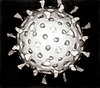
Marine viruses
Marine viruses are defined by their habitat as
When not inside a cell or in the process of infecting a cell, viruses exist in the form of independent particles called
A teaspoon of seawater typically contains about fifty million viruses.
| Part of a series of overviews on |
| Marine life |
|---|
 |
Background
| Part of a series on |
| Plankton |
|---|
 |
Viruses are now recognised as ancient and as having origins that pre-date the divergence of life into the
Opinions differ on whether viruses are a form of life or organic structures that interact with living organisms.[11] They are considered by some to be a life form, because they carry genetic material, reproduce by creating multiple copies of themselves through self-assembly, and evolve through natural selection. However they lack key characteristics such as a cellular structure generally considered necessary to count as life. Because they possess some but not all such qualities, viruses have been described as replicators[12] and as "organisms at the edge of life".[13]
The existence of viruses in the ocean was discovered through
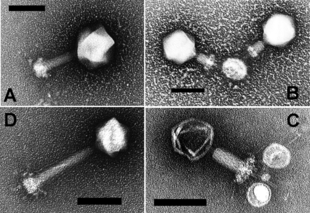
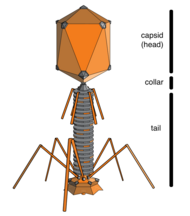
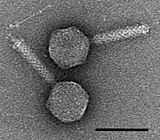
Bacteriophages
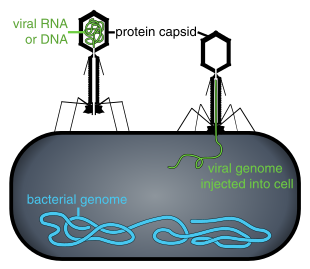

interacting with the marine Prochlorococcus MED4 bacterium
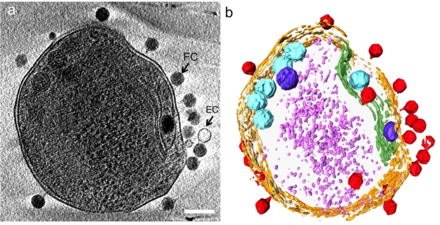
(b) same image visualised by highlighting the cell wall in orange, the plasma membrane in light yellow, the thylakoid membrane in green, carboxysomes in cyan, the polyphosphate body in blue, adsorbed phages on the sides or top of the cell in red, and cytoplasmic granules (probably mostly ribosomes) in light purple.[21]
Bacteria defend themselves from bacteriophages by producing enzymes that destroy foreign DNA. These enzymes, called
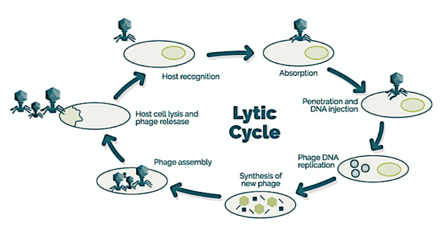
→ attachment: the phage attaches itself to the surface of the host cell
→ penetration: the phage injects its DNA through the cell membrane
→ transcription: the host cell's DNA is degraded and the cell's metabolism
is directed to initiate phage biosynthesis
→ biosynthesis: the phage DNA replicates inside the cell
→ maturation: the replicated material assembles into fully formed viral phages
→ lysis: the newly formed phages are released from the infected cell
(which is itself destroyed in the process) to seek out new host cells [26]
Microbes drive the nutrient transformations that sustain Earth's ecosystems,
For a long time, tailed phages of the order Caudovirales seemed to dominate marine ecosystems in number and diversity of organisms.[17] However, as a result of more recent research, non-tailed viruses appear to dominate multiple depths and oceanic regions.[31] These non-tailed phages also infect marine bacteria, and include the families Corticoviridae,[32]
As of September 2023, Halomonas phage vB HmeY H4907 is the first virus isolated from the deepest part of the ocean.[39]
Archaeal viruses
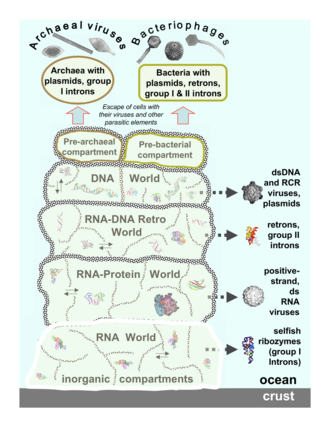
Archaean viruses replicate within
Fungal viruses
- See, Nerva L, Ciuffo M, Vallino M, Margaria P, Varese G, Gnavi G, Turina M (2016). "Multiple approaches for the detection and characterization of viral and plasmid symbionts from a collection of marine fungi". Virus Research. 219: 22–38. S2CID 53417720.
