Ligand-gated ion channel
| Neurotransmitter-gated ion-channel transmembrane region | |||||||||
|---|---|---|---|---|---|---|---|---|---|
TCDB 1.A.9 | | ||||||||
| OPM superfamily | 14 | ||||||||
| OPM protein | 2bg9 | ||||||||
| |||||||||
Ligand-gated ion channels (LICs, LGIC), also commonly referred to as ionotropic receptors, are a group of
When a
These receptor proteins are typically composed of at least two different domains: a transmembrane domain which includes the ion pore, and an extracellular domain which includes the ligand binding location (an
Cys-loop receptors

The
The prototypic ligand-gated ion channel is the nicotinic acetylcholine receptor. It consists of a pentamer of protein subunits (typically ααβγδ), with two binding sites for acetylcholine (one at the interface of each alpha subunit). When the acetylcholine binds it alters the receptor's configuration (twists the T2 helices which moves the leucine residues, which block the pore, out of the channel pathway) and causes the constriction in the pore of approximately 3 angstroms to widen to approximately 8 angstroms so that ions can pass through. This pore allows Na+ ions to flow down their electrochemical gradient into the cell. With a sufficient number of channels opening at once, the inward flow of positive charges carried by Na+ ions depolarizes the postsynaptic membrane sufficiently to initiate an action potential.
A bacterial homologue to an LIC has been identified, hypothesized to act nonetheless as a chemoreceptor.[4] This prokaryotic nAChR variant is known as the GLIC receptor, after the species in which it was identified; Gloeobacter Ligand-gated Ion Channel.
Structure
Cys-loop receptors have structural elements that are well conserved, with a large extracellular domain (ECD) harboring an alpha-helix and 10 beta-strands. Following the ECD, four transmembrane segments (TMSs) are connected by intracellular and extracellular loop structures.[7] Except the TMS 3-4 loop, their lengths are only 7-14 residues. The TMS 3-4 loop forms the largest part of the intracellular domain (ICD) and exhibits the most variable region between all of these homologous receptors. The ICD is defined by the TMS 3-4 loop together with the TMS 1-2 loop preceding the ion channel pore.[7] Crystallization has revealed structures for some members of the family, but to allow crystallization, the intracellular loop was usually replaced by a short linker present in prokaryotic cys-loop receptors, so their structures as not known. Nevertheless, this intracellular loop appears to function in desensitization, modulation of channel physiology by pharmacological substances, and posttranslational modifications. Motifs important for trafficking are therein, and the ICD interacts with scaffold proteins enabling inhibitory synapse formation.[7]
Cationic cys-loop receptors
| Type | Class | IUPHAR-recommended protein name [8] |
Gene | Previous names |
|---|---|---|---|---|
Serotonin (5-HT) |
5-HT3 | 5-HT3A 5-HT3B 5-HT3C 5-HT3D 5-HT3E |
HTR3A HTR3B HTR3C HTR3D HTR3E |
5-HT3A 5-HT3B 5-HT3C 5-HT3D 5-HT3E |
| Nicotinic acetylcholine (nAChR) |
alpha | CHRNA1 CHRNA2 CHRNA3 CHRNA4 CHRNA5 CHRNA6 CHRNA7 CHRNA9 CHRNA10 |
