Filamentous bacteriophage
| Inoviridae | |
|---|---|

| |
| Electron micrograph of shadowed filamentous bacteriophage (inovirus) | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Monodnaviria |
| Kingdom: | Loebvirae |
| Phylum: | Hofneiviricota |
| Class: | Faserviricetes |
| Order: | Tubulavirales |
| Family: | Inoviridae |
| Genera | |
Filamentous bacteriophages are a family of viruses (Inoviridae) that infect bacteria, or bacteriophages. They are named for their filamentous shape, a worm-like chain (long, thin and flexible, reminiscent of a length of cooked spaghetti), about 6 nm in diameter and about 1000-2000 nm long.[1][2][3][4][5] This distinctive shape reflects their method of replication: the coat of the virion comprises five types of viral protein, which are located in the inner membrane of the host bacterium during phage assembly, and these proteins are added to the nascent virion's DNA as it is extruded through the membrane. The simplicity of filamentous phages makes them an appealing model organism for research in molecular biology, and they have also shown promise as tools in nanotechnology and immunology.
Characteristics

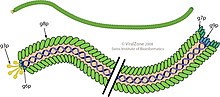
Filamentous bacteriophages are among the simplest living organisms known, with far fewer genes than the classical tailed bacteriophages studied by the phage group in the mid-20th century. The family contains 29 defined species, divided among 23 genera.[6][7] However, mining of genomic and metagenomic datasets using a machine learning approach led to the discovery of 10,295 inovirus-like sequences in nearly all bacterial phyla across virtually every ecosystem, indicating that this group of viruses is much more diverse and widespread than originally appreciated.[5]
Three filamentous bacteriophages, fd, f1 and M13, were isolated and characterized by three different research groups in the early 1960s, but they are so similar that they are sometimes grouped under the common name "Ff", which are members of genus
Structural Class I includes strains fd, f1, M13 of genus Inovirus as well as If1 (of ICTV's species Escherichia virus If1, genus Infulavirus)[11] and IKe (of ICTV's species Salmonella virus IKe, genus Lineavirus),[12] whereas Class II includes strains Pf1 (of ICTV's species Pseudomonas virus Pf1 of genus Primolicivirus),[13] and perhaps also Pf3 (of ICTV's species Pseudomonas virus Pf3 of genus Tertilicivirus),[14] Pf4[15] and PH75 (of NCBI's proposed species Thermus phage PH75, incertae sedis within Inoviridae).[16]
The DNA isolated from fd phage (of genus Inovirus) is single-stranded, and topologically a circle. That is, the DNA single strand extends from one end of the phage particle to the other and then back again to close the circle, although the two strands are not base-paired. This topology was assumed to extend to all other filamentous phages, but it is not the case for phage Pf4,[15] for which the DNA in the phage is single-stranded but topologically linear, not circular.[10] During fd phage assembly, the phage DNA is first packaged into a linear intracellular nucleoprotein complex with many copies of the phage gene 5 replication/assembly protein. The gene 5 protein is then displaced by the gene 8 coat protein as the nascent phage is extruded across the bacterial plasma membrane without killing the bacterial host.[17][18][2][19] This protein also binds with high affinity to G-quadruplex structures (although they are not present in the phage DNA) and to similar hairpin structures in phage DNA.[20]
The p1 protein of Ff phage (i. e. genus Inovirus), which is required for phage assembly at the membrane, has a membrane-spanning hydrophobic domain with the N-terminal portion in the cytoplasm and the C-terminal portion in the periplasm (the reverse of the orientation of the gene 8 coat protein). Adjacent to the cytoplasmic side of the membrane-spanning domain is a 13- residue sequence of p1 having a pattern of basic residues closely matching the pattern of basic residues near the C terminus of p8, but inverted with respect to the sequence. This assembly mechanism makes this phage a valuable system with which to study
Life cycle
Viral replication is cytoplasmic. Entry into the host cell is achieved by pilus-mediated adsorption into the host cell. Replication follows the ssDNA rolling circle model. DNA-templated transcription is the method of transcription. The virus exits the host cell by viral extrusion.[23] Viral assembly occurs at the inner membrane (in case of Gram-negative bacteria), mediated by a membrane-embedded motor protein complex.[23] This multimeric assembly complex, including p1 encoded by gene 1 (referred to as ZOT, zonula occludens toxin by researchers on Vibrio cholerae phage CTXΦ) is an ATPase containing functional and essential Walker motifs[22] that are thought to mediate the hydrolysis of ATP providing the energy for the assembly of the phage filament. Filamentous phage Cf1t from Xanthomonas campestris (of NCBI's proposed species Xanthomonas phage Cf1t, incertae sedis within Inoviridae, likely misspelled as Cflt),[24] was shown in 1987 to integrate into the host bacterial genome, and further such temperate filamentous phages have since been reported, many of which have been implicated in pathogenesis.[1]
Taxonomy
The following genera are recognized:[7]
Phylogenetic trees and clades have been increasingly used to study taxonomy[25] of Inoviridae.[1][3][5][26]
On base of metagenomical data, the family has been proposed to be split into new families Amplinoviridae, Protoinoviridae, Photinoviridae, Vespertilinoviridae, Densinoviridae, and Paulinoviridae, all within order Tubulavirales, of course.[27]
Notable members
- genus F episome.
