Lysosome
Animal cell diagram | |
|---|---|
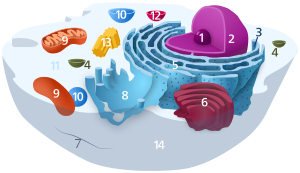 Components of a typical animal cell:
|
A lysosome (

Lysosomes are degradative organelles that act as the waste disposal system of the cell by digesting used materials in the cytoplasm, from both inside and outside the cell. Material from outside the cell is taken up through endocytosis, while material from the inside of the cell is digested through autophagy.[6] The sizes of the organelles vary greatly—the larger ones can be more than 10 times the size of the smaller ones.[7] They were discovered and named by Belgian biologist Christian de Duve, who eventually received the Nobel Prize in Physiology or Medicine in 1974.
Lysosomes contain more than 60 different enzymes, and have more than 50 membrane proteins.
In 2009, Marco Sardiello and co-workers discovered that the synthesis of most lysosomal enzymes and membrane proteins is controlled by transcription factor EB (
Etymology and pronunciation
The word lysosome (
Discovery

They succeeded in detecting the enzyme activity from the microsomal fraction. This was the crucial step in the serendipitous discovery of lysosomes. To estimate this enzyme activity, they used that of the standardized enzyme acid phosphatase and found that the activity was only 10% of the expected value. One day, the enzyme activity of purified cell fractions which had been refrigerated for five days was measured. Surprisingly, the enzyme activity was increased to normal of that of the fresh sample. The result was the same no matter how many times they repeated the estimation, and led to the conclusion that a membrane-like barrier limited the accessibility of the enzyme to its substrate, and that the enzymes were able to diffuse after a few days (and react with their substrate). They described this membrane-like barrier as a "saclike structure surrounded by a membrane and containing acid phosphatase."[18]
It became clear that this enzyme from the cell fraction came from membranous fractions, which were definitely cell organelles, and in 1955 De Duve named them "lysosomes" to reflect their digestive properties.
Originally, De Duve had termed the organelles the "suicide bags" or "suicide sacs" of the cells, for their hypothesized role in apoptosis.[22] However, it has since been concluded that they only play a minor role in cell death.[23]
Function and structure
Lysosomes contain a variety of enzymes, enabling the cell to break down various biomolecules it engulfs, including
). The enzymes responsible for this hydrolysis require an acidic environment for optimal activity.In addition to being able to break down polymers, lysosomes are capable of fusing with other organelles & digesting large structures or cellular debris; through cooperation with phagosomes, they are able to conduct autophagy, clearing out damaged structures. Similarly, they are able to break down virus particles or bacteria in phagocytosis of macrophages.
The size of lysosomes varies from 0.1
The lysosome maintains its pH differential by pumping in protons (H+ ions) from the cytosol across the membrane via proton pumps and chloride ion channels. Vacuolar-ATPases are responsible for transport of protons, while the counter transport of chloride ions is performed by ClC-7 Cl−/H+ antiporter. In this way a steady acidic environment is maintained.[25][26]
It sources its versatile capacity for degradation by import of enzymes with specificity for different substrates; cathepsins are the major class of hydrolytic enzymes, while lysosomal alpha-glucosidase is responsible for carbohydrates, and lysosomal acid phosphatase is necessary to release phosphate groups of phospholipids.
Recent research also indicates that lysosomes can act as a source of intracellular calcium.[27]
Formation

Many components of animal cells are recycled by transferring them inside or embedded in sections of membrane. For instance, in
The production of lysosomal proteins suggests one method of lysosome sustainment. Lysosomal protein genes are transcribed in the
Upon leaving the Golgi apparatus, the lysosomal enzyme-filled vesicle fuses with a late endosome, a relatively acidic organelle with an approximate pH of 5.5. This acidic environment causes dissociation of the lysosomal enzymes from the mannose 6-phosphate receptors. The enzymes are packed into vesicles for further transport to established lysosomes.[30] The late endosome itself can eventually grow into a mature lysosome, as evidenced by the transport of endosomal membrane components from the lysosomes back to the endosomes.[28]
Pathogen entry
As the endpoint of endocytosis, the lysosome also acts as a safeguard in preventing pathogens from being able to reach the cytoplasm before being degraded. Pathogens often hijack endocytotic pathways such as pinocytosis in order to gain entry into the cell. The lysosome prevents easy entry into the cell by hydrolyzing the biomolecules of pathogens necessary for their replication strategies; reduced lysosomal activity results in an increase in viral infectivity, including HIV.[31] In addition, AB5 toxins such as cholera hijack the endosomal pathway while evading lysosomal degradation.[31]
Clinical significance
Lysosomes are involved in a group of genetically inherited deficiencies, or mutations called
There is no direct medical treatment to cure LSDs.