ACHRA, ACHRD, CHRNA, CMS2A, FCCMS, SCCMS | |
| beta | β1 β2 β3 β4 |
CHRNB1 CHRNB2 CHRNB3 CHRNB4 |
CMS2A, SCCMS, ACHRB, CHRNB, CMS1D EFNL3, nAChRB2 | |
| gamma | γ | CHRNG | ACHRG | |
| delta | δ | CHRND | ACHRD, CMS2A, FCCMS, SCCMS | |
| epsilon | ε | CHRNE | ACHRE, CMS1D, CMS1E, CMS2A, FCCMS, SCCMS | |
| Zinc-activated ion channel (ZAC) |
ZAC | ZACN | ZAC1, L2m LICZ, LICZ1 |
Anionic cys-loop receptors
| Type | Class | IUPHAR-recommended protein name[8] |
Gene | Previous names |
|---|---|---|---|---|
| GABAA | alpha | GABRA1 GABRA2 GABRA3 GABRA4 GABRA5 GABRA6 |
EJM, ECA4 | |
| beta | β1 β2 β3 |
GABRB1 GABRB2 GABRB3 |
ECA5 | |
| gamma | γ2 γ3 |
GABRG1 GABRG2 GABRG3 |
CAE2, ECA2, GEFSP3 | |
| delta | δ | GABRD | ||
| epsilon | ε | GABRE | ||
| pi | π | GABRP | ||
| theta | θ | GABRQ | ||
| rho | ρ1 ρ2 ρ3 |
GABRR1 GABRR2 GABRR3 |
GABAC[9] | |
| Glycine (GlyR) |
alpha | GLRA1 GLRA2 GLRA3 GLRA4 |
STHE | |
| beta | β | GLRB |
Ionotropic glutamate receptors
The ionotropic glutamate receptors bind the neurotransmitter glutamate. They form tetramers, with each subunit consisting of an extracellular amino terminal domain (ATD, which is involved tetramer assembly), an extracellular ligand binding domain (LBD, which binds glutamate), and a transmembrane domain (TMD, which forms the ion channel). The transmembrane domain of each subunit contains three transmembrane helices as well as a half membrane helix with a reentrant loop. The structure of the protein starts with the ATD at the N terminus followed by the first half of the LBD which is interrupted by helices 1,2 and 3 of the TMD before continuing with the final half of the LBD and then finishing with helix 4 of the TMD at the C terminus. This means there are three links between the TMD and the extracellular domains. Each subunit of the tetramer has a binding site for glutamate formed by the two LBD sections forming a clamshell like shape. Only two of these sites in the tetramer need to be occupied to open the ion channel. The pore is mainly formed by the half helix 2 in a way which resembles an inverted potassium channel.

| Type | Class | IUPHAR-recommended protein name [8] |
Gene | Previous names |
|---|---|---|---|---|
| AMPA | GluA | GluA1 GluA2 GluA3 GluA4 |
GRIA1 GRIA2 GRIA3 GRIA4 |
GLUA1, GluR1, GluRA, GluR-A, GluR-K1, HBGR1 GLUA2, GluR2, GluRB, GluR-B, GluR-K2, HBGR2 GLUA3, GluR3, GluRC, GluR-C, GluR-K3 GLUA4, GluR4, GluRD, GluR-D |
| Kainate | GluK | GluK1 GluK2 GluK3 GluK4 GluK5 |
GRIK1 GRIK2 GRIK3 GRIK4 GRIK5 |
GLUK5, GluR5, GluR-5, EAA3 GLUK6, GluR6, GluR-6, EAA4 GLUK7, GluR7, GluR-7, EAA5 GLUK1, KA1, KA-1, EAA1 GLUK2, KA2, KA-2, EAA2 |
| NMDA | GluN | GluN1 NRL1A NRL1B |
GRIN1 GRINL1A GRINL1B |
GLUN1, NMDA-R1, NR1, GluRξ1 |
| GluN2A GluN2B GluN2C GluN2D |
GRIN2A GRIN2B GRIN2C GRIN2D |
GLUN2A, NMDA-R2A, NR2A, GluRε1 GLUN2B, NMDA-R2B, NR2B, hNR3, GluRε2 GLUN2C, NMDA-R2C, NR2C, GluRε3 GLUN2D, NMDA-R2D, NR2D, GluRε4 | ||
| GluN3A GluN3B |
GRIN3A GRIN3B |
GLUN3A, NMDA-R3A, NMDAR-L, chi-1 GLU3B, NMDA-R3B | ||
| ‘Orphan’ | (GluD) | GluD1 GluD2 |
GRID1 GRID2 |
GluRδ1 GluRδ2 |
AMPA receptor

The α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptor (also known as
- , etc.