- species Escherichia virus M13
- M13 bacteriophage
- f1 phage
- species Filamentous bacteriophage fd(proposal)
- fd phage
- genus Affertcholeramvirus[28]
- species Vibrio virus CTXphi
- Vibrio phage CTXphi (aka CTXφ bacteriophage)
- genus Infulavirus)[11]
- species Escherichia virus If1
- If1 phage
- genus Lineavirus[12]
- species Salmonella virus IKe
- IKe phage
- genus Primolicivirus)[13]
- species Pseudomonas virus Pf1
- Pf1 phage
- genus Tertilicivirus)[14]
- species Pseudomonas virus Pf3 — bacteriophages that infect Pseudomonas aeruginosa
- Pf3 phage
Further notable proposed species are:
- species Thermus phage PH75[16]
- PH75 phage
- species Xanthomonas phage Cf1t (likely misspelled as Cflt)[24]
- Cf1t phage
History
The filamentous particle seen in electron micrographs was initially incorrectly interpreted as contaminating bacterial pilus, but ultrasonic degradation, which breaks flexible filaments roughly in half,[29] inactivated infectivity as predicted for a filamentous bacteriophage morphology.[30] Three filamentous bacteriophages, fd, f1 and M13, were isolated and characterized by three different research groups in the early 1960s. Since these three phages differ by less than 2 percent in their DNA sequences, corresponding to changes in only a few dozen codons in the whole genome, for many purposes they can be considered to be identical.[31] Further independent characterization over the subsequent half-century was shaped by the interests of these research groups and their followers.[2]
Filamentous phages, unlike most other phages, are continually extruded through the bacterial membrane without killing the host.[19] Genetic studies on M13 using conditional lethal mutants, initiated by David Pratt and colleagues, led to description of phage gene functions.[32][33] Notably, the protein product of gene 5, which is required for synthesis of progeny single-stranded DNA, is made in large amounts in the infected bacteria,[34][35][36] and it binds to the nascent DNA to form a linear intracellular complex.[17] (The simple numbering of genes using Arabic numerals 1,2,3,4... introduced by the Pratt group is sometimes displaced by the practice of using Roman numerals I, II, III, IV... but the gene numbers defined by the two systems are the same).
Longer (or shorter) DNA can be included in fd phage, since more (or fewer) protein subunits can be added during assembly as required to protect the DNA, making the phage convenient for genetic studies.[37][38] The length of the phage is also affected by the positive charge per length on the inside surface of the phage capsid.[39] The genome of fd was one of the first complete genomes to be sequenced.[40]
The taxonomy of filamentous bacteriophage was defined by
Filamentous bacteriophage engineered to display
References
- ^ PMID 30952693.
- ^ PMID 29900501.
- ^ PMID 25670735.
- ^ PMID 29549632.
- ^ PMID 31332386.
- ^ a b c d "Inoviridae ~ ViralZone". viralzone.expasy.org. Retrieved 31 March 2021.
- ^ a b ICTV. "Virus Taxonomy: 2019 Release". Retrieved 4 July 2020.
- ^ a b NCBI: Inovirus (genus)
- ^ a b c ICTV: ICTV Taxonomy history: Inovirus. 2019 EC 51, Berlin, Germany, July 2019; Email ratification March 2020 (MSL #35).
- ^ PMID 32071243.
- ^ a b NCBI: Infulavirus (genus)
- ^ a b NCBI: Lineavirus (genus)
- ^ a b NCBI: Primolicivirus (genus)
- ^ a b NCBI: Tertilicivirus (genus)
- ^ PMID 32153575.
- ^ a b NCBI: Thermus phage PH75 (species)
- ^ PMID 4594145.
- PMID 2671388.
- ^ PMID 14127828.
- PMID 14992600.
- PMID 7752229.
- ^ PMID 28397779.
- ^ PMID 30556628.
- ^ a b NCBI: Xanthomonas phage Cf1t (species)
- PMID 32341570.
- PMID 31366885.
- ^ Simon Roux, Mart Krupovic, Rebecca A. Daly, Adair L. Borges, Stephen Nayfach, Frederik Schulz, Emiley A. Eloe-Fadrosh et al.: Cryptic inoviruses revealed as pervasive in bacteria and archaea across Earth’s biomes. In: Nature Microbiology, 22 July 2019, doi:10.1038/s41564-019-0510-x
- ^ NCBI: Affertcholeramvirus (genus)
- PMID 13894963.
- S2CID 4224468.
- PMID 22085310.
- PMID 5921643.
- PMID 5807970.
- PMID 4939035.
- PMID 5257006.
- PMID 4115107.
- S2CID 22923713.
- PMID 25941520.
- PMID 1992159.
- PMID 745987.
- PMID 5330240.
- PMID 6811498.
- PMID 2055478.
- S2CID 33495106.
- S2CID 4288812.
- PMID 4001944.
- PMID 22606037.
- S2CID 73434013.
- ^ PMID 26300850.
- PMID 25343220.
- PMID 25058851.
- ISSN 1932-7447.
- PMID 27446051.
- PMID 28357315.)
{{cite journal}}: CS1 maint: DOI inactive as of March 2024 (link
