Marine microbiome
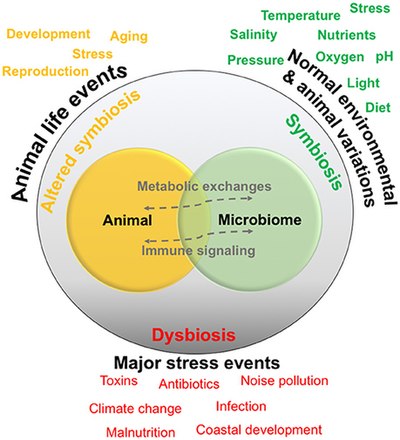
All animals on Earth form associations with microorganisms, including protists, bacteria, archaea, fungi, and viruses. In the ocean, animal–microbial relationships were historically explored in single host–symbiont systems. However, new explorations into the diversity of marine microorganisms associating with diverse marine animal hosts is moving the field into studies that address interactions between the animal host and a more multi-member microbiome. The potential for microbiomes to influence the health, physiology, behavior, and ecology of marine animals could alter current understandings of how marine animals adapt to change, and especially the growing climate-related and anthropogenic-induced changes already impacting the ocean environment.[1]
In the oceans, it is challenging to find
Background
| Part of a series on |
| Microbiomes |
|---|
 |
Within the vast
There are many outstanding questions in ecology and evolution that could be addressed by expanding the phylogenetic and ecological breadth of host-associated microbiome studies, including all possible interactions throughout the microbiome. There is strong empirical evidence and new consensus that biodiversity (i.e., the richness of species and their interactions) pervasively influences the functioning of Earth's ecosystems, including ecosystem productivity.[15][16] However, this research has focused almost exclusively on macroorganisms. Because microbial symbionts are integral parts of most living organisms,[11][17] the understanding of how microbial symbionts contribute to host performance and adaptability needs broadening.[2]
Foundations of productive ecosystems

Black circle: macronucleus, white big circle: food vacuoles, green circles: phototrophs, brown circles: chemoautotrophs, yellow ovals: heterotrophic prokaryotes