- Antagonists: CNQX, Kynurenic acid, NBQX, Perampanel, Piracetam, etc.
- CX-614, etc.
- GYKI-52,466, etc.
NMDA receptors
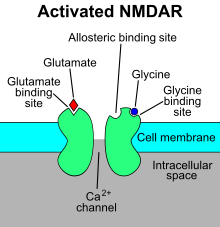
The N-methyl-D-aspartate receptor (
The name "NMDA receptor" is derived from the ligand
- Primary D-serine or glycine
- Other quinolinate, etc.
- Partial agonists : NRX-1074; 3,5-dibromo-L-phenylalanine,[15]etc.
- endogenous), etc.
ATP-gated channels

ATP-gated channels open in response to binding the nucleotide ATP. They form trimers with two transmembrane helices per subunit and both the C and N termini on the intracellular side.
| Type | Class | IUPHAR-recommended protein name [8] |
Gene | Previous names |
|---|---|---|---|---|
P2X
|
N/A | P2X1 P2X2 P2X3 P2X4 P2X5 P2X6 P2X7 |
P2RX1 P2RX2 P2RX3 P2RX4 P2RX5 P2RX6 P2RX7 |
P2X1 P2X2 P2X3 P2X4 P2X5 P2X6 P2X7 |
Clinical relevance
Ligand-gated ion channels are likely to be the major site at which
By understanding the mechanism and exploring the chemical/biological/physical component that could function on those receptors, more and more clinical applications are proven by preliminary experiments or
See also
- Action potential
- Acid-sensing ion channel
- Calcium-activated potassium channel
- Cyclic nucleotide-gated ion channel
- Voltage-dependent calcium channel
- Receptor (biochemistry)
- Inositol trisphosphate receptor
- Metabotropic receptor
- Ryanodine receptor
References
- ^ "Gene Family: Ligand gated ion channels". HUGO Gene Nomenclature Committee.
- ^ "ligand-gated channel" at Dorland's Medical Dictionary
- ISBN 978-0-87893-697-7.)
{{cite book}}: CS1 maint: multiple names: authors list (link - ^ PMID 15642096.
- PMID 26986966.
- PMID 15023997.
- ^ PMID 27330534.
- ^ PMID 18655795.
- PMID 18790874.
- S2CID 42753770.
- ISBN 9780071481274.
At membrane potentials more negative than approximately −50 mV, the Mg2+ in the extracellular fluid of the brain virtually abolishes ion flux through NMDA receptor channels, even in the presence of glutamate. ... The NMDA receptor is unique among all neurotransmitter receptors in that its activation requires the simultaneous binding of two different agonists. In addition to the binding of glutamate at the conventional agonist-binding site, the binding of glycine appears to be required for receptor activation. Because neither of these agonists alone can open this ion channel, glutamate and glycine are referred to as coagonists of the NMDA receptor. The physiologic significance of the glycine binding site is unclear because the normal extracellular concentration of glycine is believed to be saturating. However, recent evidence suggests that D-serine may be the endogenous agonist for this site.
- PMID 19605837.
- S2CID 15442694.
- PMID 10049997.
- S2CID 11672391.
- PMID 10487207.
- PMID 12173240.
- S2CID 17913232.
- S2CID 31877708.
- ^ NICE technology appraisal January 18, 2011 Azheimer's disease - donepezil, galantamine, rivastigmine and memantine (review): final appraisal determination
- S2CID 144761669.
- ISBN 978-0-07-162442-8.
- S2CID 145014277.
External links
- Ligand-Gated Ion Channel database at European Bioinformatics Institute. Verified availability April 11, 2007.
- "Revised Recommendations for Nomenclature of Ligand-Gated Ion Channels". IUPHAR Database of Receptors and Ion Channels. International Union of Basic and Clinical Pharmacology.
- www.esf.edu
- www.genenames.org
- www.guidetopharmacology.org
As of this edit, this article uses content from "1.A.9 The Neurotransmitter Receptor, Cys loop, Ligand-gated Ion Channel (LIC) Family", which is licensed in a way that permits reuse under the