The most severe and rarely found, lysosomal storage disease is
Dysfunctional lysosome activity is also heavily implicated in the biology of aging, and age-related diseases such as Alzheimer's, Parkinson's, and cardiovascular disease.[17][39]
Different enzymes present in Lysosomes
| Sr. No | Enzymes | Substrate |
|---|---|---|
| 1 | Phosphates | |
| A- Acid phosphatase | Most phosphomonoesters | |
| B- Acid phosphodiesterase | Oligonucleotides and phosphodiesterase | |
| 2 | Nucleases | |
| A- Acid ribonuclease | RNA | |
| B- Acid deoxyribonuclease | DNA | |
| 3 | Polysaccharides/ mucopolysaccharides hydrolyzing enzymes | |
| A- β-Galactosidase | Galactosides | |
| B- α-Glucosidase | Glycogen | |
| C- α-Mannosidase | Mannosides, glycoproteins | |
| D- β- Glucoronidase | Polysaccharides and mucopolysaccharides | |
| E- Lysozymes | Bacterial cell walls and mucopolysaccharides | |
| F- Hyaluronidase | Hyaluronic acids, chondroitin sulfates | |
| H- Arylsulphatase | Organic sulfates | |
| 4 | Proteases | |
| A- Cathepsin(s) | Proteins | |
| B- Collagenase | Collagen | |
| C- Peptidase | Peptides | |
| 5 | Lipid degrading enzymes | |
| A- Esterase | Fatty acyl esters | |
| B- Phospholipase | Phospholipids | |
| 6 | Sulfatases | |
| A- Arylsulfatase(A, B & G) | O- and N-Sulfate esters | |
| B- Glucosamine (N-acetyl)-6-Sulfatase/GNS | Glycosaminoglycans | |
| C- Iduronate 2-Sulfatase/IDS | O- and N-Sulfate esters |
Lysosomotropism
Weak bases with lipophilic properties accumulate in acidic intracellular compartments like lysosomes. While the plasma and lysosomal membranes are permeable for neutral and uncharged species of weak bases, the charged protonated species of weak bases do not permeate biomembranes and accumulate within lysosomes. The concentration within lysosomes may reach levels 100 to 1000 fold higher than extracellular concentrations. This phenomenon is called lysosomotropism,[41] "acid trapping" or "proton pump" effect.[42] The amount of accumulation of lysosomotropic compounds may be estimated using a cell-based mathematical model.[43]
A significant part of the clinically approved drugs are lipophilic weak bases with lysosomotropic properties. This explains a number of pharmacological properties of these drugs, such as high tissue-to-blood concentration gradients or long tissue elimination half-lives; these properties have been found for drugs such as haloperidol,[44] levomepromazine,[45] and amantadine.[46] However, high tissue concentrations and long elimination half-lives are explained also by lipophilicity and absorption of drugs to fatty tissue structures. Important lysosomal enzymes, such as acid sphingomyelinase, may be inhibited by lysosomally accumulated drugs.[47][48] Such compounds are termed FIASMAs (functional inhibitor of acid sphingomyelinase)[49] and include for example fluoxetine, sertraline, or amitriptyline.