Along with more standard examples of nutritional symbioses in animals, recent advances in
Reproduction and host development
| Part of a series of overviews on |
| Marine life |
|---|
 |
Extending beyond nutritional symbioses, microbial symbionts can alter the reproduction, development, and growth of their hosts. Specific bacterial strains in marine biofilms often directly control the recruitment of planktonic larvae and propagules, either by inhibiting settlement or by serving as a settlement cue.[29][30] For example, the settlement of zoospores from the green alga Ulva intestinalis onto the biofilms of specific bacteria is mediated by their attraction to the quorum-sensing molecule, acyl-homoserine lactone, secreted by the bacteria.[31] Classic examples of marine host–microbe developmental dependence include the observation that algal cultures grown in isolation exhibited abnormal morphologies [32] and the subsequent discovery of morphogenesis-inducing compounds, such as thallusin, secreted by epiphytic bacterial symbionts.[33] Bacteria are also known to influence the growth of marine plants, macroalgae, and phytoplankton by secreting phytohormones such as indole acetic acid and cytokinin-type hormones.[34][35][36] In the marine choanoflagellate Salpingoeca rosetta, both multicellularity and reproduction are triggered by specific bacterial cues, offering a view into the origins of bacterial control over animal development (reviewed by Woznica and King.[37] The benefit to the bacteria, in return, is that they receive physical space to colonize at particular points in the water column typically accessible only to planktonic microbes. Perhaps the best-studied example of intimate host–microbe interactions controlling animal development is the Hawaiian bobtail squid Euprymna scolopes.[38] It lives in a mutualistic symbiosis with the bioluminescent bacteria Aliivibrio fischeri. The bacteria are fed a solution of sugars and amino acids by the host and, in return, provide bioluminescence for countershading and predator avoidance.[7] This mutualism with microbes provides a selective advantage for the squid in predator–prey interactions. Another invertebrate example can be found in tubeworms, in which Hydroides elegans metamorphosis is mediated by a bacterial inducer and mitogen-activated protein kinase (MAPK) signaling in biofilms.[39][2]
Biofouling and microbial community assembly
(G) a humpback whale breaching and (H) a SEM image of a humpback's skin surface associated bacteria, with arrows indicating two different cell morphologies.[1]
Some host-associated microbes produce compounds that prevent biofouling and regulate microbiome assembly and maintenance in many marine organisms, including sponges, macroalgae, and corals.[40][41] For example, tropical corals harbor diverse bacteria in their surface mucus layer that produce quorum-sensing inhibitors and other antibacterial compounds as a defense against colonization and infection by potential microbial pathogens.[3] Epiphytic bacteria of marine macroalgae excrete a diverse chemical arsenal capable of selectively shaping further bacterial colonization and deterring the settlement of biofouling marine invertebrates such as bryozoans.[34][42] As in corals, these diverse, microbially secreted compounds include not only bactericidal and bacteriostatic antibiotics but also compounds like halogenated furanones, cyclic dipeptides, and acyl-homoserine lactone mimics that disrupt bacterial quorum sensing and inhibit biofilm formation.[43] The bacteria likely are able to utilize the carbon-rich exudates from their hosts.[44][45] For example, in the case of giant kelp, the alga emits approximately 20% of primary production as dissolved organic carbon.[45] Whereas these prior examples illustrate how the microbiomes can protect hosts from surface colonization, a similar phenomenon has also been observed internally in the shipworm Bankia setacea, in which symbionts produce a boronated tartrolon antibiotic thought to keep the wood-digesting cecum clear of bacterial foulants.[46] By producing antimicrobial compounds, these microbes are able to defend their niche space to prevent other organisms from crowding them out.[2]
Biogeochemical cycling
Host-associated microbiomes also influence biogeochemical cycling within ecosystems with cascading effects on biodiversity and ecosystem processes. For example, microbial symbionts comprise up to 40% of the biomass of their sponge hosts.
These examples demonstrate the importance of microbial symbioses for the functioning of ocean ecosystems. Understanding symbioses with this same level of detail in the context of complex communities (i.e., whole microbiomes) remains ripe for exploration and, indeed, requires a more integrated framework from the fields of microbiology, evolutionary biology, community ecology, and oceanography. Individual taxa within the microbiome may help hosts withstand a wide range of environmental conditions, including those predicted under scenarios of climate change. Next, we explore two different avenues of how interdisciplinary collaborations could advance this line of research.[2]
Examples
The microbiomes of diverse marine animals are currently under study, from simplistic organisms including sponges[55] and ctenophores [56] to more complex organisms such as sea squirts[57] and sharks.[58][1]
The relationship between the
The gutless marine
Corals
Corals are one of the more common examples of an animal host whose symbiosis with microalgae can turn to dysbiosis, and is visibly detected as bleaching. Coral microbiomes have been examined in a variety of studies, which demonstrate how variations in the ocean environment, most notably temperature, light, and inorganic nutrients, affect the abundance and performance of the microalgal symbionts, as well as calcification and physiology of the host.[68][69][1]
Studies have also suggested that resident bacteria, archaea, and fungi additionally contribute to nutrient and organic matter cycling within the coral, with viruses also possibly playing a role in structuring the composition of these members, thus providing one of the first glimpses at a multi-domain marine animal symbiosis.
Sponges
Sponges are common members of the ocean's diverse benthic habitats and their abundance and ability to filter large volumes of seawater have led to the awareness that these organisms play critical roles in influencing benthic and pelagic processes in the ocean.[76] They are one of the oldest lineages of animals, and have a relatively simple body plan that commonly associates with bacteria, archaea, algal protists, fungi, and viruses.[4] Sponge microbiomes are composed of specialists and generalists, and complexity of their microbiome appears to be shaped by host phylogeny.[77] Studies have shown that the sponge microbiome contributes to nitrogen cycling in the oceans, especially through the oxidation of ammonia by archaea and bacteria.[78][79] Most recently, microbial symbionts of tropical sponges were shown to produce and store polyphosphate granules,[80] perhaps enabling the host to survive periods of phosphate depletion in oligotrophic marine environments.[81] The microbiomes of some sponge species do appear to change in community structure in response to changing environmental conditions, including temperature[82] and ocean acidification,[83][84] as well as synergistic impacts.[85]