Eukaryote viruses

Marine protists
By 2015, about 40 viruses affecting
Phycodnaviridae play important ecological roles by regulating the growth and productivity of their algal hosts. Algal species such Heterosigma akashiwo and the genus Chrysochromulina can form dense blooms which can be damaging to fisheries, resulting in losses in the aquaculture industry.[53] Heterosigma akashiwo virus (HaV) has been suggested for use as a microbial agent to prevent the recurrence of toxic red tides produced by this algal species.[54] The coccolithovirus Emiliania huxleyi virus 86, a giant double-stranded DNA virus, infects the ubiquitous coccolithophore Emiliania huxleyi.[50][51] This virus has one of the largest known genomes among marine viruses.[55] Phycodnaviridae cause death and lysis of freshwater and marine algal species, liberating organic carbon, nitrogen and phosphorus into the water, providing nutrients for the microbial loop.[56]
The virus-to-prokaryote ratio, VPR, is often used as an indicator of the relationship between viruses and hosts. Studies have used VPR to indirectly infer virus impact on marine microbial productivity, mortality, and biogeochemical cycling.[57] However, in making these approximations, scientists assume a VPR of 10:1, the median observed VPR in the surface ocean.[57][18] The actual VPR varies greatly depending on location, so VPR may not be the accurate proxy for viral activity or abundance as it has been treated.[57][58]
Marine invertebrates
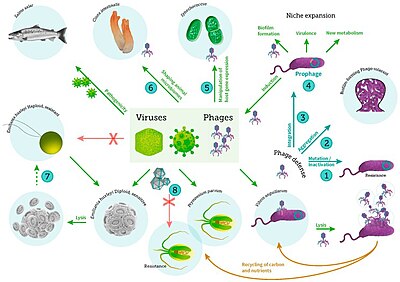
including viral infection of bacteria, phytoplankton and fish[59]
Marine invertebrates are susceptible to viral diseases.[60][61][62] Sea star wasting disease is a disease of starfish and several other echinoderms that appears sporadically, causing mass mortality of those affected.[63] There are around 40 different species of sea stars that have been affected by this disease. In 2014 it was suggested that the disease is associated with a single-stranded DNA virus now known as the sea star-associated densovirus (SSaDV); however, sea star wasting disease is not fully understood.[64]
Marine vertebrates
In 1984,
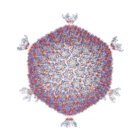
Giant marine viruses
Most viruses range in length from about 20 to 300 nanometers. This can be contrasted with the length of bacteria, which starts at about 400 nanometers. There are also giant viruses, often called giruses, typically about 1000 nanometers (one micron) in length. All giant viruses belong to the phylum Nucleocytoviricota (NCLDV), together with poxviruses. The largest known of these is Tupanvirus. This genus of giant virus was discovered in 2018 in the deep ocean as well as a soda lake, and can reach up to 2.3 microns in total length.[70]
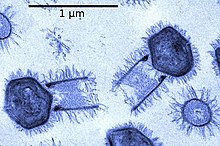
The discovery and subsequent characterization of giant viruses has triggered some debate concerning their evolutionary origins. The two main hypotheses for their origin are that either they evolved from small viruses, picking up DNA from host organisms, or that they evolved from very complicated organisms into the current form which is not self-sufficient for reproduction.[72] What sort of complicated organism giant viruses might have diverged from is also a topic of debate. One proposal is that the origin point actually represents a fourth domain of life,[73][74] but this has been largely discounted.[75][76]
Virophages
Virophages are small, double-stranded DNA viruses that rely on the co-infection of giant viruses. Virophages rely on the viral replication factory of the co-infecting giant virus for their own replication. One of the characteristics of virophages is that they have a parasitic relationship with the co-infecting virus. Their dependence upon the giant virus for replication often results in the deactivation of the giant viruses. The virophage may improve the recovery and survival of the host organism. Unlike other satellite viruses, virophages have a parasitic effect on their co-infecting virus. Virophages have been observed to render a giant virus inactive and thereby improve the condition of the host organism.
