Major capsid protein VP1
| Major capsid protein VP1 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 A rendering of an icosahedral viral capsid comprising 72 pentamers of murine polyomavirus VP1, colored such that areas of the surface closer to the interior center appear blue and areas further away appear red. Rendered from PDB: 1SIE. | |||||||||
| Identifiers | |||||||||
| Symbol | VP1 | ||||||||
| Pfam | PF00718 | ||||||||
| InterPro | IPR000662 | ||||||||
| |||||||||
Major capsid protein VP1 is a
icosahedral structure consisting of 360 VP1 molecules organized into 72 pentamers. VP1 molecules possess a surface binding site that interacts with sialic acids attached to glycans, including some gangliosides, on the surfaces of cells to initiate the process of viral infection. The VP1 protein, along with capsid components VP2 and VP3, is expressed from the "late region" of the circular viral genome.[1][2][3]
Structure
VP1 is the major structural component of the
disulfide bonds.[7]

The VP1 protein monomer is primarily composed of
disordered and forms interactions between neighboring pentamers in the assembled capsid. The flexibility of the C-terminal arm will enable it to adopt different conformations in the six distinct interaction environments imposed by the symmetry of the icosahedral assembly.[4][8] The C-terminus also contains a basic nuclear localization sequence,[5]: 316 while the N-terminus - which is oriented toward the center of the assembled capsid - contains basic residues that facilitate non-sequence-specific interactions with DNA.[9]

Function and trafficking
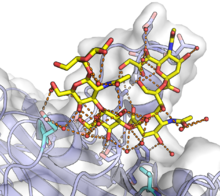
The VP1 protein is responsible for initiating the process of infecting a cell by binding to
disulfide bonds, likely occur in the ER. For some polyomaviruses, VP1 has been detected reaching the nucleus along with the viral genome, though it is unclear how the genomic DNA disengages from VP1.[12]
All of the capsid proteins are expressed from the late region of the viral genome, so named because expression occurs only late in the infection process. VP1 has a
oligomerization to form capsids occurs in the nucleus.[5]
: 316–17