Ambroxol is a lysosomotropic drug of clinical use to treat conditions of productive cough for its mucolytic action. Ambroxol triggers the exocytosis of lysosomes via neutralization of lysosomal pH and calcium release from acidic calcium stores.[50] Presumably for this reason, Ambroxol was also found to improve cellular function in some disease of lysosomal origin such as Parkinson's or lysosomal storage disease.[51][52]
Systemic lupus erythematosus
Impaired lysosome function is prominent in systemic lupus erythematosus preventing macrophages and monocytes from degrading neutrophil extracellular traps[53] and immune complexes.[54][55][56] The failure to degrade internalized immune complexes stems from chronic mTORC2 activity, which impairs lysosome acidification.[57] As a result, immune complexes in the lysosome recycle to the surface of macrophages causing an accumulation of nuclear antigens upstream of multiple lupus-associated pathologies.[54][58][59]
Controversy in botany
By scientific convention, the term lysosome is applied to these vesicular organelles only in animals, and the term vacuole is applied to those in plants, fungi and algae (some animal cells also have vacuoles). Discoveries in plant cells since the 1970s started to challenge this definition. Plant vacuoles are found to be much more diverse in structure and function than previously thought.[60][61] Some vacuoles contain their own hydrolytic enzymes and perform the classic lysosomal activity, which is autophagy.[62][63][64] These vacuoles are therefore seen as fulfilling the role of the animal lysosome. Based on de Duve's description that "only when considered as part of a system involved directly or indirectly in intracellular digestion does the term lysosome describe a physiological unit", some botanists strongly argued that these vacuoles are lysosomes.[65] However, this is not universally accepted as the vacuoles are strictly not similar to lysosomes, such as in their specific enzymes and lack of phagocytic functions.[66] Vacuoles do not have catabolic activity and do not undergo exocytosis as lysosomes do.[67]
See also
- Peroxisome
- Cathelicidin
- Antimicrobial peptides
- Innate immune system
- TMEM106B
References
- PMID 33803964.
- vacuoles, see § Controversy in botany
- PMID 28524.
- PMID 23609508.
- ^ Holtzclaw FW, et al. (2008). AP* Biology: to Accompany Biology (8th AP ed.). Pearson Benjamin Cummings.
- ^ S2CID 187669426.
- ISBN 978-0-387-28957-1.
- PMID 25668017.
- ^ "Lysosomal Enzymes". www.rndsystems.com. R&D Systems. Retrieved 4 October 2016.
- ^ PMID 30397314.
- ^ PMID 32597833.
- S2CID 24493663.
- PMID 24668941.
- ^ S2CID 20353685.
- ^ PMID 23185029.
- PMID 23416930.
- ^ PMID 27125853.
- ^ Susana Castro-Obregon (2010). "The Discovery of Lysosomes and Autophagy". Nature Education. 3 (9): 49.
- S2CID 30307451.
- PMID 13357540.
- PMID 18567941.
- ^ Hayashi, Teru, and others. "Subcellular Particles." Subcellular Particles., 1959.
- PMID 19473965.
- ISBN 978-1-58890-175-0.
- PMID 22335796.
- PMID 23712550.
- PMID 25720963.
- ^ ISBN 978-0-8153-3218-3.
- PMID 17077125.
- ^ ISBN 978-0-7167-3136-8.
- ^ PMID 15827185.
- PMID 21723623.
- PMID 22647656.
- PMID 23165354.
- PMID 23510062.
- PMID 23114583.
- PMID 28502768.
- ISBN 978-0815340720.
- PMID 30936119.
- OCLC 857764171.
- PMID 4606365.
- PMID 7532261.
- PMID 18504571.
- S2CID 7258546.
- S2CID 24735371.
- S2CID 25784783.
- PMID 18027916.
- PMID 21909365.
- PMID 20502000.
- PMID 26560688.
- PMID 24771397.
- PMID 24574503.
- PMID 20439745.
- ^ PMID 27035940.
- PMID 17643939.
- S2CID 23685272.
- PMID 29866702.
- PMID 27059595.
- PMID 28235864.
- PMID 10213780.
- PMID 16006126.
- .
- PMID 12226358.
- PMID 21980143.
- PMID 9596630.
- ISBN 978-0306-4-3126-5.
- ISBN 978-0-643-09944-9.
External links
 This article incorporates NCBI. Archived from the originalon 8 December 2009.
This article incorporates NCBI. Archived from the originalon 8 December 2009.- 3D structures of proteins associated with lysosome membrane
- Hide and Seek Foundation For Lysosomal Research
- Lysosomal Disease Network, a research consortium funded by the NIH through its NCATS/Rare Diseases Clinical Research Network
- Self-Destructive Behavior in Cells May Hold Key to a Longer Life
- Mutations in the Lysosomal Enzyme–Targeting Pathway and Persistent Stuttering
- Animation showing how lysosomes are made, and their function