All known virophages are grouped into the family
Most of these virophages were discovered by analyzing metagenomic data sets. In metagenomic analysis, DNA sequences are run through multiple bioinformatic algorithms which pull out certain important patterns and characteristics. In these data sets are giant viruses and virophages. They are separated by looking for sequences around 17 to 20 kbp long which have similarities to already sequenced virophages. These virophages can have linear or circular double-stranded DNA genomes.[80] Virophages in culture have icosahedral capsid particles that measure around 40 to 80 nanometers long.[81] Virophage particles are so small that electron microscopy must be used to view these particles. Metagenomic sequence-based analyses have been used to predict around 57 complete and partial virophage genomes[82] and in December 2019 to identify 328 high-quality (complete or near-complete) genomes from diverse habitats including the human gut, plant rhizosphere, and terrestrial subsurface, from 27 distinct taxonomic clades.[83]
A giant marine virus
Role of marine viruses
Although marine viruses have only recently been studied extensively, they are already known to hold critical roles in many ecosystem functions and cycles.
Viral shunt
The dominant hosts for viruses in the ocean are marine microorganisms, such as bacteria.
In this way, marine viruses are thought to play an important role in nutrient cycles by increasing the efficiency of the
Viral activity also enhances the ability of the biological pump to
The

Viruses are the most abundant biological entity in marine environments.
Limiting algal blooms
Microorganisms make up about 70% of the marine biomass.[104] It is estimated viruses kill 20% of the microorganism biomass each day and that there are 15 times as many viruses in the oceans as there are bacteria and archaea. Viruses are the main agents responsible for the rapid destruction of harmful algal blooms,[105] which often kill other marine life.[106] Scientists are exploring the potential of marine cyanophages to be used to prevent or reverse eutrophication. The number of viruses in the oceans decreases further offshore and deeper into the water, where there are fewer host organisms.[107]
Gene transfer
Marine bacteriophages often contain auxiliary metabolic genes, host-derived genes thought to sustain viral replication by supplementing host metabolism during viral infection.[108] These genes can impact multiple biogeochemical cycles, including carbon, phosphorus, sulfur, and nitrogen.[109][110][111][112]
Viruses are an important natural means of
Marine habitats
Along the coast
At the ocean surface

In the water column
Marine viral activity presents a potential explanation of the paradox of the plankton proposed by George Hutchinson in 1961.[116] The paradox of the plankton is that many plankton species have been identified in small regions in the ocean where limited resources should create competitive exclusion, limiting the number of coexisting species.[116] Marine viruses could play a role in this effect, as viral infection increases as potential contact with hosts increases.[4] Viruses could therefore control the populations of plankton species that grow too abundant, allowing a wide diversity of species to coexist.[4]
In sediments
Marine bacteriophages play an important role in deep sea ecosystems. There are between 5x1012 and 1x1013 phages per square metre in deep sea sediments and their abundance closely correlates with the number of prokaryotes found in the sediments. They are responsible for the death of 80% of the prokaryotes found in the sediments, and almost all of these deaths are caused by cell lysis (bursting). This allows nitrogen, carbon, and phosphorus from the living cells to be converted into dissolved organic matter and detritus, contributing to the high rate of nutrient turnover in deep sea sediments. Because of the importance of deep sea sediments in biogeochemical cycles, marine bacteriophages influence the carbon, nitrogen and phosphorus cycles. More research needs to be done to more precisely elucidate these influences.[117]
In hydrothermal vents
Polar regions
In addition to varied topographies and in spite of an extremely cold climate, the polar aquatic regions are teeming with
Distribution
Viruses are highly host specific.[128] A marine virus is more likely to infect cooccurring organisms, those that live in the same region the virus lives in.[129] Therefore, biogeography is an important factor in a virion's ability to infect.