Cetaceans

The access of microbial samples from the gut out of marine mammals is limited because most species are rare, endangered, and deep divers. There are different techniques for sampling the cetacean's gut microbiome. The most common is collecting fecal samples from the environment and taking a probe from the center that is non-contaminated.[88] Besides there are studies from rectal swabs and rare studies from stranded dead or living animals direct from the intestine.[89][90][91]
The outermost epidermal layer, i.e. the skin, is the first barrier that protects the individual from the outside world and the epidermal microbiome on it is considered an indicator not only of the health of the animal but is also considered an ecological indicator that shows the state of the surrounding environment. Knowing the microbiome of the skin of marine mammals under ''normal'' conditions has allowed us to understand how these communities are different from the free microbial communities found in the sea and how they can change according to abiotic and biotic variations, and also ''communities vary between healthy and sick individuals''.[92]
Cetaceans are in danger because they are affected by multiple stress factors which make them more vulnerable to various diseases. These animals have been noted to show high susceptibility to airway infections, but very little is known about their respiratory microbiome. Therefore, the sampling of the exhaled breath or "blow" of the cetaceans can provide an assessment of the state of health. Blow is composed of a mixture of microorganisms and organic material, including lipids, proteins , and cellular debris derived from the linings of the airways which, when released into the relatively cooler outdoor air, condense to form a visible mass of vapor, which can be collected. There are various methods for collecting exhaled breath samples, one of the most recent is through the use of aerial drones. This method provides a safer, quieter, and less invasive alternative and often a cost-effective option for monitoring fauna and flora. Once obtained, the blow samples are taken to the laboratory and we proceed with the amplification and sequencing of the respiratory tract microbiota. The use of aerial drones has been more successful with large cetaceans due to slow swim speeds and larger blow sizes.[87][93][94][95][96][86][97][98][99]
Marine holobionts
Reef-building corals are holobionts that include the coral itself (a eukaryotic invertebrate within class Anthozoa), photosynthetic dinoflagellates called zooxanthellae (Symbiodinium), and associated bacteria and viruses.[100] Co-evolutionary patterns exist for coral microbial communities and coral phylogeny.[101]
References
- ^ doi:10.3389/fmars.2017.00222..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ PMID 31710600..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ PMID 23363627.
- ^ PMID 27103626.
- PMID 1722662.
- PMID 18764872.
- ^ S2CID 21583331.
- PMID 31132110.
- PMID 30842763.
- S2CID 9729436.
- ^ PMID 23391737.
- PMID 21443739.
- S2CID 10101482.
- PMID 31213707.
- S2CID 4459856.
- S2CID 4333166.
- PMID 30510004.
- PMID 29610704.
- PMID 30723123.
- S2CID 4328049.
- S2CID 90469901.
- PMID 29379177.
- PMID 28835538.
- S2CID 42138428.
- PMID 20124339.
- S2CID 14401680.
- PMID 31239380.
- PMID 19828317.
- S2CID 14731587.
- PMID 26834723.
- PMID 17080611.
- S2CID 85817449.
- S2CID 28850526.
- ^ .
- S2CID 4462055.
- .
- PMID 29331767.
- PMID 24995875.
- S2CID 23501584.
- PMID 23278392.
- ^ PMID 29523192..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - S2CID 83963124.
- S2CID 34459128.
- ^ S2CID 85962482.
- ^ PMID 28099457.
- PMID 23288898.
- S2CID 6720678.
- PMID 25825737.
- PMID 29632094.
- hdl:1956/4610.
- S2CID 195355739.
- S2CID 27806510.
- PMID 27775707.
- .
- .
- .
- .
- .
- .
- .
- doi:10.1038/35077067.
- ^ .
- .
- .
- .
- .
- PMID 28915923.
- ^ Dubinsky, Z. and Jokiel, P.L. (1994) "Ratio of energy and nutrient fluxes regulates symbiosis between zooxanthellae and corals". Pacific Science, 48(3): 313–324.
- .
- .
- .
- .
- .
- ^ USGS scientists publish long-read microbiome sequences from temperate coral, providing community resource for probe and primer design, United States Geological Survey, 6 March 2019.
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- PMID 31384703..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- ^ S2CID 86518859.
- ^ .
- PMID 31210263.
- PMID 33776947.
- PMID 26839246.
- S2CID 226302293.
- PMID 23761254.
- PMID 29034331.
- PMID 31260051.
- PMID 23765934.
- PMID 32614926.
- PMID 28341851.
- PMID 23757234.
- PMID 29865228.
- S2CID 24127308.
- PMID 30467310.
- PMID 25621279.
- PMID 29244764.Creative Commons Attribution 4.0 International License
 Modified text was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/
Modified text was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ - PMID 30249182..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Further references
- Stal, L. J. and Cretoiu, M. S. (Eds.) (2016) The marine microbiome: an untapped source of biodiversity and biotechnological potential Springer. ISBN 9783319330006.
- Marine Microbiome and Biogeochemical Cycles in Marine Productive Areas. Frontiers Media S.A. 2020. OCLC 1291256407.


![Coral holobiont[102]](http://upload.wikimedia.org/wikipedia/commons/thumb/2/26/Trophic_connections_of_the_coral_holobiont_in_the_planktonic_food_web.jpg/271px-Trophic_connections_of_the_coral_holobiont_in_the_planktonic_food_web.jpg)
![Seagrass holobiont[103]](http://upload.wikimedia.org/wikipedia/commons/thumb/3/35/Processes_within_the_seagrass_holobiont.webp/225px-Processes_within_the_seagrass_holobiont.webp.png)
![Sponge holobiont[41]](http://upload.wikimedia.org/wikipedia/commons/thumb/6/6b/The_sponge_holobiont.webp/347px-The_sponge_holobiont.webp.png)
![Climate change and the rhodolith holobiont[104]](http://upload.wikimedia.org/wikipedia/commons/thumb/f/f7/Climate_change_stressors_and_rhodolith_holobiont_fitness.webp/474px-Climate_change_stressors_and_rhodolith_holobiont_fitness.webp.png)