implications for Human Disease". Center for Infectious Disease Research & Policy, University of Minnesota. 2007-06-27. Retrieved 2007-09-14.
- PMID 1579108.
- PMID 20568566.
- PMID 19524497.
- PMID 19618626.
- PMID 14745020.
- PMID 19540800.
- S2CID 26392163.
- ^ "Ten things you need to know about pandemic influenza". World Health Organization. 14 October 2005. Archived from the original on 23 September 2009. Retrieved 26 September 2009.
- PMID 20421481.
- PMID 15602562.
- PMID 20007665.
- ^ "ECDC Daily Update – Pandemic (H1N1) 2009 – January 18, 2010" (PDF). European Centre for Disease Prevention and Control. 2010-01-18. Archived from the original (PDF) on January 22, 2010. Retrieved 2010-01-18.
- (PDF) from the original on Apr 9, 2024 – via Zenodo.
- PMID 10807575.
- PMID 16537638.
- PMID 1579108.
- S2CID 15968981.
- PMID 11825952.
- PMID 16586359.
- PMID 6309999.
- PMID 23408893.
- PMID 23942954.
- PMID 25355894.
- ^ a b c Spickler AR (February 2016). "Influenza" (PDF). The Center for Food Security and Public Health. Iowa State University. p. 7.
- from the original on Jan 21, 2022.
Further reading
- Hoyle, L. (1969). The Influenza Viruses. Virology Monographs. Vol. 4. Springer-Verlag. OCLC 4053391.
External links
- Health-EU Portal: EU work to prepare a global response to influenza.
- Influenza Research Database: Database of influenza genomic sequences and related information.
- European Commission—Public Health: EU coordination on Pandemic (H1N1) 2009
- 3D Influenza-virus-related structures from the EM Data Bank (EMDB)
- Viralzone: Orthomyxoviridae
- Virus Taxonomy: 2020 Release: International Committee on Taxonomy of Viruses (ICTV)