Knowledge of this variation in viral populations across spatiotemporal and other environmental gradients is supported by viral morphology, as determined by transmission electron microscopy (TEM). Non-tailed viruses appear to be dominant in multiple depths and oceanic regions, followed by the Caudovirales
Metagenomic approaches to assess viral diversity are often limited by a lack of reference sequences, leaving many sequences unannotated.[133] However, viral contigs are generated through direct sequencing of a viral fraction, typically generated after 0.02-um filtration of a marine water sample, or through bioinformatics approaches to identify viral contigs or viral genomes from a microbial metagenome. Novel tools to identify putative viral contigs, such as VirSorter[134] and VirFinder,[135] allow for the assessment of patterns of viral abundance, host range, and functional content of marine bacteriophage.[136][137]
See also
References
- S2CID 2917222.
- ^ These Tiny Organisms Have Some Really Weird Shapes, National Geographic, 12 November 2016.
- ^ ISBN 978-0-7637-2932-5.
- ^ )
- ^ PMID 16984643..
{{cite journal}}: CS1 maint: multiple names: authors list (link) Modified text was copied from this source, which is available under a Creative Commons Attribution 2.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 2.0 International License - S2CID 4370363.
- ISBN 978-0-12-375146-1.
- PMID 16494962.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 20660197.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - ^ PMID 12941415.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 26965225.
- PMID 26965225.
- ^ Rybicki, EP (1990). "The classification of organisms at the edge of life, or problems with virus systematics". South African Journal of Science. 86: 182–186.
- ^ PMID 15884981.
- PMID 8919794.
- PMID 9872758.
- ^ PMID 15884981.
- ^ PMID 10704475.
- S2CID 4271861.
- ^ Shors pp. 595–97
- ^ PMID 28281671.
- PMID 8336674.
- S2CID 3888761.
- PMID 18703739.
- S2CID 42827598.
- ^ How do bacteriophages reproduce? University of Barcelona. Retrieved 12 July 2020.
- S2CID 2844984.
- S2CID 32998525.
- S2CID 20738145.
- PMID 32518192.
- ^ PMID 23635867.
- PMID 17634101.
- PMID 22020507.
- PMID 22808158.
- S2CID 4462007.
- ^ Scientists Find New Type of Virus in World's Oceans: Autolykiviridae, on: sci-news, 25 January 2018
- ^ Never-Before-Seen Viruses With Weird DNA Were Just Discovered in The Ocean, on: sciencealert, 25 January 2018
- ^ NCBI: Autolykiviridae (family) – unclassified dsDNA viruses
- PMID 37728551.
- PMID 19158076.
- S2CID 9915859.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - S2CID 20018642.
- S2CID 27481111.
- PMID 16545108.
- PMID 24909109.
- ISBN 978-1-4051-3645-7.
- ISBN 978-4-431-55130-0.
- PMID 24278736...
 Modified text was copied from this source, which is available under a Creative Commons Attribution 3.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 3.0 International License - PMID 22360532.
- ^ a b "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ^ a b ICTV. "Virus Taxonomy: 2014 Release". Retrieved 15 June 2015.
- PMID 25349393.
- S2CID 23152411.
- PMID 10049839.
- ^ Largest known viral genomes Giantviruses.org. Accessed: 11 June 2020.
- ISBN 9780470026472.
- ^ doi:10.1101/025544.
- S2CID 3463306.
- PMID 29057790.
- S2CID 30161955.
- ISBN 9781118024584.
- PMID 28189502.
- ^ Dawsoni, Solaster. "Sea Star Species Affected by Wasting Syndrome." Pacificrockyintertidal.org Seastarwasting.org (n.d.): n. pag. Ecology and Evolutionary Biology. Web.
- ^ "Sea Star Wasting Syndrome | MARINe". eeb.ucsc.edu. Retrieved 3 June 2018.
- ^ ISBN 978-0-12-511340-3.
- ^ New Brunswick to help Chile beat disease Fish Information and Services
- ^ Fact Sheet - Atlantic Salmon Aquaculture Research Archived December 29, 2010, at the Wayback Machine Fisheries and Oceans Canada. Retrieved 12 May 2009.
- .
- ^ S2CID 4658457.
- PMID 29487281.
- PMID 28710447.
- ^ Bichell RE. "In Giant Virus Genes, Hints About Their Mysterious Origin". All Things Considered.
- .
- PMID 22482024.
- S2CID 206655792.
- PMID 30837339.
- PMID 30897185.
- ^ S2CID 206530677.
- ^ S2CID 4458402.
- PMID 25184667.
- S2CID 14196910– via Springer.
- PMID 29021524.
- PMID 31823797.
- PMID 30897185...
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - PMID 20974979.
- PMID 20974979.
- S2CID 30191542.
- ^ Herndl G, Briand F, eds. (2003). Ecology of marine viruses. CIESM Workshop Monograph 21 [1]
- ISBN 978-1-55581-307-9.
- S2CID 134263396.
- ^ Weitz JS, Wilhelm SW (2013). "An ocean of viruses". The Scientist. 27 (7): 35–39.
- S2CID 4370363.
- JSTOR 1313569.
- S2CID 4658457.
- S2CID 4370363.
- .
- PMID 17298376.
- ^ Robinson, Carol, and Nagappa Ramaiah. "Microbial heterotrophic metabolic rates constrain the microbial carbon pump." The American Association for the Advancement of Science, 2011.
- .
- PMID 31749771.
- S2CID 1260399.
- ^ Tsai, An-Yi, Gwo-Ching Gong, and Yu-Wen Huang. "Importance of the Viral Shunt in Nitrogen Cycling in Synechococcus Spp. Growth in Subtropical Western Pacific Coastal Waters." Terrestrial, Atmospheric & Oceanic Sciences25.6 (2014).
- JSTOR 1313569.
- PMID 29784790.
- ^ S2CID 4370363.
- ^ "Harmful Algal Blooms: Red Tide: Home|CDC HSB". www.cdc.gov. Retrieved 19 December 2014.
- ^ S2CID 4658457.
- .
- PMID 27088500.
- PMID 24200126.
- S2CID 692770.
- PMID 25171894.
- PMID 11536914.
- S2CID 20194876.
- ^ PMID 30813345...
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ S2CID 86353285.
- S2CID 4331430.
- PMID 22084639.
- PMID 25279954.
- ISSN 0967-0637.
- PMID 22457982.
- S2CID 10737747.
- ^ PMID 30813316..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - PMID 21130655.
- S2CID 33598792.
- S2CID 32607904.
- S2CID 2927907.
- PMID 23968968.
- PMID 23178671.
- PMID 17634101.
- PMID 22020507.
- ^ PMID 22808158.
- PMID 23468974.
- PMID 26038737.
- PMID 28683828.
- S2CID 4466854.
- PMID 28677677.



![Cryo-electron micrograph of the CroV giant virus [71] scale bar=0.2 µm](http://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/CroV_TEM_%28cropped%29.jpg/270px-CroV_TEM_%28cropped%29.jpg)